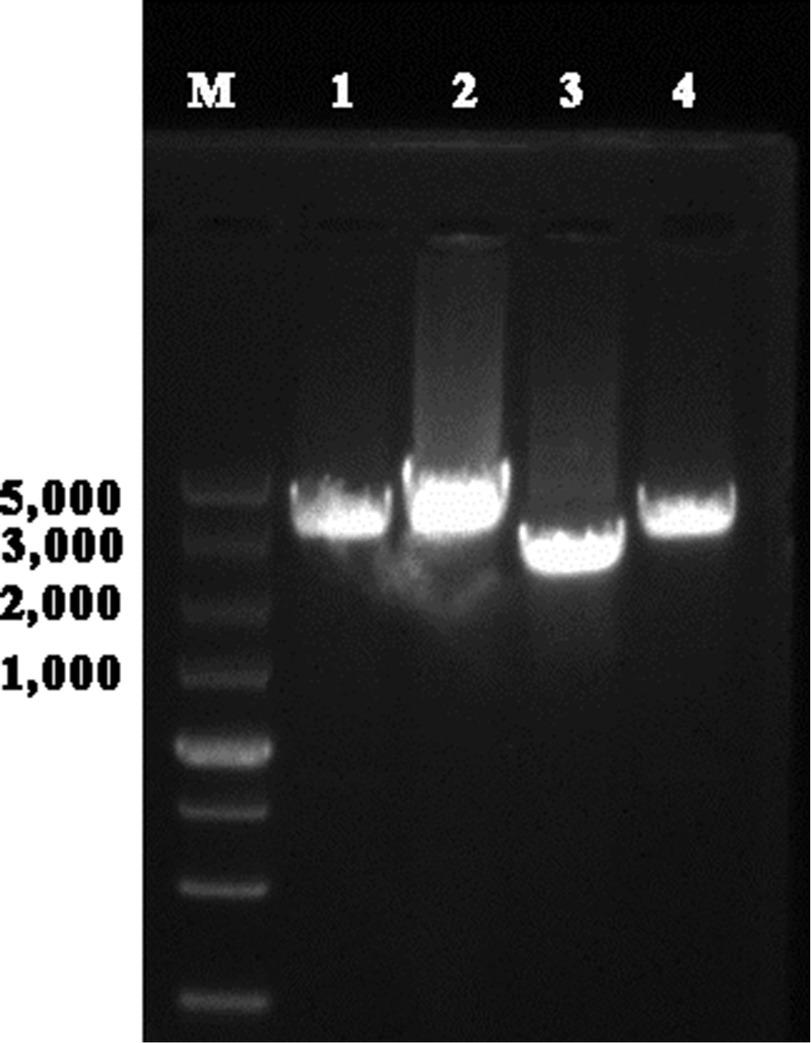
Figure 1
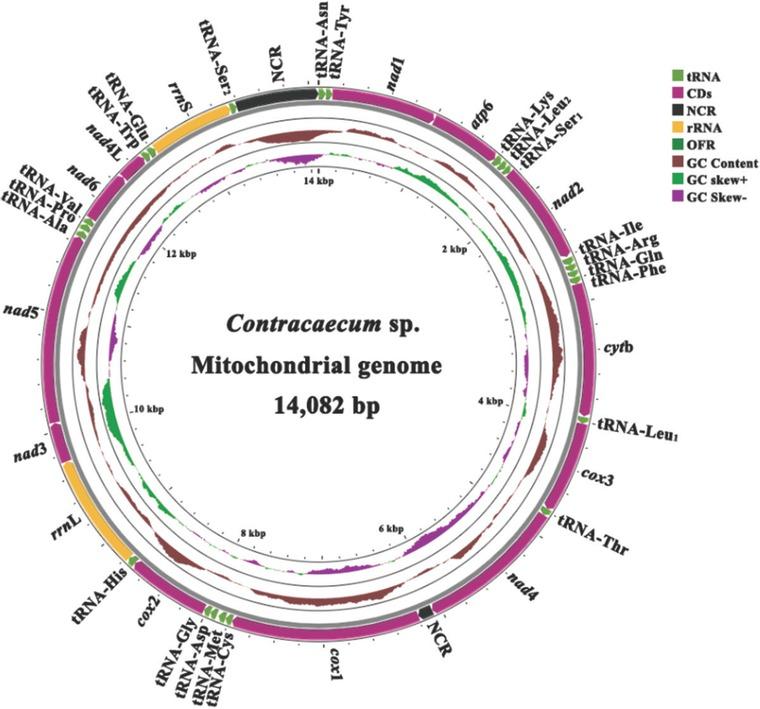
Figure 2

Figure 3

Figure S1
Nucleotide composition and skews of Contracaecum sp_ mitochondrial genome_
| Nucleotide frequency (%) | |||||||
|---|---|---|---|---|---|---|---|
| Gene | A | G | T | C | A + T (%) | AT-skew | GC-skew |
| atp6 | 22.0 | 22.0 | 49.1 | 6.9 | 71.1 | -0.380 | 0.526 |
| cox1 | 19.5 | 21.8 | 47.2 | 11.5 | 66.7 | -0.416 | 0.307 |
| cox2 | 21.2 | 22.1 | 46.5 | 10.1 | 67.7 | -0.373 | 0.372 |
| cox3 | 18.9 | 20.9 | 49.8 | 10.4 | 68.7 | -0.449 | 0.333 |
| cytb | 19.7 | 22.0 | 47.6 | 10.7 | 67.3 | -0.415 | 0.343 |
| nad1 | 19.5 | 20.5 | 50.5 | 9.5 | 70.0 | -0.444 | 0.364 |
| nad2 | 20.7 | 18.2 | 54.6 | 6.5 | 75.3 | -0.451 | 0.474 |
| nad3 | 20.0 | 21.4 | 54.4 | 4.2 | 74.4 | -0.464 | 0.674 |
| nad4 | 21.4 | 17.0 | 50.9 | 10.7 | 72.3 | -0.408 | 0.226 |
| nad4L | 22.9 | 17.3 | 55.0 | 4.8 | 77.9 | -0.411 | 0.569 |
| nad5 | 21.2 | 18.8 | 51.9 | 8.1 | 73.1 | -0.420 | 0.398 |
| nad6 | 20.7 | 13.3 | 58.2 | 7.8 | 78.9 | -0.475 | 0.261 |
| rrnS | 30.2 | 19.7 | 40.4 | 9.7 | 70.6 | -0.143 | 0.340 |
| rrnL | 27.3 | 17.5 | 48.3 | 6.9 | 75.6 | -0.277 | 0.436 |
| 22 tRNA | 31.5 | 18.7 | 40.8 | 9.0 | 72.3 | -0.129 | 0.352 |
| NCR | 37.4 | 10.3 | 46.7 | 5.6 | 84.1 | -0.111 | 0.290 |
| Total | 23.5 | 19.0 | 48.7 | 8.9 | 72.2 | -0.350 | 0.364 |
Amino acid frequency of Contracaecum sp_ mitochondrial PCGs_
| Amino acid | Codon | Number | RSCU (%) | Amino acid | Codon | Number | RSCU (%) |
|---|---|---|---|---|---|---|---|
| Phe | TTT | 480 | 1.92 | Tyr | TAT | 154 | 1.84 |
| Phe | TTC | 19 | 0.08 | Tyr | TAC | 13 | 0.16 |
| Leu | TTA | 199 | 2.3 | Stop | TAA | 5 | 1.43 |
| Leu | TTG | 216 | 2.5 | Stop | TAG | 2 | 0.57 |
| Leu | CTT | 76 | 0.88 | His | CAT | 54 | 1.86 |
| Leu | CTC | 2 | 0.02 | His | CAC | 4 | 0.14 |
| Leu | CTA | 10 | 0.12 | Gln | CAA | 20 | 0.98 |
| Leu | CTG | 16 | 0.18 | Gln | CAG | 21 | 1.02 |
| Ile | ATT | 214 | 1.92 | Asn | AAT | 100 | 1.79 |
| Ile | ATC | 9 | 0.08 | Asn | AAC | 12 | 0.21 |
| Met | ATA | 76 | 0.86 | Lys | AAA | 35 | 0.71 |
| Met | ATG | 101 | 1.14 | Lys | AAG | 63 | 1.29 |
| Val | GTT | 219 | 2.61 | Asp | GAT | 62 | 1.65 |
| Val | GTC | 13 | 0.16 | Asp | GAC | 13 | 0.35 |
| Val | GTA | 49 | 0.59 | Glu | GAA | 32 | 0.84 |
| Val | GTG | 54 | 0.64 | Glu | GAG | 44 | 1.16 |
| Ser | TCT | 139 | 3.08 | Cys | TGT | 53 | 1.96 |
| Ser | TCC | 6 | 0.13 | Cys | TGC | 1 | 0.04 |
| Ser | TCA | 14 | 0.31 | Trp | TGA | 21 | 0.57 |
| Ser | TCG | 5 | 0.11 | Trp | TGG | 53 | 1.43 |
| Pro | CCT | 66 | 3.11 | Arg | CGT | 33 | 3.88 |
| Pro | CCC | 7 | 0.33 | Arg | CGC | 1 | 0.12 |
| Pro | CCA | 9 | 0.42 | Arg | CGA | 0 | 0 |
| Pro | CCG | 3 | 0.14 | Arg | CGG | 0 | 0 |
| Thr | ACT | 89 | 3.24 | Ser | AGT | 121 | 2.68 |
| Thr | ACC | 6 | 0.22 | Ser | AGC | 2 | 0.04 |
| Thr | ACA | 9 | 0.33 | Ser | AGA | 36 | 0.8 |
| Thr | ACG | 6 | 0.22 | Ser | AGG | 38 | 0.84 |
| Ala | GCT | 72 | 2.5 | Gly | GGT | 112 | 2.22 |
| Ala | GCC | 24 | 0.83 | Gly | GGC | 21 | 0.42 |
| Ala | GCA | 11 | 0.38 | Gly | GGA | 23 | 0.46 |
| Ala | GCG | 8 | 0.28 | Gly | GGG | 46 | 0.91 |
Primers used for assembly validation_
| Name | Sequence (5'-3') | Size (bp) |
|---|---|---|
| yeluF1 | AGTTGTTGAAGAAGGAGCAGTT | |
| yeluR1 | CTAAACATTGACCTAACCACCT | 3,564 bp |
| yeluF2 | AGGTGGTTAGGTCAATGTTTAG | |
| yeluR2 | ACAGAGTAAACATCAGGGAAAT | 3,900 bp |
| yeluF3 | TTGGATTTCCCTGATGTTTACT | |
| yeluR3 | CAAACTAAACATACTGCCAACA | 2,816 bp |
| yeluF4 | TTGGTCAACAAGATGGTCGTAA | |
| yeluR4 | AACTGCTCCTTCTTCAACAACT | 3,757 bp |
Organization of the complete mt genome of Contracaecum sp_ from Beijing, China_
| Gene/region | Strand | Positions | Size (bp) | Number of aaa | Ini/Ter codons | Anticodons | In |
|---|---|---|---|---|---|---|---|
| tRNA-Asn (N) | H | 1–60 | 60 | GTT | 0 | ||
| tRNA-Tyr (Y) | H | 61–116 | 56 | GTA | 0 | ||
| nad1 | H | 117–989 | 873 | 290 | TTG/TAG | 0 | |
| atp6 | H | 993–1,591 | 599 | 199 | ATT/TA | +3 | |
| tRNA-Lys (K) | H | 1,592–1,653 | 62 | TTT | 0 | ||
| tRNA-Leu2 (L2) | H | 1,654–1,708 | 55 | TAA | 0 | ||
| tRNA-Ser1 (S1) | H | 1,709–1,759 | 51 | TCT | 0 | ||
| nad2 | H | 1,760–2,605 | 846 | 281 | TTG/TAA | 0 | |
| tRNA-Ile (I) | H | 2,619–2,678 | 60 | GAT | +13 | ||
| tRNA-Arg (R) | H | 2,679–2,732 | 54 | GCG | 0 | ||
| tRNA-Gln (Q) | H | 2,733–2,787 | 55 | TTG | 0 | ||
| tRNA-Phe (F) | H | 2,788–2,846 | 59 | GAA | 0 | ||
| Cytb | H | 2,847–3,953 | 1,107 | 368 | TTG/TAA | 0 | |
| tRNA-Leu1 (L1) | H | 3,961–4,017 | 57 | TAG | +7 | ||
| cox3 | H | 4,018–4,782 | 766 | 255 | TTG/T | 0 | |
| tRNA-Thr (T) | H | 4,783–4,843 | 60 | TGT | 0 | ||
| nad4 | H | 4,844–6,073 | 1,230 | 409 | TTG/TAA | 0 | |
| Intergenic region | H | 6,074–6,195 | 122 | 0 | |||
| cox1 | H | 6,196–7,771 | 1,576 | 525 | TTG/T | 0 | |
| tRNA-Cys (C) | H | 7,772–7,829 | 58 | GCA | 0 | ||
| tRNA-Met (M) | H | 7,831–7,890 | 60 | CAT | +1 | ||
| tRNA-Asp (D) | H | 7,907–7,963 | 57 | GTC | +16 | ||
| tRNA-Gly (G) | H | 7,965–8,021 | 57 | TCC | +1 | ||
| cox2 | H | 8,022–8,713 | 692 | 230 | TTG/TA | 0 | |
| tRNA-His (H) | H | 8,714–8,778 | 65 | GTG | 0 | ||
| rrnL | H | 8,779–9,737 | 959 | 0 | |||
| nad3 | H | 9,738–10,073 | 336 | 111 | TTG/TAG | 0 | |
| nad5 | H | 10,077–11,659 | 1,583 | 527 | ATT/TA | +3 | |
| tRNA-Ala (A) | H | 11,660–11,716 | 57 | TGC | 0 | ||
| tRNA-Pro (P) | H | 11,724–11,780 | 57 | TGG | +7 | ||
| tRNA-Val (V) | H | 11,781–11,837 | 57 | TAC | 0 | ||
| nad6 | H | 11,838–12,272 | 435 | 144 | TTG/TAA | 0 | |
| nad4L | H | 12,275–12,505 | 231 | 76 | ATT/TAA | +2 | |
| tRNA-Trp (W) | H | 12,506–12,563 | 58 | TCA | 0 | ||
| tRNA-Glu (E) | H | 12,565–12,624 | 60 | TTC | +1 | ||
| rrnS | H | 12,625–13,335 | 711 | 0 | |||
| tRNA-Ser2 (S2) | H | 13,336–13,391 | 56 | TGA | 0 | ||
| Noncoding region | H | 13,392–14,082 | 691 | 0 |
Mitochondrial genome sequences of nematodes of superfamily Ascaridoidea and Heterakoidea were sequenced completely before the present study and used for phylogenetic analysis_
| Superfamily | Family | Species | Size (bp) | GenBank accession No. |
|---|---|---|---|---|
| Ascaridoidea | Anisakidae | Anisakis pegreffii | 14,002 | NC_034329 |
| Anisakis simplex | 13,899 | MK820679 | ||
| Contracaecum ogmorhini Canada | 14,010 | KU558727 | ||
| Contracaecum ogmorhini Australia | 14,019 | KU558725 | ||
| Contracaecum ogmorhini South Africa | 14,012 | KU558726 | ||
| Contracaecum osculatum | 13,823 | NC_024037 | ||
| Contracaecum rudolphii | 14,022 | NC_014870 | ||
| Pseudoterranova decipiens | 13,962 | NC_031645 | ||
| Pseudoterranova decipiens s.l. | 13,965 | KU558722 | ||
| Pseudoterranova krabbei | 13,948 | NC_031646 | ||
| Pseudoterranova bulbosa | 13,957 | KU558720 | ||
| Pseudoterranova cattani | 13,950 | KU558721 | ||
| Ascarididae | Ascaris lumbricoides | 14,281 | NC_016198 | |
| Ascaris lumbricoides China | 14,303 | HQ704900 | ||
| Ascaris ovis | 14,288 | NC_036666 | ||
| Ascaris suum | 14,284 | NC_001327 | ||
| Ascaris suum China | 14,311 | HQ704901 | ||
| Ascaris sp. Chimpanzee | 14,268 | KC839986 | ||
| Ascaris sp. gibbon | 14,274 | KC839987 | ||
| Baylisascaris ailuri | 14,657 | HQ671080 | ||
| Baylisascaris procyonis | 14,781 | NC_016200 | ||
| Baylisascaris schroederi | 14,778 | NC_015927 | ||
| Baylisascaris transfuga | 14,898 | NC_015924 | ||
| Ophidascaris baylisi | 14,784 | MW880927 | ||
| Ophidascaris sp. | 14,660 | MK106624 | ||
| Parascaris equorum | 13,899 | NC_036427 | ||
| Parascaris univalens | 13,920 | NC_024884 | ||
| Toxascaris leonina | 14,310 | NC_023504 | ||
| Heterocheilidae | Ortleppascaris sinensis | 13,828 | NC_036669 | |
| Toxocaridae | Toxocara canis | 14,322 | NC_010690 | |
| Toxocara canis Australia | 14,163 | EU730761 | ||
| Toxocara cati | 14,029 | NC_010773 | ||
| Toxocara malaysiensis | 14,266 | NC_010527 | ||
| Cucullanidae | Cucullanus robustus | 13,972 | NC_016128 | |
| Heterakoidea | Ascaridiidae | Ascaridia columbae | 13,931 | NC_021643 |
| Ascaridia galli | 13,977 | NC_021642 | ||
| Ascaridia sp. | 13,862 | JX624730 | ||
| Heterakidae | Heterakis beramporia | 14,012 | NC_029838 | |
| Heterakis dispar | 13,995 | NC_042411 | ||
| Heterakis gallinarum | 13,973 | NC_029839 | ||
| Oxyuroidea | Oxyuridae | Enterobius vermicularis | 14,010 | EU281143 |
| Wellcomia siamensis | 14,128 | NC_016129 |