Walnut (Juglans regia L.) is a tree species of the family Juglandaceae with high ecological and economic value. The vast geographical area spanning from Central Asia to the Himalayas is considered the gene center and evolutionary origin of this species (Pollegioni et al., 2015). The natural walnut forests in this region are of critical global bio-diversity importance, providing unique genetic resources that harbor the species’ adaptive potential in the face of climate change (Beer et al., 2008; Shah et al., 2025).
The Arslanbob walnut forests in southwestern Kyrgyzstan, covering approximately 600,000 hectares, form the world’s largest natural walnut forest ecosystem (Zhursunbek et al., 2019). Extending from 1200–2000 meters above sea level, these forests provide a critical habitat not only for J. regia but also for the region’s endemic flora and fauna. In terms of ecosystem services, these forests perform vital functions such as preventing soil erosion, regulating the water cycle, sequestering carbon, and stabilizing the local climate (Muratbek Kyzy and Gözlekçi, 2017; Zhaanbaev, 2023).
However, this unique ecosystem is facing severe threats. Uncontrolled livestock grazing, illegal logging, habitat fragmentation, and the effects of climate change are endangering the integrity and genetic diversity of the Arslanbob forests (Muratbek Kyzy and Gözlekçi, 2017). The increasing anthropogenic pressures in recent years are hindering natural regeneration and disrupting the population structure. Beer et al. (2008) reported, based on palynological data, that these forests were shaped by both natural and human influences, but that modern-day human pressure has reached unsustainable levels.
The loss of genetic diversity weakens ecosystem resilience by reducing the capacity of species to adapt to changing environmental conditions (Einollahi and Khadivi, 2024). Morphological diversity, as a phenotypic reflection of genetic diversity, is a critical indicator for evaluating the adaptive potential of populations. Nut and seed characteristics are important traits that are shaped by natural selection pressures and reflect local adaptation (Bernard et al., 2018). The systematic documentation of these traits provides fundamental data for developing in-situ conservation strategies and for identifying priority genetic materials for ex-situ conservation.
Historically, more than 220 genotypes were identified in the walnut forests of Kyrgyzstan by Prutenko and Shevchenko between 1953 and 1973, but very few of these (6 genotypes) could be put under protection (Venglovsky and Mamatzhanov, 2000). Previous studies are outdated, and there is a lack of recent data on the specific morphometric characteristics of this population under current climatic conditions. Given the habitat loss and degradation over the last 50 years, there is concern that a significant portion of these genetic resources may have been lost. This highlights the urgent need to assess and conserve the existing genetic diversity.
Global climate change scenarios predict an increase in temperature and changes in precipitation regimes in Central Asia. It has been reported that the annual average temperature has increased significantly in the region, with a 20% reduction in snow depth over the last 70 years (Fallah et al., 2024). These changes are expected to particularly affect mountain forest ecosystems with narrow ecological tolerance. As a result of climate change, the vertical upward movement of ecosystem zones (forests, grasslands, etc.) in mountainous regions has been observed (Saidaliyeva et al., 2024). Populations with high genetic diversity are at an advantage in adapting to these changes. The greatest adaptive response is expected in situations where populations are large, have high genetic variability, selection is strong, and there is an ecological opportunity for better-adapted genotypes to establish themselves (Alberto et al., 2013). Therefore, a comprehensive documentation of the existing diversity is of critical importance for future conservation and restoration efforts (Shah et al., 2025).
This study was conducted to evaluate the morphological diversity of natural walnut populations selected from the Arslanbob walnut forests in the Jalal-Abad region of Kyrgyzstan, to reveal intra-population variation, and to identify critical genotypes for conservation priorities. The aim is for the findings to: (i) reveal the current genetic diversity status of the Arslanbob forest ecosystem, (ii) provide basic data for in-situ conservation strategies, (iii) identify priority genetic materials for ex-situ conservation programs, and (iv) contribute to the development of adaptive management strategies to increase ecosystem resilience against climate change.
This study was conducted between July and September 2021 in the Arslanbob walnut forest ecosystem (40°44′N, 72°55′E) in the Jalal-Abad region of Kyrgyzstan (Fig. 1). The research was carried out in a valley within this unique ecosystem, which is described as the world’s largest natural walnut forest, covering an area of approximately 600,000 hectares. The region is located at an altitude of 1200–2000 meters and exhibits continental climate characteristics. According to data from the nearest meteorological station representative of the study region (Hardy et al., 2018), the area is characterized by a continental climate with marked altitudinal variation in precipitation and temperature. Annual precipitation ranges from approximately 535 mm at lower elevations (900 m) to 1090 mm at higher elevations (1750 m), with peaks in spring and autumn; August and September are the driest months. Mean temperatures in January and July are −3.1°C and 20.5°C, respectively, and winter minima can reach as low as −20°C (Hardy et al., 2018).

View of the Arslanbob walnut-forest region in the Jalal-Abad region, Kyrgyzstan.
It is reported that approximately 90% of the Arslanbob forests have been lost over the last 50 years (Kruzman, 2024). These forests have a global conservation priority due to their high biodiversity and endemic species (Orozumbekov et al., 2015). Furthermore, this region is considered the gene center for many nut species, offering critical genetic resources for future food security (Gaisberger et al., 2020).
A selection based on phenotypic superiority approach was applied to maximize the representation of genetic diversity, encompassing different elevation gradients and microhabitat characteristics. Since these trees represent wild-type populations with potential for domestication, the selection criteria prioritized individuals displaying traits suitable for cultivation and improvement, including: (1) tree vigor and health status (absence of major diseases and pests), (2) high nut yield potential based on visual assessment of nut load, (3) superior nut quality characteristics such as large nut size and thin shell, (4) ease of kernel extraction, (5) kernel plumpness and color quality, (6) representation of different elevation zones (1200–2000 m) to capture adaptive variation, and (7) spatial distribution across the study area to maximize genetic diversity. Trees showing signs of severe stress, disease, or mechanical damage were excluded from the selection. Twenty-eight mature walnut trees (J. regia L.) from the natural population were selected, maintaining a distance of at least 50 meters between them (Fig. 2). This distance criterion was determined to minimize the possibility of genetic relatedness and to maximize intra-population diversity.

Physical appearance of the studied walnut genotypes.
A total of eight quantitative morphological characters were evaluated in the nut samples from 28 genotypes selected from the Arslanbob walnut forests in the Jalal-Abad region of Kyrgyzstan. At harvest time, 30 nut samples collected from each genotype were separated from their green hulls, washed, and dried (Kırca et al., 2014). The measured nut characteristics were nut weight (g), kernel weight (g), kernel percentage (%), nut length (mm), nut width (mm), nut thickness (mm), shell thickness (mm), and nut shape index (length/width). Measurements were performed using a digital caliper (Asimeto 306-06-0, Hong Kong) and a precision analytical balance (Precisa XB 220 A, Switzerland; 0.01 g precision). The qualitative characteristics assessed were kernel color, kernel plumpness (well-filled, medium), shell roughness (smooth, medium, rough), shell color (yellow, light yellow, dark yellow, brown), and ease of kernel extraction (easy, medium, difficult) (Khadivi-Khub et al., 2015).
The morphological characterization was conducted according to the descriptor lists for walnut established by Bioversity International (IPGRI) and UPOV, taking into account both quantitative and qualitative traits to comprehensively reveal phenotypic diversity (Bioversity International, 2017; UPOV, 2017).
The statistical analyses of the obtained morphological data were performed using RStudio (Version 4.2.1) software (R Core Team, 2024). Prior to ANOVA, the normality of data was assessed using the Shapiro-Wilk test, and homogeneity of variances was tested using Levene's test (p > 0.05 indicating homogeneity). Analysis of variance (ANOVA) was applied to determine the differences among walnut genotypes, and Tukey’s multiple comparison test (p<0.05) was used for mean separation (Hothorn et al., 2008). Pearson correlation analysis was conducted to evaluate the relationships among pomological traits, and the correlation matrix was visualized (Wei and Simko, 2021).
Principal Component Analysis (PCA) was performed and a biplot graph was created to determine the distribution of genotypes and the relationships among traits (Lê et al., 2008; Kassambara and Mundt, 2020). Hierarchical Cluster Analysis (HCA) was conducted using Ward’s method to group the genotypes based on their similarities (Murtagh and Legendre, 2014). Percentages of qualitative traits were calculated and visualized as bar plots using the ‘ggplot2’ package. All graphs were created using the ‘ggplot2’ package (Wickham, 2016).
The conservation priorities of the genotypes were evaluated based on the criteria of morphological superiority, genetic representation, and adaptive potential. Priority genotypes for ex-situ conservation were identified based on both high performance and the capacity to represent population diversity at a maximum level. The conservation strategies were developed in accordance with the IUCN Red List of Ecosystems criteria (IUCN, 2017).
In this section, the morphological characterization of 28 walnut genotypes from the Arslanbob forest ecosystem is presented. Initially, the quantitative variation observed across genotypes for key pomological traits is described, followed by the application of multivariate statistical approaches (PCA and HCA) to reveal the underlying structure of genetic diversity. Subsequently, trait correlations are examined to identify potential trade-offs and synergies relevant to breeding and conservation strategies. Finally, qualitative traits are assessed, and the findings are discussed in relation to published literature on walnut germplasm, with particular emphasis on the implications for in-situ and ex-situ conservation under climate change.
Significant differences among 28 genotypes were detected for all quantitative pomological traits (p < 0.001; Tab. 1). Nut weight ranged from 4.8 g (G13) to 11.1 g (G27), with G27, G15, G23, and G28 forming the top group, while G13, G10, G1, and G9 were lowest. This nut weight range is lower than those reported for selected genotypes and cultivars in Turkey (12.35–20.88 g; Demir et al., 2019) and also below the mean values reported for Iranian walnut populations (mean 12.99 g; range 7.71–20.11 g; Arab et al., 2019), reflecting the wild and non-selected nature of the Arslanbob population. Kernel weight varied between 2.2 g (G13) and 5.1 g (G15), with G15 and G23 leading; these values are also lower than those reported for seedling-originated but selected genotypes (2.84–7.97 g; Einollahi and Khadivi, 2024) and Iranian populations (3.83–8.97 g; mean 6.5 g; Arab et al., 2019). Kernel percentage ranged from 33.6% (G27) to 57.7% (G24), with G24 and G19 notably high (>50%); this range overlaps substantially with previous reports for both seedling-originated genotypes (34.41–59.18%; Einollahi and Khadivi, 2024) and broader Persian walnut populations (38.31–67.36%; Arab et al., 2019), indicating that high kernel recovery can be achieved even in natural forest stands. Nut dimensions also spanned wide intervals (e.g., length 24.6–31.7 mm; width 22.8–30.5 mm; thickness 25.9–38.4 mm). Although these dimensions are generally smaller than those reported for selected or cultivated material (Demir et al., 2019; Einollahi and Khadivi, 2024), they fall within the lower limits of the ranges reported for natural and seedling-derived walnut populations (Arab et al., 2019). Shell thickness varied from 1.9 mm (G10) to 4.0 mm (G27); this range is notably higher than shell thickness values reported for Iranian populations (0.98–2.63 mm; mean 1.70 mm; Arab et al., 2019) and seedling-originated selected genotypes (0.88–2.60 mm; Einollahi and Khadivi, 2024), suggesting a population-level tendency toward thicker shells. Shape index ranged from 0.84 (G19) to 1.06 (G23).
Analysis of variance for some pomological traits of selected walnut genotypes (n = 30 per genotype).
| Genotypes | Nut weight (g) | Kernel weight (g) | Kernel ratio (%) | Nut length (mm) | Nut width (mm) | Nut thickness (mm) | Shell thickness (mm) | Shape index |
|---|---|---|---|---|---|---|---|---|
| G 1 | 7.13±0.73 g | 3.30±0.49 d-i | 46.18±3.50 d-g | 27.52±1.02 f-i | 27.08±0.99 f-i | 27.52±1.27 no | 2.23±0.27 k-n | 1.01±0.02 b-e |
| G 2 | 8.66±1.01 c-e | 3.25±0.55 e-i | 37.59±4.86 j-l | 28.30±1.31 d-f | 27.96±1.23 c-g | 33.01±1.46 c-e | 2.71±0.49 f-i | 0.93±0.03 h-k |
| G 3 | 8.77±0.71 c | 3.99±0.55 b-d | 45.35±3.07 e-h | 29.40±1.33 c-e | 30.34±1.02 a | 31.31±1.47 e-i | 2.36±0.32 h-l | 0.96±0.04 f-i |
| G 4 | 7.31±0.52 fg | 3.44±0.39 d-h | 46.98±3.40 d-g | 27.86±0.72 e-h | 27.30±0.90 f-i | 27.81±0.94 l-n | 2.33±0.45 h-l | 1.01±0.02 b-d |
| G 5 | 8.36±0.69 c-f | 3.61±0.44 c-g | 43.08±3.41 e-j | 27.94±0.92 e-g | 29.37±1.30 a-c | 29.48±1.95 i-m | 2.35±0.39 h-l | 0.95±0.04 f-i |
| G 6 | 9.98±0.92 b | 3.99±0.63 b-d | 39.89±4.28 h-k | 28.26±1.37 d-f | 28.24±1.19 c-g | 33.52±1.54 b-d | 2.95±0.34 ef | 0.92±0.03 i-k |
| G 7 | 8.64±0.86 c-e | 3.54±0.50 c-g | 40.83±2.78 g-j | 27.12±0.91 f-j | 27.48±0.94 e-h | 32.76±1.01 c-f | 2.47±0.22 g-k | 0.90±0.02 j-l |
| G 8 | 8.15±2.01 c-g | 3.45±1.17 d-h | 41.37±5.47 e-j | 29.91±1.31 b-d | 29.07±1.15 a-e | 30.13±1.74 g-k | 2.26±0.43 i-n | 1.01±0.04 b-e |
| G 9 | 7.29±1.08 g | 3.79±0.70 c-f | 51.80±4.60 b-d | 28.38±1.29 d-f | 27.52±1.23 e-g | 28.28±1.50 k-n | 2.01±0.23 l-n | 1.02±0.04 bc |
| G 10 | 6.09±0.58 h | 2.79±0.41 hi | 45.66±4.64 e-g | 25.43±1.13 kl | 25.12±1.20 jk | 27.93±1.01 mn | 1.87±0.14 n | 0.96±0.03 f-h |
| G 11 | 8.14±1.38 c-g | 3.58±0.88 c-g | 43.94±7.53 e-j | 28.73±1.55 c-f | 27.74±1.50 d-g | 30.09±1.81 h-k | 2.20±0.37 k-n | 0.99±0.02 b-f |
| G 12 | 7.67±0.93 d-g | 2.92±0.43 g-i | 38.61±6.87 i-l | 28.35±1.15 d-f | 27.63±1.05 e-g | 29.51±1.71 i-m | 2.24±0.28 k-n | 0.99±0.05 b-f |
| G 13 | 4.77±1.11 i | 2.17±0.37 i | 47.6±13.35 de | 24.62±1.21 l | 22.80±1.26 l | 25.90±1.39 o | 1.90±0.31 mn | 1.01±0.04 bc |
| G 14 | 8.12±0.48 c-g | 2.79±0.33 hi | 34.26±2.74 kl | 27.84±1.36 f-h | 25.99±0.51 h-j | 29.08±1.07 j-n | 3.56±0.13 bc | 1.01±0.04 bc |
| G 15 | 10.74±0.63 ab | 5.13±0.51 a | 47.66±3.16 c-f | 29.61±2.43 cd | 29.95±2.20 ab | 38.43±1.77 a | 2.98±0.46 ef | 0.87±0.03 lm |
| G 16 | 7.78±0.92 c-g | 3.73±0.62 c-f | 47.68±2.84 de | 26.16±1.29 i-k | 26.84±1.27 g-i | 33.36±1.93 b-d | 2.43±0.32 h-k | 0.87±0.03 lm |
| G 17 | 8.73±0.59 cd | 3.93±0.44 b-f | 44.94±3.45 e-i | 28.31±1.27 d-f | 28.60±1.43 b-f | 34.78±1.71 bc | 2.23±0.41 j-n | 0.90±0.02 kl |
| G 18 | 7.10±0.64 gh | 2.99±0.51 g-i | 41.91±4.18 e-j | 28.63±1.61 c-f | 28.03±0.83 c-g | 29.85±1.31 i-l | 2.30±0.41 h-m | 0.99±0.04 b-g |
| G 19 | 7.50±0.86 e-g | 4.25±0.69 bc | 56.39±4.70 ab | 26.29±1.28 h-k | 27.80±1.18 c-g | 35.18±1.83 b | 2.85±0.36 e-g | 0.84±0.03 m |
| G 20 | 8.63±0.70 cd | 3.53±0.45 d-g | 40.73±2.23 g-j | 27.30±0.80 f-j | 26.05±1.03 h-j | 30.78±1.18 f-j | 3.24±0.36 c-e | 0.97±0.02 e-h |
| G 21 | 8.16±0.93 c-g | 3.23±0.52 f-i | 39.53±4.03 h-k | 27.62±1.09 f-i | 27.17±0.92 f-i | 32.08±1.22 d-h | 3.51±0.24 b-d | 0.94±0.04 h-k |
| G 22 | 7.98±0.87 c-g | 3.29±0.50 f-h | 41.12±3.04 g-j | 29.68±2.11 cd | 29.83±2.40 ab | 33.22±2.01 cd | 3.14±0.46 de | 0.94±0.05 h-j |
| G 23 | 10.5±0.98 ab | 4.66±0.70 ab | 44.39±5.00 e-ı | 31.72±1.30 a | 30.41±1.31 a | 29.29±2.46 j-n | 3.76±0.44 ab | 1.06±0.05 a |
| G 24 | 7.31±0.73 g | 4.22±0.56 bc | 57.69±3.34 a | 25.99±1.20 jk | 27.67±1.34 e-g | 32.04±1.50 d-g | 2.16±0.30 k-n | 0.87±0.02 lm |
| G 25 | 7.64±0.70 d-g | 2.61±0.48 i | 34.08±4.66 kl | 27.05±1.05 f-j | 24.08±1.05 kl | 28.46±1.22 k-n | 2.74±0.43 f-h | 1.03±0.03 ab |
| G 26 | 7.45±0.58 fg | 3.98±0.62 b-e | 53.75±8.62 a-c | 26.49±1.27 g-k | 25.78±0.79 ij | 30.63±1.13 g-j | 2.03±0.16 k-n | 0.94±0.05 g-k |
| G 27 | 11.07±1.09 a | 3.73±0.68 c-f | 33.64±4.91 l | 30.05±1.27 bc | 29.18±1.35 a-d | 32.97±1.34 c-e | 3.95±0.49 a | 0.97±0.04 d-h |
| G 28 | 10.03±1.18 b | 4.22±0.83 bc | 41.82±5.90 f-j | 31.18±1.80 ab | 30.54±1.60 a | 32.95±2.80 c-e | 2.65±0.46 f-j | 0.98±0.05 c-g |
| p-value | *** | *** | *** | *** | *** | *** | *** | *** |
Averages within the same column followed by the same letter do not differ statistically significantly (p>0.05).
The *** symbol indicates that the differences between genotypes for the relevant trait are significant at the p<0.001 level.
Overall, the wide trait ranges observed in the Arslanbob population are consistent with the high genetic diversity reported for Persian walnut populations derived from seed propagation and natural regeneration (Arab et al., 2019; Khadivi Khub, 2014; Einollahi and Khadivi, 2024). From a breeding and conservation perspective, the coexistence of high kernel percentage and moderate shell thickness in several genotypes (e.g., G19, G24, G26) indicates immediate potential for quality-focused selection, whereas large-fruited, high-weight genotypes (G15, G23, G27, G28) represent promising material for yield-oriented improvement. The comparatively lower nut and kernel weights observed in Arslanbob relative to cultivated material can be attributed to the absence of artificial selection and to ecological influences, as ecological effects on walnut fruit traits have been clearly demonstrated in cultivar-based studies conducted under different environments (Bayazıt et al., 2024; Bayazıt et al., 2025). Given the documented cultivar × environment interactions for nut and kernel traits and the increasing climatic variability affecting walnut-growing regions, the maintenance of such within-population morphological diversity in natural walnut forests is critical for long-term conservation and adaptive breeding strategies.
The first two principal components explained 74.4% of total variance (PC1=42.9%; PC2=31.5%; Fig. 3). PC1 loaded strongly on nut size and mass (nut and kernel weight, width, thickness), positioning G15, G17, G28, and G22 on the positive side; small/light genotypes (G13, G10) loaded negatively. PC2 captured the kernel yield vs. shell/shape axis, with nut length, shell thickness, and shape index positive, and kernel percentage negative. Thus, G23–G27 tended toward longer nuts and/or thicker shells, while G19–G24 showed higher kernel percentages.

Principal Component Analysis (PCA) Biplot Graph of Selected Walnut Genotypes Based on Studied Pomo-logical Traits (n = 30 per genotype)
This two-axis structure (size–mass vs. kernel yield–shell) is consistent with findings from other walnut germplasm studies, in which PC1 usually captures overall nut or seed size, whereas PC2 separates kernel proportion from shell thickness or hardness (Khadivi Khub et al., 2015; Hakimi et al., 2024; Sharma et al., 2025). Functionally, PC2 underscores a commercial and mechanical trade-off: very thin shells risk transport damage; very thick shells reduce cracking efficiency and net yield. Identifying genotypes with favorable positions on both axes is valuable for combining market-preferred kernel yield with robust shell integrity in breeding pipelines.
HCA grouped genotypes into three main clusters (Fig. 4). Cluster I aggregated high kernel-percentage types (G19, G24, G26) alongside small-sized genotypes (G10, G13). Cluster II represented a heterogeneous, central group with moderate values. Cluster III brought together large and heavy genotypes (G15, G23, G27, G28) with thicker shells.
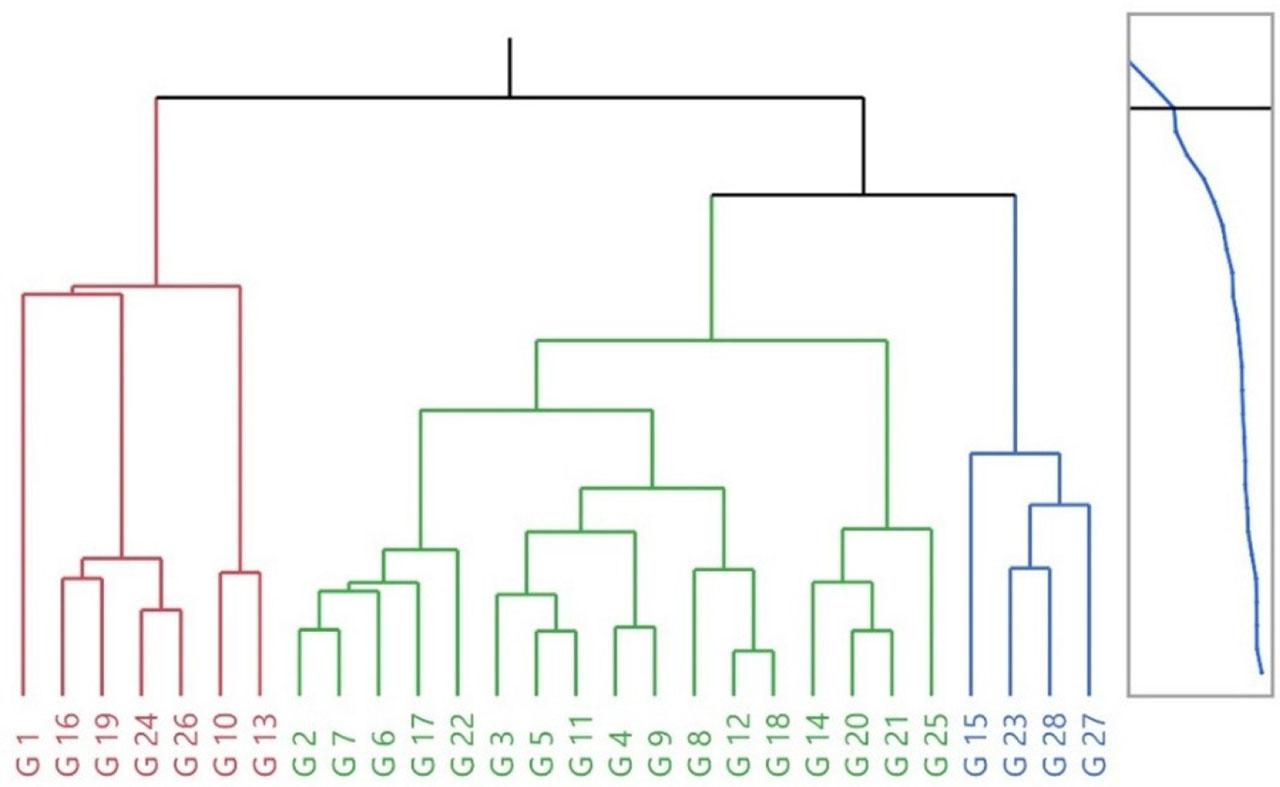
Dendrogram of hierarchical cluster analysis of selected walnut genotypes based on studied pomological traits (n = 30 per genotype).
Consistency between HCA groupings and PCA positions suggests a robust underlying structure. For strategy: Cluster I genotypes are prime for quality traits (kernel%), but require attention to shell robustness; Cluster III suits yield/size targets and can be crossed with Cluster I to combine desirable attributes. Considering reported genetic erosion risks due to preferential harvest of thin-shelled/high-kernel types in Kyrgyz forests (Orozumbekov et al., 2015; Cantarello et al., 2014), in-situ protection of Cluster I is a near-term conservation priority, complemented by ex-situ safety-duplication.
Correlation analysis (Fig. 5) showed that nut weight was positively correlated with kernel weight (r = 0.62, p < 0.001), nut length (r = 0.76, p < 0.001), nut width (r = 0.73, p < 0.001), nut thickness (r = 0.62, p < 0.001), and shell thickness (r = 0.66, p < 0.001). Kernel weight was positively associated with kernel ratio (r = 0.42, p < 0.05) and showed strong positive correlations with nut width (r = 0.65, p < 0.001) and nut thickness (r = 0.58, p < 0.001), whereas its correlations with nut length (r = 0.32) and shell thickness (r = 0.22) were not significant. Kernel ratio was negatively correlated with shell thickness (r = −0.53, p < 0.01) and tended to decrease with increasing nut length (r = −0.37). Nut length and nut width were strongly correlated (r = 0.83, p < 0.001). In addition, nut length was positively correlated with shell thickness (r = 0.44, p < 0.05), and nut width was moderately correlated with nut thickness (r = 0.54, p < 0.01).

Correlation matrix of the studied pomological traits of walnut genotypes (n = 30 per genotype). *: 0.05, **:0.01, ***: 0.001
Consistent with previous walnut germplasm studies reporting similar linkages between nut size, kernel traits and shell characteristics (Einollahi and Khadivi, 2024; Khadivi et al., 2019; Arab et al., 2019; Karadeniz, 2011; Karadeniz and Çorumlu, 2014; Demir et al., 2019; Karlıdağ et al., 2019; Kırca, 2025; Amiri et al., 2010), these relationships imply that (i) selection for heavier nuts tends to co-select larger kernels and thicker shells; (ii) targeting higher kernel percentage may require concurrent constraints on shell thickening and nut elongation; and (iii) dimensional traits move together, with potential consequences for cracking efficiency and whole-kernel recovery. Under climate warming and potential water stress (Fallah et al., 2024; Saidaliyeva et al., 2024), size-related traits may shift, so maintaining these covariance structures is likely important for preserving population-level resilience (IUCN, 2017).
Qualitative assessments (Fig. 6) indicated 53.6% yellow and 46.4% light-yellow kernel color; 100% ‘plump’ kernels; 57.14% smooth shells (vs. 35.71% medium, 7.14% rough); shell color predominantly yellow (50%) followed by light yellow (25%), dark yellow (21.43%), brown (3.57%); and 71.43% easy kernel extraction (vs. 17.86% medium, 10.71% difficult).
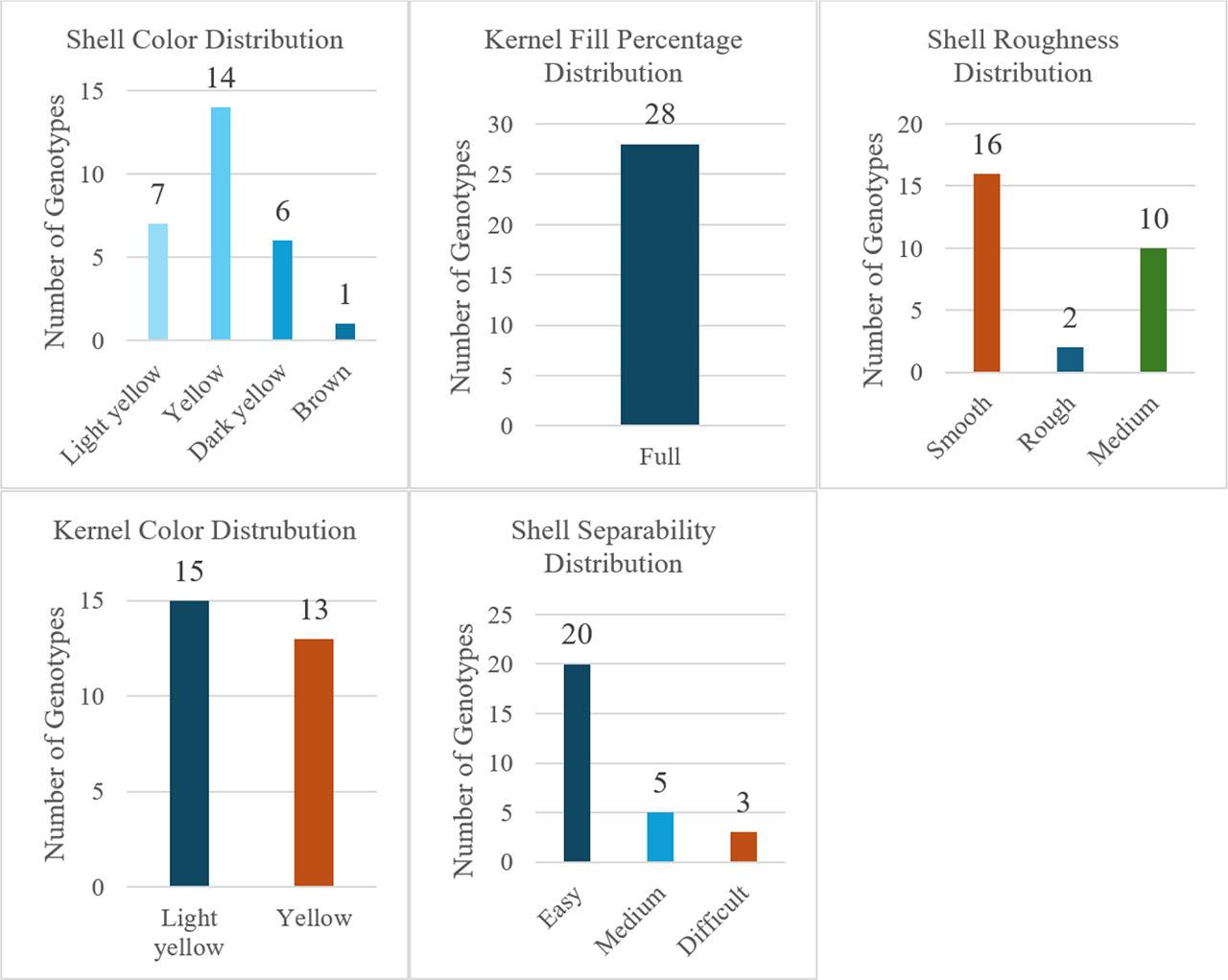
Qualitative pomological traits of the studied walnut genotypes.
Light-colored kernels, smooth shells, and easy extraction are premium market traits (Vahdati et al., 2020; Koç et al., 2019). Their relatively high frequencies here may reflect both natural and human-mediated selection patterns observed in Kyrgyz walnut-fruit forests (Orozumbekov et al., 2015). Genotypes G15, G16, G17, G18, G19, G22, G23, and G26 combined superior qualitative profiles with quantitative merit, making them strong candidates for both direct evaluation (elite type testing) and parental use in crossing schemes. Conserving these types in-situ and ex-situ is recommended to secure trait combinations valuable under changing climates and markets. While morphological traits can be influenced by environmental factors, the integration of molecular markers—such as RAPDs—is essential for environment-independent genetic identification and phylogenetic relationships among genotypes (Erdoğan and Aygün, 2016). Future molecular studies will be critical for formal registration and protection of these wild genetic resources.
In this study, a comprehensive pomological characterization was performed on 28 genotypes selected from the Arslanbob walnut forest in Kyrgyzstan. High variation was found for quantitative traits, particularly for nut weight (4.8–11.1 g), kernel percentage (33.6–57.7%), and shell thickness (1.9–4.0 mm). Notably, genotypes G15, G16, G17, G18, G19, G22, G23, and G26 showed superior traits with a high kernel percentage (>45%) and suitable shell thickness (<3.0 mm). PCA analysis, explaining 74.4% of the total variation, revealed that nut size/weight and kernel yield/shell thickness were the main sources of variation.
The rich genetic diversity found in the Arslanbob walnut forests requires urgent conservation measures in the face of threats from climate change and habitat loss. The identified superior genotypes can be evaluated as direct cultivar candidates or used in future breeding programs. While this study provides an important foundation for the conservation and sustainable use of Kyrgyzstan’s walnut genetic resources, it is recommended that it be supported by molecular characterization and environmental adaptation studies.