Fig. 1
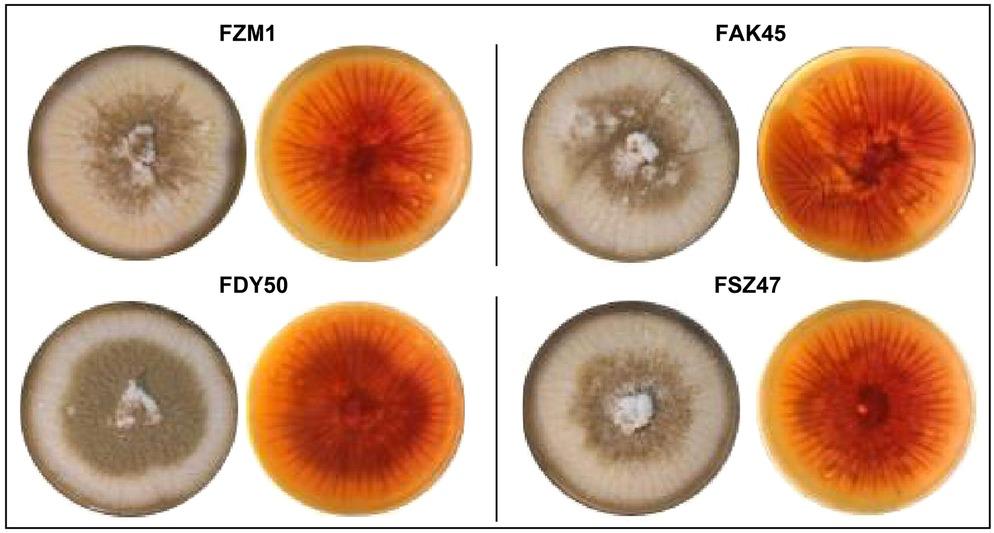
Fig. 2
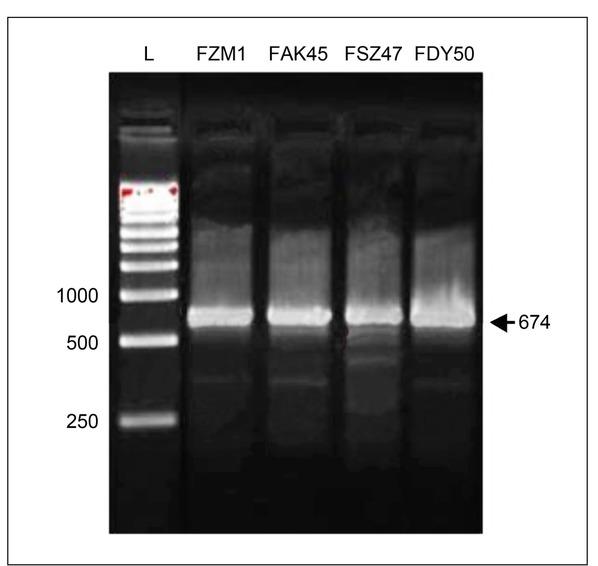
Fig. 3
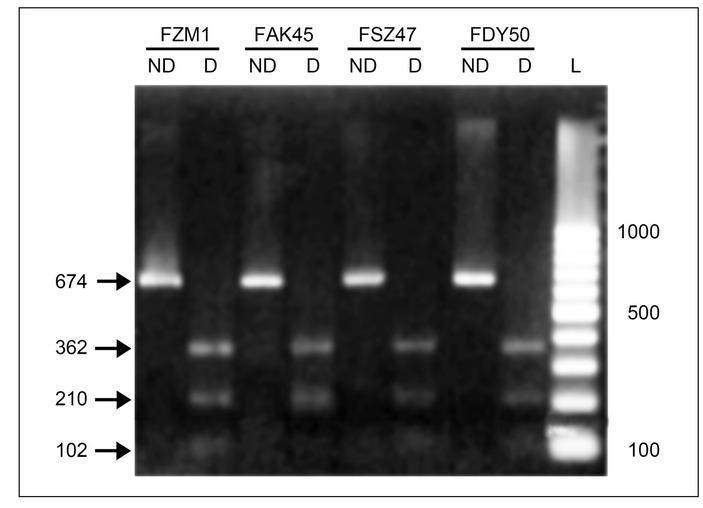
Fig. 4
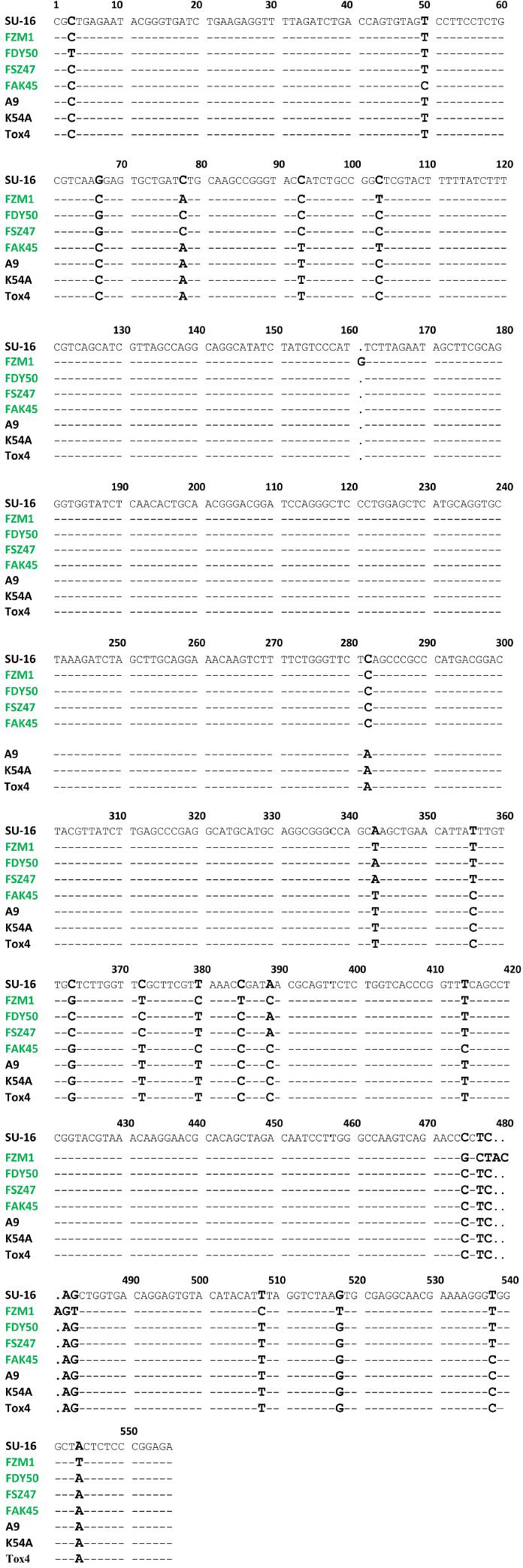
Fig. 5
Nucleotide variations in aflR-aflS(J) intergenic region sequences of Aspergillus flavus isolates_
| Nucleotide position | Nucleotide variation | Isolate |
|---|---|---|
| 103, 385, 478 | C with T (substitution) | FZM1: Aspergillus flavus strain FZM1 aflR-aflS(J) |
| 161 | Insertion of G | |
| 380, 477, 508 | T with C (substitution) | |
| 475 | C with G (inversion) | |
| 479–481 | Insertion of ACA | |
| 482 | A with G (substitution) | |
| 483 | G with T (substitution) | |
| 544 | A with T (inversion) | |
| 50 | T with C (substitution) | FAK45: Aspergillus flavus strain FAK45 aflR-aflS(J) |
| 103 | C with T (substitution) | |
| 3 | C with T (substitution) | FDY50: Aspergillus flavus strain aflR-aflS(J) |
| 5, 28 | G with A (substitution) |
Genbank accession numbers of the Aspergillus flavus isolates_
| Isolate | Accession number | Strain |
|---|---|---|
| FZM1 | OL944584.1 | A. flavus strain FZM1 aflR-aflJ intergenic region, partial sequence |
| FAK45 | OL944586.1 | A. flavus strain FAK45 aflR-aflJ intergenic region, partial sequence |
| FSZ47 | OL944587.1 | A. flavus strain FSZ47 aflR-aflJ intergenic region, partial sequence |
| FDY50 | OL944588.1 | A. flavus strain FDY50 aflR-aflJ intergenic region, partial sequence |
Aspergillus flavus isolates identity based on the BLAST NCBI data_
| Strain | Identity (%) |
|---|---|
| FZM1 | Aspergillus flavus strain A9 chromosome 3 (97.48%) |
| FAK45 | Aspergillus flavus strain A9 chromosome 3 (98.12%) |
| FSZ47 | Aspergillus flavus strain SU-16 chromosome 3 (99.37%) |
| FDY50 | Aspergillus flavus strain SU-16 chromosome 3 (99.67%) |
Genbank accession numbers of the Aspergillus flavus reference strains_
| Accession number | Strain |
|---|---|
| CP051037.1 | A. flavus strain A9 chromosome 3 |
| CP051085.1 | A. flavus strain K54A chromosome 3 |
| CP051045.1 | A. flavus strain Tox4 chromosome 3 |
| CP047251.1 | A. flavus strain SU-16 chromosome 3 |