Fig. 1.
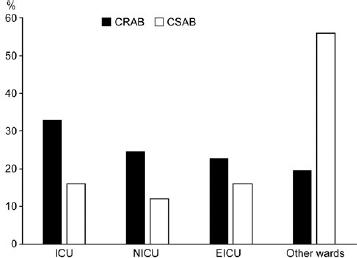
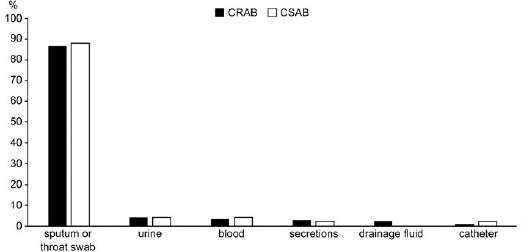
Fig. 2.
Primers used in this study_
| Primer | Primer sequence (5’–3’) | Target gene | References |
|---|---|---|---|
| intF | CCAAGCTCTCGGGTAACATC | intI1 | Wang et al. 2023 |
| P2R | CCCGAGGCATAGACTGTA | ||
| 5CS | GGCATCCAAGCAGCAAG | variable region | Wang et al. 2023 |
| 3CS | AAGCAGACTTGACCTGA | ||
| AACF | ATCTCATATCGTCGAGTGG | aac(6’)-Ib | Wang et al. 2023 |
| AACR | TGCGTGTTCGCTCGAATGC | ||
| AADAF | GCAGCGCAATGACATTCTTG | aadA1 | Wang et al. 2023 |
| AADAR | ATCCTTCGGCGCGATTTTG | ||
| CATF | TTACTCTGGCTACTATCAC | catB8 | Wang et al. 2023 |
| CATR | TGATGGCATAAGGCTCTAC | ||
| KPC-F | CGTCTAGTTCTGCTGTCTTG | blaKPC | Wang et al. 2023 |
| KPC-R | CTTGTCATCCTTGTTAGGCG | ||
| VIM-F | GATGGTGTTTGGTCGCATA | blaVIM | Wang et al.2023 |
| VIM-R | CGAATGCGCAGCACCAG | ||
| NDM-F | GGTTTGGCGATCTGGTTTTC | blaNDM | Wang et al. 2023 |
| NDM-R | CGGAATGGCTCATCACGATC | ||
| OXA-23-like-F | CCGTCGTTTACGACATTCA | blaOXA-23-like | Wang et al.2023 |
| OXA-23-like-R | AAAGAGCGCATTGCTTTGAT | ||
| IMP-F | GGAATAGAGTGGCTTAAYTCTC | blaIMP | Wang et al.2023 |
| IMP-R | GGTTTAAYAAAACAACCACC | ||
| bfmS-F | ATATATGCGGGGCTGGTAATTC | bfms | Cai et al. 2017 |
| bfmS-R | ATGCAGGTGCTTTTTTATTGGT | ||
| cusE-F | ATGCATGTTCTCTGGACTGATGTTGAC | cusE | Cai et al. 2017 |
| cusE-R | CGACTTGTACCGTGACCGTATCTTGATAAG | ||
| bap-F | CGTTTCCTGGGTCTGATGTATT | bap | Cai et al. 2017 |
| bap-R | GTTATTGAAGGCTTCTTTAGTG | ||
| abal-F | GTGGCTCAAGACAGAGAATC | abal | Wang and Ye 2021 |
| abal-R | TCAATCATCATTGGTGGACC | This study |
Virulence- and resistance-gene-carrying status of CRAB and CSAB_
| Genes | CRAB (n = 219) | CSAB (n = 50) | p |
|---|---|---|---|
| abal + | 204 | 19 | 0* |
| – | 15 | 31 | |
| bfmS + | 140 | 31 | 0.798 |
| – | 79 | 19 | |
| bap + | 193 | 5 | 0* |
| – | 26 | 45 | |
| cusE + | 170 | 24 | 0* |
| – | 49 | 26 | |
| blaOXA-23-like + | 218 | 0 | 0* |
| – | 1 | 50 |
Resistance rates of CRAB and CSAB isolates to commonly used antibiotics_
| Antibiotics | CRAB (n = 219) | CSAB (n = 50) | p | ||
|---|---|---|---|---|---|
| No. | Rate (%) | No. | Rate (%) | ||
| CAZ | 218 | 99.54 | 6 | 12.00 | 0.000* |
| CIP | 217 | 99.09 | 6 | 12.00 | 0.000* |
| TOB | 206 | 94.06 | 6 | 12.00 | 0.000* |
| DOX | 216 | 98.63 | 6 | 12.00 | 0.000* |
| SXT | 166 | 75.80 | 8 | 16.00 | 0.000* |
| LVX | 175 | 79.91 | 5 | 10.00 | 0.000* |
| AMK | 131 | 59.82 | 6 | 12.00 | 1.000 |
| FEP | 169 | 77.17 | 6 | 12.00 | 0.384 |
| MNO | 78 | 35.62 | 2 | 4.00 | 0.000* |
| TGC | 11 | 5.02 | 0 | 0.00 | 0.226 |
| COL | 3 | 1.37 | 0 | 0.00 | 1.000 |
Correlation between class 1 integron and antibiotic resistant rates in 219 CRAB isolates_
| Antibiotics | intI1(+) (n = 75) | intI1(−) (n = 144) | p | ||
|---|---|---|---|---|---|
| No. | Rate (%) | No. | Rate (%) | ||
| CAZ | 75 | 100 | 143 | 99.31 | 1.000 |
| CIP | 75 | 100 | 142 | 98.61 | 0.548 |
| TOB | 75 | 100 | 132 | 91.67 | 0.009* |
| DOX | 74 | 98.67 | 142 | 98.61 | 1.000 |
| SXT | 73 | 97.33 | 93 | 64.58 | 0.000* |
| LVX | 62 | 82.67 | 113 | 78.47 | 0.462 |
| AMK | 58 | 77.33 | 78 | 54.17 | 0.001* |
| FEP | 44 | 58.67 | 125 | 86.81 | 0.000* |
| MNO | 36 | 48 | 42 | 29.17 | 0.006* |
| TGC | 1 | 1.33 | 10 | 6.94 | 0.103 |
| COL | 1 | 1.33 | 2 | 1.39 | 1.000 |
Correlation between class 1 integron and virulence and resistance genes in 219 CRAB isolates_
| Genes | intI1(+) (n = 75) | intI1(–) (n = 144) | p |
|---|---|---|---|
| abal + | 75 | 129 | 0.003* |
| – | 0 | 15 | |
| bfmS + | 53 | 87 | 0.134 |
| – | 22 | 57 | |
| bap + | 72 | 121 | 0.009* |
| – | 3 | 23 | |
| cusE + | 53 | 116 | 0.088 |
| – | 22 | 28 | |
| blaOXA-23-like + | 75 | 143 | 1.000 |
| – | 0 | 1 |
Correlations between virulence genes and antibiotic resistance in 219 CRAB isolates_
| Antibiotics | abal | bfmS | cusE | bap | ||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| + (n = 204) | – (n = 15) | p | + (n = 140) | – (n = 79) | p | + (n = 170) | – (n = 49) | p | + (n = 193) | – (n = 26) | p | |
| MNO | 72 | 6 | 0.782 | 47 | 31 | 0.4 | 59 | 19 | 0.6 | 68 | 10 | 0.747 |
| DOX | 201 | 15 | 1 | 139 | 77 | 0.296 | 168 | 48 | 0.534 | 191 | 25 | 0.317 |
| SXT | 154 | 12 | 1 | 98 | 68 | 0.008* | 124 | 42 | 0.066 | 145 | 21 | 0.528 |
| CIP | 202 | 15 | 1 | 139 | 78 | 1 | 168 | 49 | 1 | 192 | 25 | 0.224 |
| LVX | 163 | 12 | 1 | 109 | 66 | 0.313 | 141 | 34 | 0.037* | 152 | 23 | 0.246 |
| TOB | 191 | 15 | 0.607 | 129 | 77 | 0.142 | 161 | 45 | 0.494 | 185 | 21 | 0.011* |
| AMK | 125 | 6 | 0.105 | 79 | 52 | 0.173 | 96 | 35 | 0.06 | 119 | 12 | 0.13 |
| TGC | 11 | 0 | 1 | 9 | 2 | 0.335 | 9 | 2 | 1 | 11 | 0 | 0.369 |
| CAZ | 203 | 15 | 1 | 139 | 79 | 1 | 169 | 49 | 1 | 192 | 26 | 1 |
| FEP | 156 | 13 | 0.74 | 112 | 57 | 0.184 | 141 | 28 | 0* | 148 | 21 | 0.641 |
| COL | 3 | 0 | 1 | 2 | 1 | 1 | 2 | 1 | 0.534 | 2 | 1 | 0.317 |
Results of multilocus sequence typing (MLST)_
| Oxford | ST | gltA | gyrB | gdhB | recA | cpn60 | gpi | rpoD |
| 3272 | 1 | 12 | 56 | 1 | 1 | 385 | 26 | |
| Pasteur | ST | cpn60 | fusA | gltA | pyrG | recA | rplB | rpoB |
| 2520 | 3 | 2 | 2 | 2 | 3 | 4 | 95 |