In recent years, the percentage of elderly people in the human population has grown rapidly. Data from the Turkish Statistical Agency indicate that 8.7 % of the population was older than 65 in 2018, compared to 4.7 % in 1980 [1]. According to the Statistical Agency, by 2080, people over 65 will make up a quarter of the entire population. The number of Alzheimer’s disease (AD) patients has increased due to the aging population. According to AD International’s (ADI) annual report, approximately 50 million people suffer from AD worldwide [2], [3]. Furthermore, they predict that this number will double every 20 years. As AD is a growing problem, developing new and effective methods for early diagnosis and treatment is becoming increasingly important and popular. Because brain tissue and nerve cells are lost, patients experience memory loss and cognitive impairment.
Globally, by 2050, one in 85 people will have AD or another form of dementia [4], [5]. Other than therapies that slow disease progression, no alternative medication has been identified to stop or cure the disease, despite predictions that nursing and treatment expenses will rise dramatically with the increasing number of patients [6]. Since AD is a degenerative brain disease, nerve cells and tissue gradually disappear as the illness progresses [7]. Therefore, an essential component of early AD diagnosis is the identification of mild cognitive impairment (MCI), which is considered a precursor to AD [8]. Magnetic resonance imaging (MRI) and other forms of brain imaging, which allow for the visualization of the structure and function of the brain, have been used in the medical diagnosis of brain disorders [9].
To diagnose AD dementia, doctors assess the signs and symptoms of AD and use various tests. Physicians may also prescribe memory tests, brain imaging studies, or other laboratory tests [10]. By excluding other illnesses that might cause similar symptoms, these tests help medical professionals diagnose patients. MRI can detect brain abnormalities associated with MCI [11] and can also help predict which individuals with MCI may eventually develop AD. MRI scans can identify irregularities, such as a reduction in the size of certain brain regions, primarily affecting the parietal and temporal lobes [12]. AD is a neurological condition that causes memory loss and dementia. Accurate and early diagnosis remains a significant challenge in medical practice, as traditional diagnostic methods often detect AD in its later stages. To address these challenges, a novel AD-HOLDER model has been proposed for detecting AD using MRI and positron emission tomography (PET) images. MRI and PET imaging offer valuable insights into brain structure and function, but manually analyzing these complex datasets is time-consuming and prone to errors. Machine learning (ML) [13] techniques automate the detection process, leveraging large datasets to identify subtle patterns associated with AD progression. The main contributions of this research are as follows:
The novel AD-HOLDER model integrates MRI and PET images, leveraging both structural and statistical features. The dual-input images improve the ability of the model to capture complete neurological patterns crucial for early AD diagnosis.
A Histograms of Oriented Gradients (HOG) method is introduced, combining residual learning and multi-branch convolution to extract rich spatial-contextual features. This approach effectively fuses complementary features from MRI and PET, improving the quality of representations used in classification.
A graph-based segmentation (GBS) approach is employed to precisely segment abnormal regions, improving detection specificity (SPE) and providing localized insights into affected brain areas, as demonstrated by significant gains in dice index (DI) compared to traditional segmentation models.
The remainder of the paper is organized as follows: The literature on AD diagnosis and classification is reviewed in Part 2; the proposed approach is presented in Part 3; the experimental and assessment findings are shown in Part 4; and the study is concluded and future research is discussed in Part 5.
Many categorization strategies have been developed in various studies for the diagnosis and detection of AD. This section summarizes current research on AD diagnostic and detection systems using traditional ML and deep learning (DL) techniques.
In 2021, Battineni et al. [14] proposed an ML approach associated with brain research that provided a more accurate examination of AD. The system uses longitudinal brain MRI data to classify patients into two groups: AD or non-AD. Six distinct supervised classifiers are used to categorize individuals. With an accuracy (ACC) rate of 97.58 %, the results demonstrate that the gradient boosting strategy outperforms the other models.
In 2023, Rallabandi and Seetharaman [15] proposed an Inception-ResNet wrapper Convolutional Neural Network (CNN) model by combining information from PET and structural MRI imaging to detect AD. The DL model enhanced the automated imaging diagnostic tool’s ACC and ability to distinguish mild cognitive impairment (MCI) and AD from healthy controls (HC) using dual imaging modalities. The proposed model performs as the best classifier, detecting HC, MCI, and AD with accuracies of 95.5 %, 94.1 %, and 95.7 %, respectively.
In 2023, Odusami et al. [16] proposed integrating neuroimaging information from MRI and PET images to diagnose early AD. Effective binary categorization of AD is achieved using a specific 3-in-channel technique that extracts the most descriptive information from combined PET and MRI data. According to the testing results, the proposed model achieved a 73.90 % classification ACC using the AD Neuroimaging Initiative (ADNI) database. The results were obtained with a model of Explainable Artificial Intelligence (XAI).
In 2022, AlSaeed and Omar [17] presented a pre-trained CNN model based on ResNet50 for AD diagnosis using MR images. The efficacy of a CNN utilizing standard Softmax, Support Vector Machine (SVM), and Random Forest (RF) is then assessed using a range of parameters. The results demonstrated that the model outperformed previous state-of-the-art models, with models using the ADNI dataset exhibiting an ACC range of 85.7 % to 99 %.
In 2022, Amini et al. [18] demonstrated the use of single-nucleotide polymorphism markers (SNPs) to detect AD using PET images. Furthermore, SVM, Linear Discriminant Analysis (LDA), K-Nearest Neighbors (KNN), and CNN approaches were used to classify AD based on the frequency of alterations in brain tissue in PET images. The proposed SNPs appear to have a stronger correlation with quantitative traits than the ApoE gene SNPs, based on the data. In terms of the categorization outcome, CNN achieves the highest ACC at 91.1 %.
In 2022, AA et al. [19] proposed a Computer-Aided Diagnosis (CAD) system that distinguishes between AD and NC patients based on features extracted from 18FDG-PET images. To extract these features, several 2D slices were taken from the FDG-PET images. According to the results, the proposed CAD system performs admirably compared to existing approaches and detailed in the literature, achieving ACC, sensitivity, and SPE of 96 %, 94 %, and 96 %, respectively.
In 2021, Murugan et al. [20] presented a CNN architecture for AD categorization. Using the standard Kaggle dataset for dementia stage categorization, the model was trained and validated. It can accurately identify brain areas linked to AD and serves as a powerful decision support tool for doctors, helping them estimate the severity of AD based on the degree of dementia. The ACC of the model, evaluated using the ADNI dataset, was 84.83 %, demonstrating its robustness.
In 2024, Castellano et al. [21] proposed a unimodal and multimodal framework utilizing amyloid PET scans and 2D and 3D MRI images. Models that learn representations from volumetric data are more effective than those using 2D images. Additionally, compared to single-modality approaches, the model’s performance is significantly enhanced by mixing multiple modalities. Gradient-weighted Class Activation Mapping (Grad-CAM) predicts areas associated with AD, highlighting its potential as a tool for understanding the disease’s underlying causes.
In 2023, Marwa et al. [22] created a DL based pipeline for AD stage detection and classification. The proposed analysis method uses 2D T1-weighted MR brain images and a CNN architecture. The pipeline provides both global and local categorization (i.e., normal vs. AD vs. MCI), offering a fast and accurate AD diagnosis. The method further classifies MCI into three categories: Mild Dementia (MD), Moderate Dementia (MoD), and Very Mild Dementia (VMD). In 2021, Massalimova and Varol [23] used structural MRI and diffusion tensor imaging (DTI) data from the OASIS-3 dataset to classify AD, MCI, and normal cognition using a multi-modal DL technique. The results show that the agnostic model achieved an ACC of 0.96 on structural MRI and DTI images.
In 2025, Muksimova et al. [24] developed FusionNet for AD diagnosis by creatively combining longitudinal imaging data with multi-modal data. The model provides a robust framework for early identification and continuous monitoring of AD by integrating various data sources, including MRI, PET, and CT scans, and by analyzing changes over time. FusionNet achieved 94 % ACC, with 92 % precision (PRE) and 93 % recall (REC) rates.
In 2025, Mousavi et al. [25] proposed deep CNNs for AD diagnosis and categorization using MRI data. Training was enhanced through hyperparameter adjustment and generalization. Overfitting was prevented by using a dynamic learning rate and early termination. The Xception model demonstrated high PRE, REC, and F1-score (F1) with an ACC of 96.89 and values of 0.97.
According to the literature review, the effectiveness of existing methods is limited by their reliance on the laborious process of physical feature extraction and classification. Most of the studies mentioned above use MRI and PET images, which reduces performance rates. To address these challenges, early-stage AD detection and classification are necessary. In this research, a novel AD-HOLDER model is proposed for detecting AD using dual images.
In this study, the AD-HOLDER model is proposed for detecting the AD using MRI and PET images. Features are extracted using the HOG method, which captures essential patterns and edges from the images. The Light Gradient Boosting Machine (LGBM) classifier is then used to classify images as normal or abnormal. Finally, the abnormal region is segmented using a GBS model to specifically detect the region for early diagnosis of AD. The proposed AD-HOLDER methodology is shown in Fig. 1.
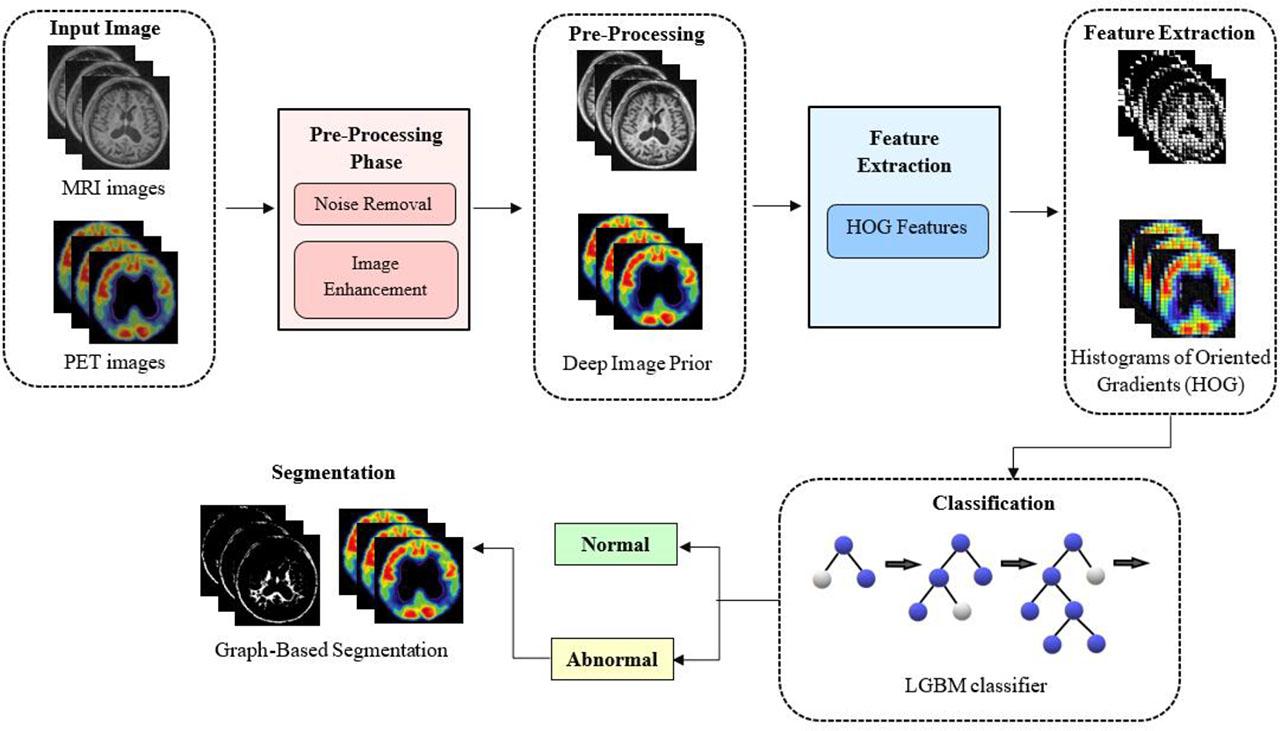
The proposed AD-HOLDER methodology.
In this study, the OASIS dataset [26] consists of MRI and PET images from a longitudinal collection of 150 subjects aged 60 to 96 years. Each individual performed a total of 373 imaging sessions, with at least one year between visits. For each patient, three or four distinct T1-weighted MRI images were obtained in a single session. The subjects include both men and women, and all are right-handed. During this analysis, 72 research participants were categorized as non-demented. At the time of their first visits, 64 of the included subjects – 51 of whom had mild to moderate AD were classified as demented, and their condition remained that way for all future scans. A total of fourteen additional participants were initially diagnosed as non-demented but later developed mental health issues.
In this section, MRI and PET images are denoised using the Deep Image Prior (DIP) filter [27] to improve the quality of both MRI and PET images for efficient BS classification. A randomly initialized DIP filter takes random noise as input and reconstructs the corrupted image. Despite the corruption, the internal structure of the filter effectively adapts to fit the clear image. Let x0 denote the corrupted image; DIP proposes the following optimization:
Here, T represents the DIP filter, z is the random noise input, x̂ is the recovered (denoised) image, and θ∗ represents the optimized filter parameters. This approach does not require conventional training data pairs. Instead, Stochastic Gradient Langevin Dynamics (SGLD) is used for posterior inference, improving denoising performance while avoiding early stopping.
The MRI and PET image restoration process estimates and removes the noise component from the input image, thereby preserving the essential structure of the image. A DIP filter with 64 filters, a 3 × 3 kernel size, and ReLU activation functions was used in the denoising step. The Adam optimizer was used to optimize the filter for up to 200 epochs at a learning rate of 0.001, with early stopping implemented after 20 consecutive epochs if validation loss did not improve. This DIP configuration effectively reduced noise artifacts while preserving clinically relevant anatomical details, ensuring high-quality inputs for subsequent feature extraction. Filters produce a clear image output that reduces noisy distortions while maintaining the integrity of significant features, improving downstream tasks such as segmentation and classification. Data augmentation, including multi-angle rotation, adding Gaussian noise, improving and reducing brightness, and horizontal and vertical mirroring, was used to increase categorization ACC.
The features are extracted using the HOG method [28], which captures essential patterns and edges from the images. HOG was used to extract structural features such as edges, contours, and textures relevant for distinguishing normal from abnormal brain tissues. A cell size of 50×50 pixels was selected to balance local detail capture and computational efficiency. In medical imaging applications, larger cells provide improved robustness against intensity variations while retaining discriminative edge information with similar parameter settings. Experimental investigations showed that 50×50 cells offer the optimal balance between ACC and processing cost. We empirically evaluated different cell sizes (20×20, 30×30, and 50×50) and found that 50×50 yielded the highest classification ACC with reduced computational cost. HOGs represented each item as a single value vector by moving the window detector across the image, rather than computing a collection of feature vectors for different image areas. To obtain a HOG feature, the image’s scale is modified while the HOG descriptor is calculated for each location.
To compute the HOG feature, the image’s gradients must be determined. Image gradients are defined as directed increases in pixel intensity along the x- and y-axes. A pixel’s gradient vector at position (y, x) is described by (5).
Here, gx and gy are the gradients in the x and y directions, respectively, and f (x, y) is the pixel intensity at coordinates x and y. The gradient’s phase, θ(x, y), and magnitude, M(x, y), can then be determined using the following formulas.
Here, gx and gy are the gradients in the x and y directions, respectively.
The LGBM classifier [29] is used to classify normal and abnormal cases using MRI and PET images. Ensemble learning includes techniques such as gradient boosting. Fig. 2 shows the architecture diagram of the HOG-based LGBM classifier. In the ensemble boosting strategy, models are constructed sequentially by continuously minimizing the error of previously trained models. When splitting and propagating a tree, it is important to select the leaf that minimizes the loss the most. LGBM uses a histogram-based method to find the best split candidates. It employs gradient-based one-side sampling (GOSS) to determine the importance of image occurrences for training, focusing more on data samples with larger gradients and less on those with smaller gradients.
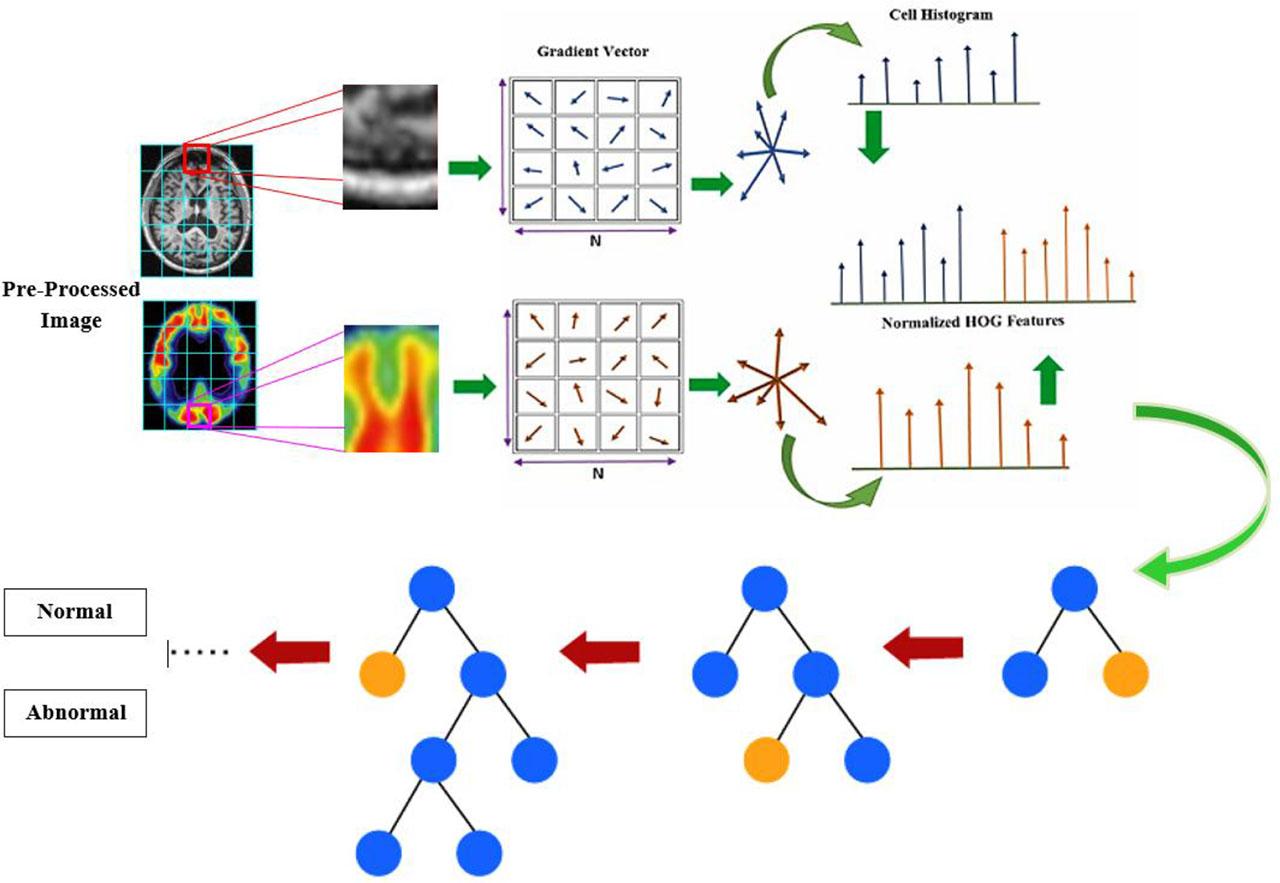
Architecture diagram of the HOG-based LGBM model.
It is assumed that data with smaller gradients have previously undergone extensive training, as they contain fewer errors. GOSS proposes discarding these less-informative data points and using the remaining ones to determine information gain when establishing the best splits. However, this changes the original data distribution and creates a bias towards samples with larger gradients. To address this, GOSS retains all samples with large gradients and randomly samples data with small gradients. When computing information gain, GOSS applies a constant multiplier to the weights of data instances with moderate gradients, as the sample remains skewed toward data with strong gradients. Additionally, LGBM regulates dataset sparsity using Exclusive Feature Bundling, which combines features almost losslessly by removing incompatible aspects and retaining the most useful components.
Table 1 presents the hyperparameter settings of the LGBM classifier. The proposed AD-HOLDER model performance was evaluated using ACC, PRE, REC, SPE, and F1. To optimize performance, the number of estimators was set to 500, the learning rate to 0.05, and the maximum tree depth to 8. The feature fraction was set to 0.8 to reduce feature correlation and improve generalization. Regularization terms were incorporated through λL1 = 0.1 and λL2 = 0.2 to control model complexity and minimize overfitting. The OASIS dataset was split into 75 % for training and 25 % for testing, with cross-validation performed on the training set to ensure robustness and prevent overfitting. Cross-validation confirmed consistent results across folds, indicating strong generalization and minimal overfitting.
Hyperparameter settings of the LGBM classifier.
| Hyper parameter | Value |
|---|---|
| Number of estimators | 500 |
| Learning rate | 0.05 |
| Maximum depth | 8 |
| Feature fraction | 0.8 |
| λL1(L1 reg) | 0.1 |
| λL2(L2 reg) | 0.2 |
| Cross-validation folds | 3 and 5 |
The abnormal region is segmented using the GBS model [30] to specifically detect the region for early diagnosis of AD. GBS is a technique for segmenting images to enhance their quality by using graphs. Here, E is the set of edges composed of two vertices (vi, vj), G = (V, E) is an undirected graph, and wij is edge weight. Edge weights in GBS indicate the degree of dissimilarity between two pixels in an image.
The term min (w(e)) in (9) denotes the lowest possible edge weight that unites two separate components, D1 and D2, and is also known as “min (w(e)).” It is necessary to use the least edge weight because employing a quantile, such as the median, results in an NP-hard computational problem.
The minimum internal difference, MInt(D1, D2) is defined as
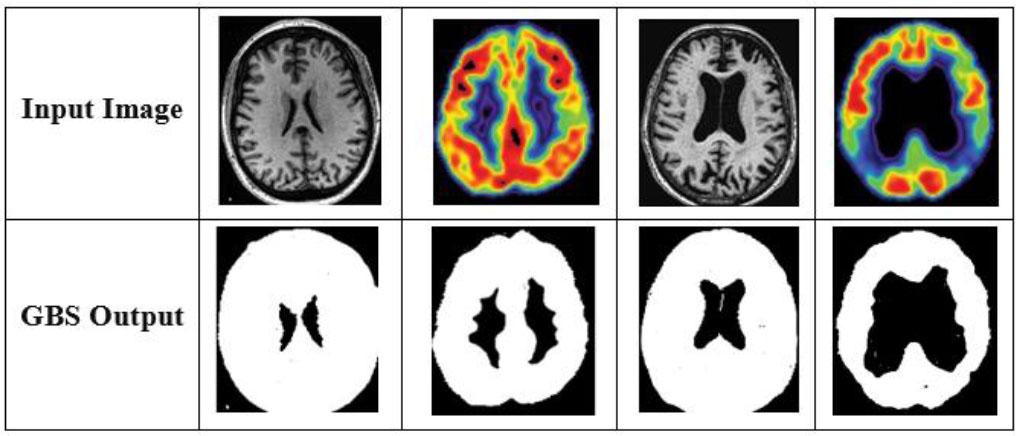
Visual examples of segmented regions.
The greatest edge weight in the MST is indicated by Int (D) in (12). According to the GBS algorithm, the default threshold function is specified as
In the GBS model, k serves as a control-scale parameter that influences the preference for merging components. Specifically, this threshold determines the minimum dissimilarity required between two regions for a boundary to be established. A higher k value promotes the formation of larger and more coherent segments by allowing greater intra-region variation before splitting occurs. This technique is particularly important for accurately describing pathological regions in brain images, where preserving structural consistency is essential for reliable disease localization and diagnosis.
The experiments, analysis, and use of the following resources, tools, and settings are described in this section: a) Setting Used; b) Scikit-learn ML libraries with Python 3. Additionally, a study of the proposed AD-HOLDER model using a HOG-based LGBM architecture is included. The OASIS dataset is divided into two subsets for the research: 75 % of the images are used for training, and the remaining 25 % are used for testing. The percentage of patients accurately diagnosed as not having AD is a measure of diagnostic ACC.
Fig. 4 illustrates the results of the proposed AD-HOLDER classification pipeline using features. Column 1 contains input dual images collected from the OASIS dataset. These dual modalities capture both structural and statistical features of the brain, which are crucial for early AD identification. Column 2 shows the enhanced images using a DIP filter to improve image quality. Features are extracted using HOG in column 3. Structural features are derived from the MRI images, representing anatomical aspects such as tissue loss. Statistical features are extracted from the PET images, capturing metabolic changes typical of Alzheimer’s progression. The dual features extracted from MRI and PET images are passed to an LGBM classifier, which categorizes them as normal or abnormal, as shown in column 4. The abnormal cases are fed into the next phase for tumor segmentation in column 5.
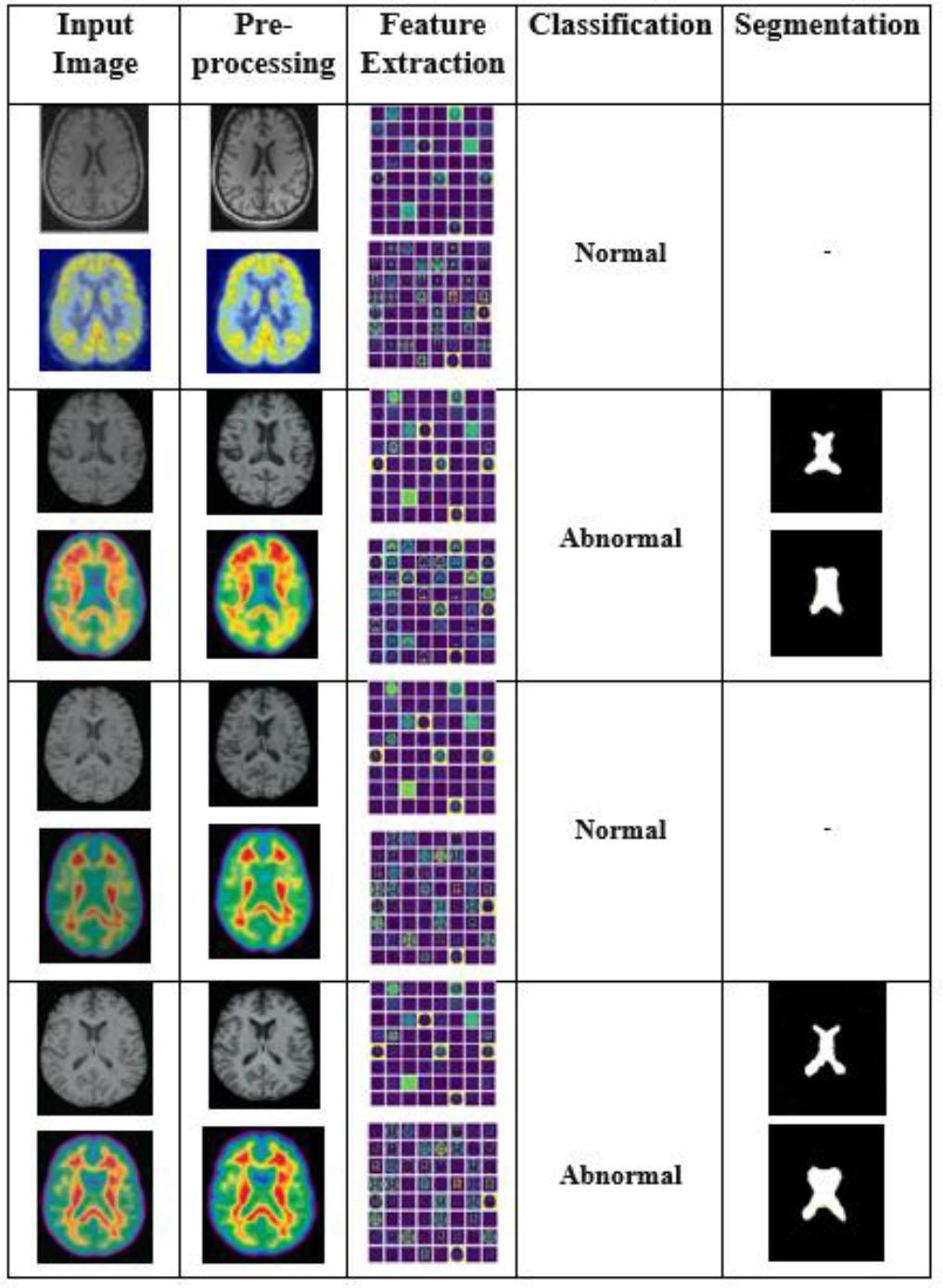
Experimental results of the proposed AD-HOLDER methodology.
The effectiveness of the AD-HOLDER approach for categorizing instances of AD is evaluated using a set of assessment criteria, including SPE, PRE, REC, ACC, and F1.
Here, Ntrue and Nfalse represent true-positives and true-negatives, while Ptrue and Pfalse denote false negatives and false positives for the MRI and CT images.
Table 2 presents the efficiency of the AD-HOLDER model using the specific parameters ACC, PRE, SPE, REC, and F1. As shown in Fig. 5, the performance of the classification for the normal and abnormal classes is illustrated across different evaluation metrics. The AD-HOLDER model achieved an ACC of 99.12 %, SPE of 97.79 %, PRE of 97.57 %, REC of 92.60 %, and F1 of 96.55 % in identifying the two types of Alzheimer’s. This model attains higher ACC in identifying the normal class than the abnormal class. The overall average ACC achieved by the proposed AD-HOLDER model is 99.12 % for AD classification based on the collected OASIS dataset.
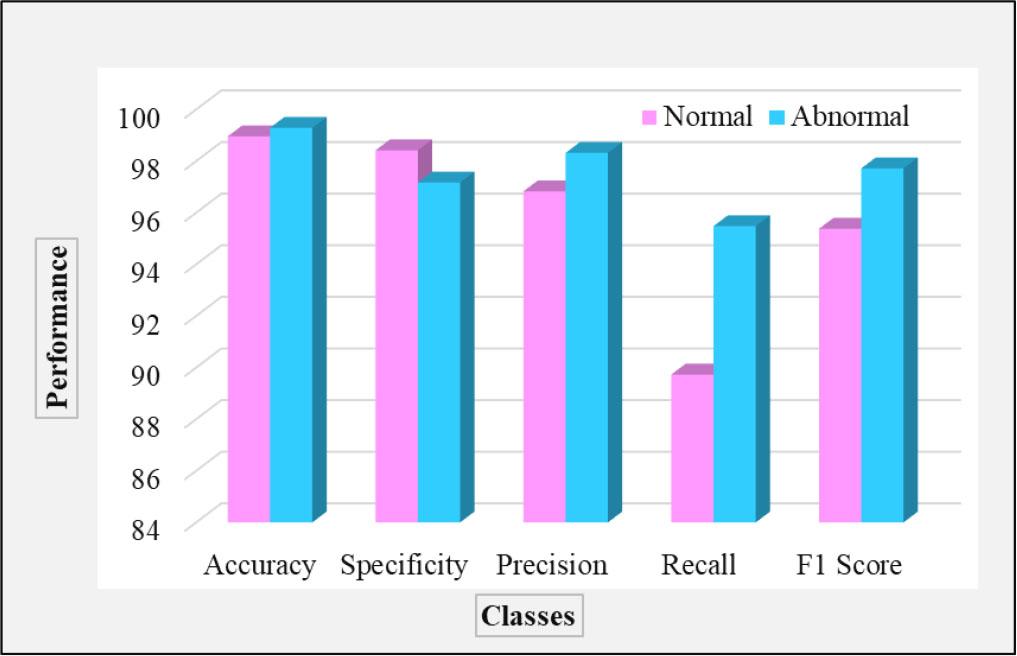
Performance analysis for two-class classification.
Efficiency assessment of the proposed AD-HOLDER model.
| Classes | ACC | SPE | PRE | REC | F1 |
|---|---|---|---|---|---|
| Normal | 98.96 | 98.42 | 96.83 | 89.73 | 95.38 |
| Abnormal | 99.29 | 97.17 | 98.32 | 95.48 | 97.72 |
The ACC curve (Fig. 6) and loss curve (Fig. 7) of the proposed model over 100 epochs show its performance in classifying AD cases. The training curve is slightly higher than the testing curve, indicating effective learning and a steady improvement in ACC over time. The proposed model consistently reduces classification errors, demonstrating its effectiveness. The testing and training curves are nearly aligned, indicating good generalization without overfitting. These results suggest that the proposed model is well-trained, with minimal loss and excellent ACC.
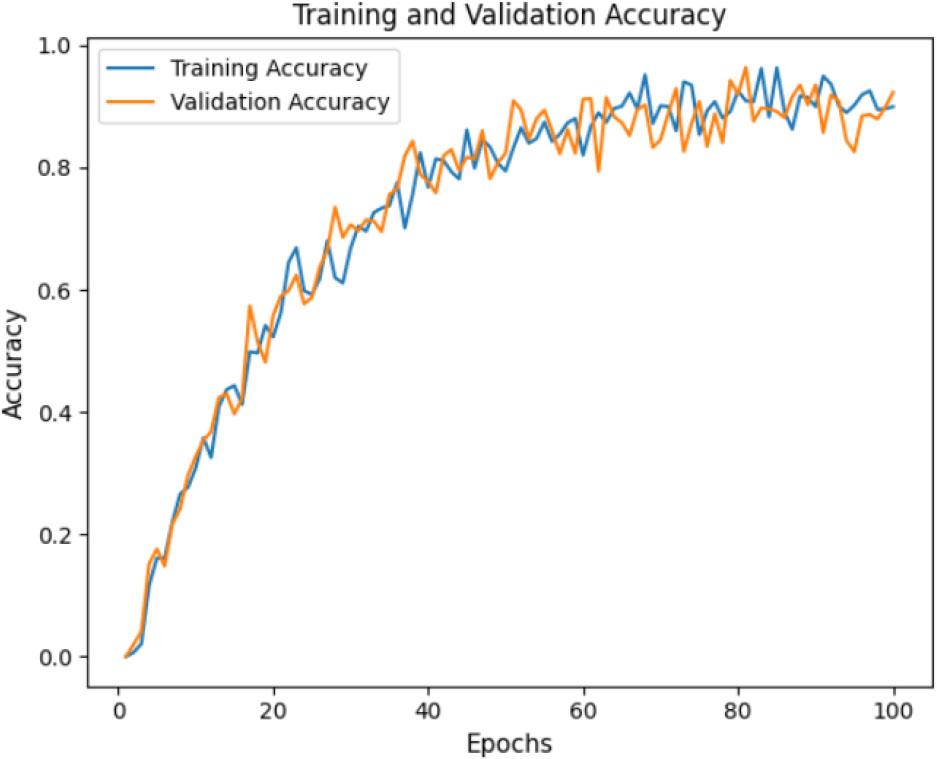
ACC curve for the proposed AD-HOLDER model.
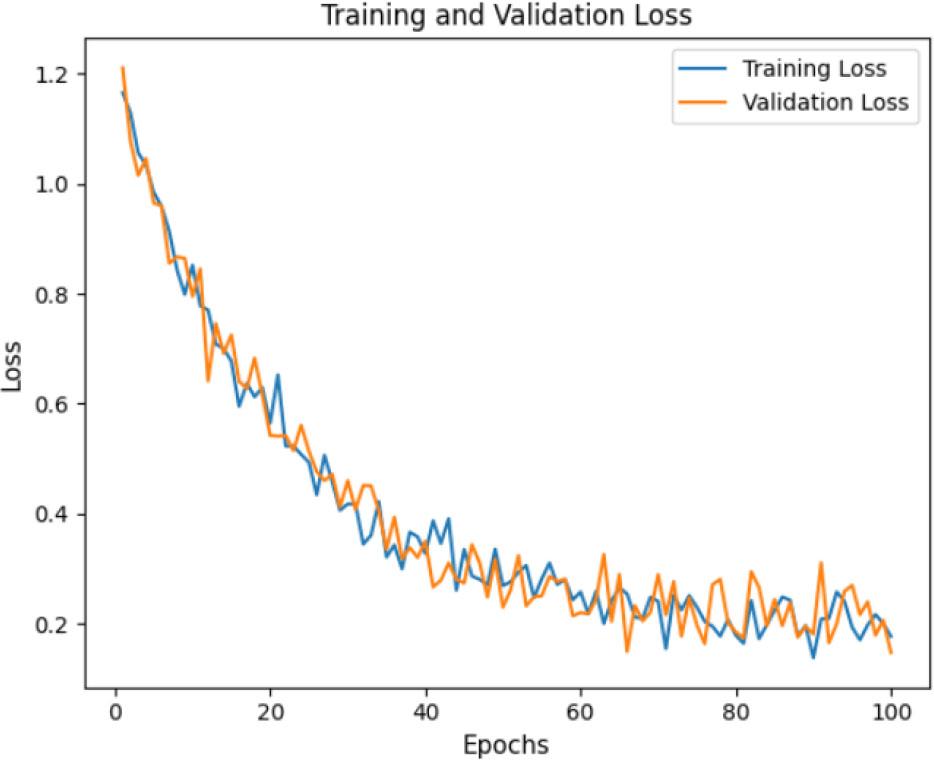
Loss curve for the proposed AD-HOLDER model.
Fig. 8 shows the classification efficiency of the proposed AD-HOLDER model for AD detection across two stages: normal and abnormal. The confusion matrix (CM)provides clinically relevant insights into the model’s performance. A false positive occurs when a normal case is misclassified as abnormal, which may lead to unnecessary anxiety. A false negative (FN) occurs when an abnormal case is misclassified as normal, causing delays in diagnosis and intervention, potentially accelerating disease progression. Minimizing false positives is crucial for reducing unnecessary evaluations, but it is also vital for early and accurate detection of AD. The CM shows the model’s high ACC, correctly classifying 98.96 % of normal and 99.29 % of abnormal samples, with low misclassification rates of 1.04 % and 0.71 %, respectively.
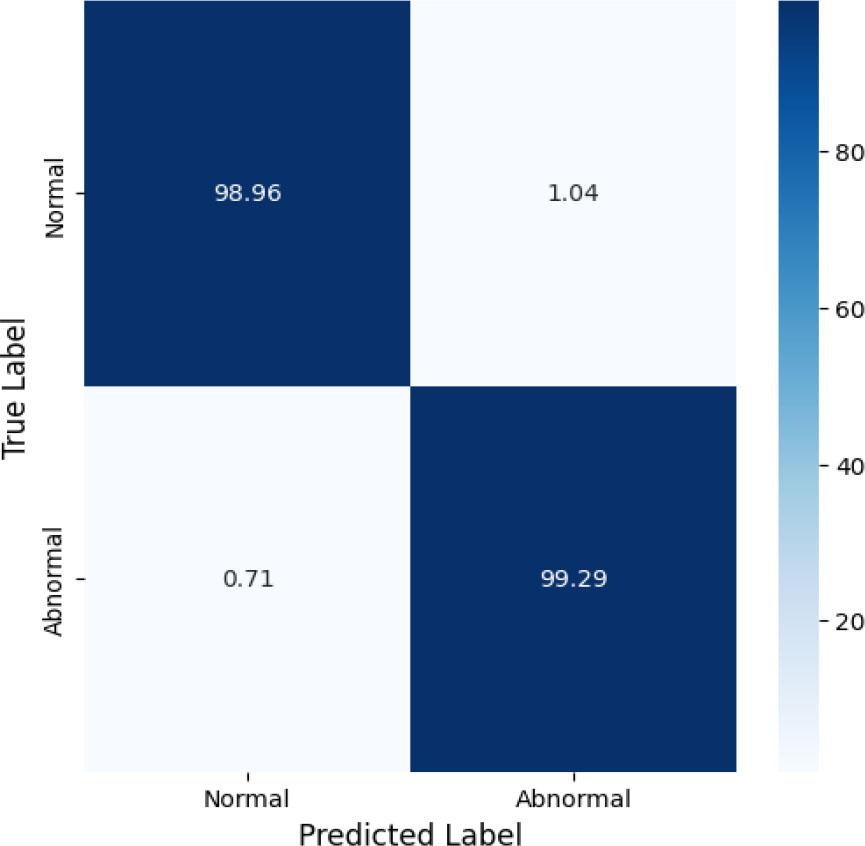
CM for two-class classification.
Fig. 9 illustrates the ROC curve of the proposed AD-HOLDER framework across Alzheimer’s stages. According to the area under curve (AUC), each curve represents the difference between the true positive rates (TPR) and false positive rates (FPR) for a particular class. The model performs poorly for the normal class (AUC = 0.98) and best for the abnormal class (AUC = 0.99), as estimated with TPR and FPR parameters. The high AUC values indicate strong discrimination capacity at every level, with only slightly lower ACC under the collected dataset.
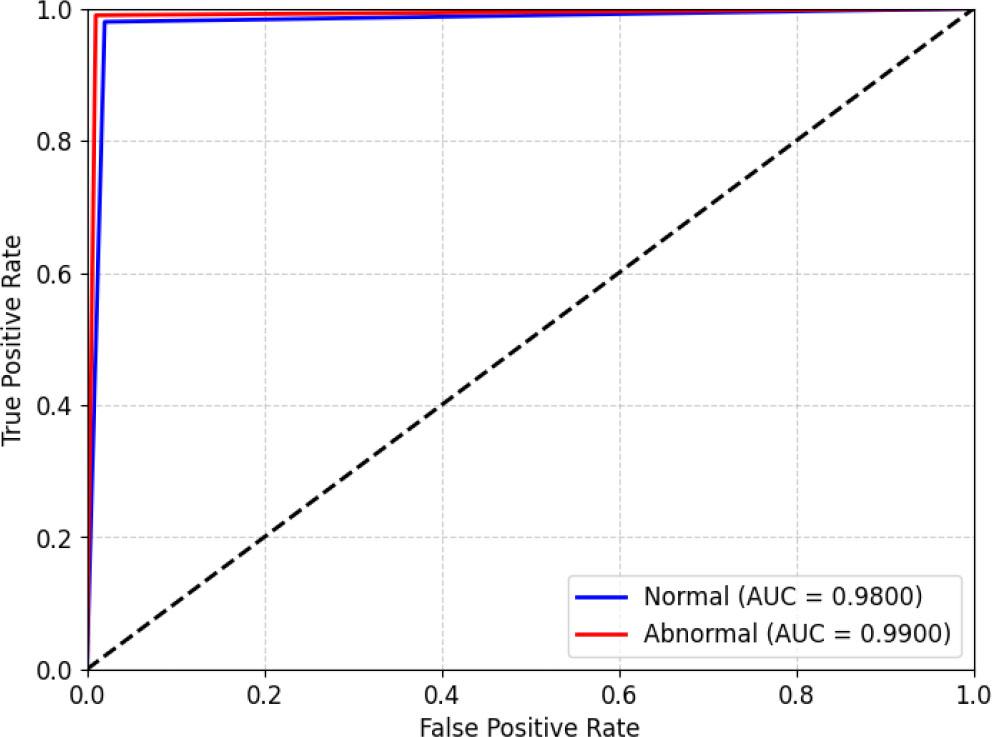
ROC curve of the proposed AD-HOLDER classification model.
Table 3 presents the AD-HOLDER model cross-validation results using 3-fold and 5-fold methods based on the OASIS dataset. For 3-fold cross-validation, the dataset was divided into three subsets, with 70 % used for training and 30 % for testing in each iteration to ensure every sample was evaluated. For 5-fold cross-validation, 80 % of the dataset was used for training and 20 % for testing, providing a robust estimate of generalization ability. Compared to the fixed 75 %-25 % split, cross-validation minimizes the risk of selection bias and yields more reliable performance evaluation. The consistent results across both 3-fold and 5-fold validation confirm the stability and robustness of the proposed model.
Cross-validation results of the proposed AD-HOLDER model.
| Metric | 3-fold cross-validation [%] | 5-fold cross-validation [%] |
|---|---|---|
| ACC | 98.27 | 98.59 |
| SPE | 96.80 | 96.14 |
| PRE | 96.17 | 96.73 |
| REC | 89.27 | 90.83 |
| F1 | 94.68 | 94.91 |
The efficacy of conventional models was evaluated to verify that the proposed AD-HOLDER model achieves a high level of ACC. The comparison of ML classifiers was conducted to assess whether the proposed model obtains better ACC. The comparative assessment was performed between the proposed model and four ML networks: KNN, Naive Bayes, Decision Tree, and RF. The performance assessment was carried out using ACC, SPE, PRE, REC, and F1 for each ML method. The ACC of the proposed AD-HOLDER model is 99.14 %, which is higher than that of the classical ML methods.
Table 4 presents a comparison of different denoising methods on MRI and PET images. Traditional filters such as Median, Gaussian, Bilateral, and Non-Local Mean (NLM) improve the peak signal-to-noise ratio (PSNR) and structural similarity index (SSIM) values compared to noisy inputs; however, they often fail to preserve fine structural details and edges in MRI and PET images. The proposed DIP method achieves the highest PSNR (31.82 dB) and SSIM (0.917), indicating superior noise suppression and structural preservation. These results confirm the effectiveness of DIP denoising for enhancing MRI and PET images prior to feature extraction.
Quantitative comparison of denoising methods on MRI and PET images.
| Method | PSNR [dB] | SSIM |
|---|---|---|
| Median filter | 27.42 | 0.712 |
| Gaussian filter | 28.06 | 0.815 |
| Bilateral filter | 28.75 | 0.822 |
| NLM filter | 29.74 | 0.858 |
| DIP (proposed) | 31.82 | 0.917 |
Table 5 presents a comparative assessment of existing classification networks. The proposed LGBM achieves the highest ACC (99.12 %) and F1 (96.55 %), demonstrating its superior performance. Decision trees and Naive Bayes perform moderately, while KNN performs lowest on all criteria, indicating its comparatively low efficacy. LGBM outperforms the other architectures across all metrics, achieving the highest ACC (99.12 %), SPE (97.79 %), PRE (97.57 %), REC (92.60 %), and F1 (96.55 %). The LGBM network achieves better results compared to the other detection networks.
Comparative analysis between traditional ML networks.
| Techniques | ACC | SPE | PRE | REC | F1 |
|---|---|---|---|---|---|
| KNN | 90.63 | 89.39 | 90.37 | 85.92 | 87.93 |
| CNN | 93.95 | 94.21 | 92.49 | 87.37 | 90.62 |
| Naive Bayes | 92.84 | 91.83 | 88.93 | 86.75 | 89.51 |
| Decision Tree | 93.92 | 93.72 | 94.93 | 88.23 | 92.73 |
| ViT | 96.38 | 92.63 | 94.26 | 90.18 | 95.29 |
| RF | 95.29 | 95.90 | 95.27 | 91.35 | 94.62 |
| LGBM | 99.12 | 97.79 | 97.57 | 92.60 | 96.55 |
The comparison was conducted with different DL-based segmentation networks using several parameters, as shown in Table 6. The proposed GBS network increases the overall DI by 26.34 %, 11.39 %, 7.66 %, and 5.38 % compared to U-net, V-net, Nested V-net, and SegNet, respectively. The proposed GBS approach outperforms all compared methods, achieving the highest ACC (99.12 %), DI (90.74 %), and intersection over union (IoU) (82.7 %). Traditional segmentation DL-networks did not perform as well as the GBS network.
Comparison of traditional segmentation algorithms [%].
| Methods | ACC | DI | IoU |
|---|---|---|---|
| U-net | 91.37 | 71.82 | 58.2 |
| V-net | 93.75 | 81.46 | 69.9 |
| Nested V-net | 95.01 | 84.28 | 72.9 |
| SegNet | 92.37 | 86.10 | 73.5 |
| GBS (ours) | 99.12 | 90.74 | 82.7 |
Fig. 10 displays the visualization outcomes of the different segmentation methods. In this evaluation, the GBS was compared with conventional networks such as V-Net and U-Net. GBS reduces processing complexity and yields better segmentation results than current segmentation techniques.
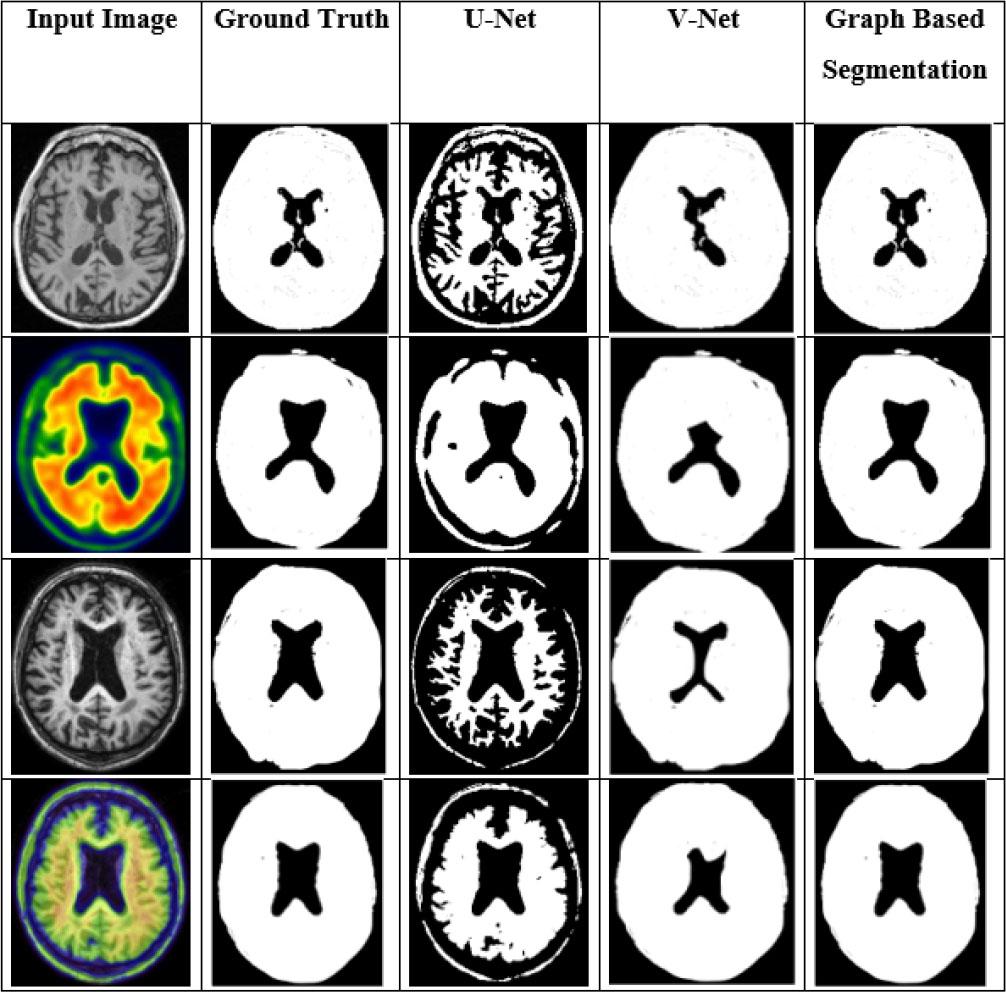
Comparison results of different segmentation techniques.
Table 7 presents a comparison of various existing methods with the proposed AD-HOLDER model using the collected OASIS dataset.
Comparison of existing models and the proposed AD-HOLDER model.
| Authors | Methods | ACC [%] | PRE [%] | REC [%] | F1 [%] | p-value |
|---|---|---|---|---|---|---|
| Battineni, G. [14] | GBA | 97.58 | 94.89 | 88.26 | 90.36 | 0.041 |
| Odusami, M., et al. [16] | XAI | 73.90 | 92.67 | 85.29 | 88.82 | 0.37 |
| Hamdi, M. [19] | CNN | 96.00 | 95.92 | 90.51 | 93.52 | 0.42 |
| Proposed model | AD-HOLDER model | 99.12 | 97.57 | 92.60 | 96.55 | 0.029 |
Battineni (2021) employed a Gradient Boosting Algorithm (GBA), achieving 97.58 % ACC. Odusami et al. (2023) used XAI, achieving 73.90 % ACC. Hamdi (2022) implemented a CNN, which attained a slightly lower accuracy (96 %). The proposed AD-HOLDER model increases the overall ACC by 1.55 %, 25.44 %, and 3.14 % for the GBA, XAI, and CNN, respectively. The reported p-values from paired t-tests are all below 0.05, indicating that the performance improvements of the AD-HOLDER model are statistically significant compared to existing approaches. The proposed AD-HOLDER model also increases the F1 by 6.85 %, 8.70 %, and 3.23 % for the GBA, XAI, and CNN, respectively. Based on this overall comparative analysis, the proposed AD-HOLDER model achieves better ACC and statistical significance.
Fig. 11 illustrates the process of detecting AD using the proposed AD-HOLDER model. A patient first visits a hospital, where MRI and PET images are collected in real time for diagnosis. These images are then processed by the AD-HOLDER model, which analyzes them to predict AD locations. In the hospital, prediction results assist the doctor in refining the patient’s diagnosis and recommending a treatment plan. This streamlined process enables real-time diagnosis, supporting timely and accurate diagnoses. The approach focuses on specific AD types, such as normal and abnormal, to simplify diagnosis and improve ACC. However, because it relies on a specific MRI and PET image dataset, the proposed model may not be generalizable to broader clinical settings and requires high-quality imaging for accurate classification. Moreover, while the proposed AD-HOLDER model categorizes AD into normal and abnormal cases, it does not distinguish between multiple disease stages, limiting its ability to provide a detailed analysis of disease progression.
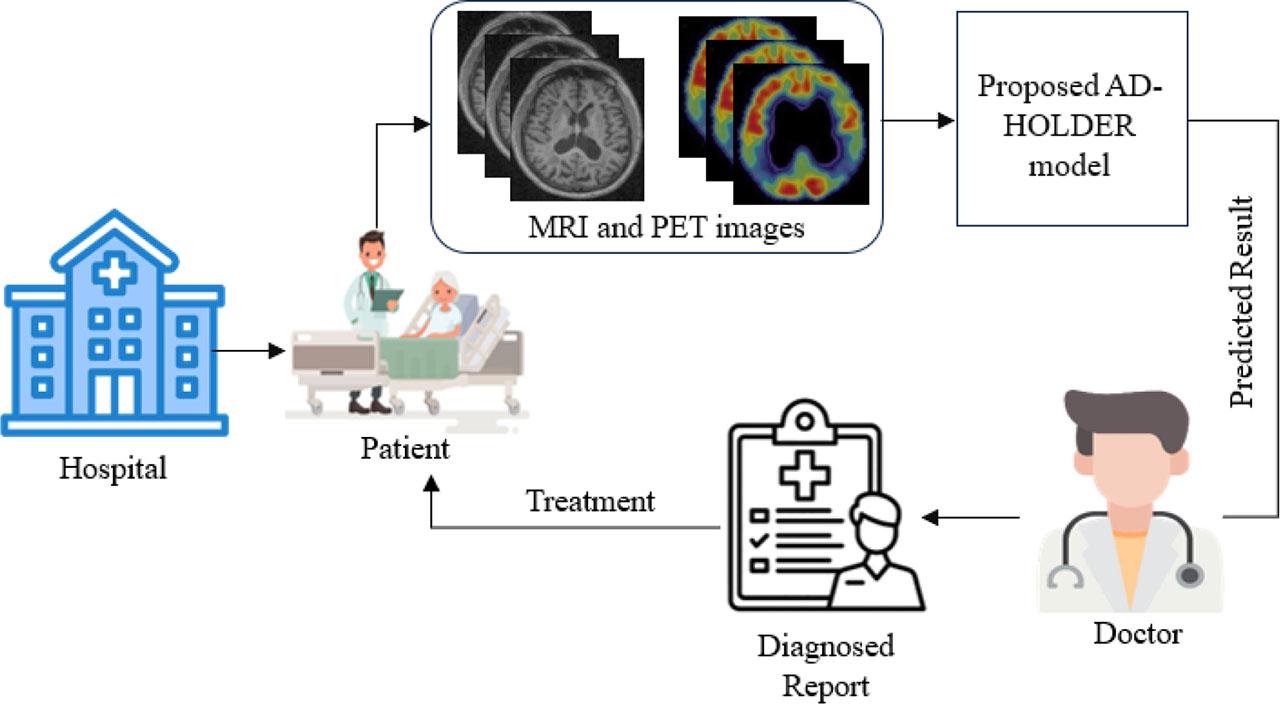
Real-time clinical setting of the proposed AD-HOLDER model.
In this study, the AD-HOLDER model is proposed for detecting AD using MRI and PET images. Initially, the input images are denoised using a deep image priors filter for noise removal and image enhancement. Next, features are extracted using the HOG method, which captures essential patterns and edges from the images. Then, the LGBM classifier is used to classify the normal and abnormal cases. Finally, the abnormal region is segmented using a GBS model to specifically detect the region for early diagnosis of AD. The proposed AD-HOLDER model achieves a classification ACC of 99.12 % through machine learning. The proposed LGBM achieves the highest ACC (99.12 %) and F1 (96.55 %), indicating its superior performance. The proposed AD-HOLDER model increases overall ACC by 1.55 %, 25.44 %, and 3.14 % for the GBA, XAI, and CAD system, respectively. The proposed AD-HOLDER model categorizes AD into normal and abnormal cases, without distinguishing multiple disease stages, making it insufficient for detailed progression analysis. Future work will extend the proposed model to classify AD into multiple stages, allowing a more nuanced understanding of the disease’s progression. Additionally, the model’s diagnostic ACC could be enhanced by incorporating multi-modal data, such as genetic and clinical information.