The increasing resistance of microbes to antibiotics poses one of the greatest challenge for modern medicine, particularly in the context of hospital-acquired and systemic infections. In this state, it is imperative to swiftly and precisely identify both the pathogen and its drug susceptibility profile, to implement effective treatment. Phenotypic antimicrobial susceptibility testing (AST) is widely regarded as the gold standard in the field of antimicrobial resistance diagnostics. This is since it relies on the assessment of microbial growth or its inhibition in the presence of an antibiotic (Lin et al. 2025). The primary benefit of this approach is its capacity to discern resistance irrespective of its underlying mechanism. However, a significant constraint pertains to the system’s protracted turnaround time, which customarily extends to approximately 24 hours (Astudillo et al. 2024; Elbehiry and Abalkhail 2025). Meanwhile, DNA-based and enzymatic assays offer quicker results, but are limited in scope, insofar as they can detect only specific resistance mechanisms. This highlights the pressing need for novel tools that facilitate rapid and comprehensive diagnostics, which are crucial for the timely and effective administration of therapy (Idelevich et al. 2018; Elbehiry and Abalkhail 2025). In response to the limitations of alternative methods and the growing demand for more effective solutions, Matrix-Assisted Laser Desorption/Ionization - Time of Flight with mass spectrometry (MALDI-TOF MS) has emerged as a pivotal technology in microbial diagnostics. This approach has the potential to transform the realm of diagnostics by providing rapid and precise methods for identifying micro-organisms. Furthermore, it can expand the horizons of research by opening new avenues for assessing the susceptibility of microorganisms to antibiotics, thereby facilitating more informed and effective treatment options. MALDI-TOF MS is a highly sensitive method, often requiring only a small amount of biological material for analysis. The basis of this technique is mass spectrometry, an advanced analytical technique that enables the quantitative detection of ionized molecules by measuring their mass-to-charge ratio (m/z). As asserted by Calderaro and Chezzi (2024) and Sogawa et al. (2021), the generation of a unique spectrum by each molecule enables precise identification of the composition of the sample (Sogawa et al. 2021; Calderaro and Chezzi 2024). The purpose of this article is to present the current possibilities of using MALDI-TOF MS technology in the determination of drug susceptibility of microorganisms and in the search for new target sites in their cellular structures for existing and newly developed antimicrobial compounds. This study focuses on the most recent advancements in the field of resistance to biomarker detection, incorporating the utilization of state-of-the-art proteomic methodologies. Furthermore, it explores prospective avenues for the advancement of this technology within the broader context of the escalating issue of antibiotic resistance.
This review is a critical and comparative in nature attempt, focusing on clinical usefulness, especially in the context of predicting the effectiveness of therapies for multidrug-resistant microorganism infections, technical limitations, costs associated with data analysis or reagent selection, as well as participation of a spectrum analysis specialist, and barriers to the implementation of selected MALDI-TOF MS methods for the diagnosis of resistance to various antimicrobial drugs.
Previous studies on resistance detection using MALDI-TOF MS have evaluated a relatively narrow range of antibiotics and species, with a clear focus on selected, clinically important combinations of antimicrobials (Table 1). Most studies have focused on detecting proteins associated with β-lactam resistance (particularly cephalosporin and carbapenem resistance) in Enterobacterales and non-fermenting bacilli such as Pseudomonas aeruginosa and Acinetobacter baumannii. These studies have evaluated both β-lactamase activity (including ESBL and carbapenemase types) and complex MDR phenotypes (Sorensen et al. 2020; Astudillo et al. 2024; Elbehiry and Abalkhail 2025).
Examples of protein biomarkers of resistance detected using MALDI-TOF MS
| Type of antibiotic resistance | Pathogen species | Biomarker (m/z, protein) | Related gene/mechanism | Phenotypic significance | Estimated effectiveness | Restrictions/remarks | Ref. |
|---|---|---|---|---|---|---|---|
| Klebsiella pneumoniae carbapenemase (KPC) | Klebsiella pneumoniae | Plasmid peaks associated with the presence of blaKPC, no single universal m/z value | Presence of plasmids carrying blaKPC | Correlation with the phenotype of KPC-dependent carbapenem resistance | High specificity and good sensitivity for detecting KPC-associated protein biomarkers | Reliability is limited by plasmid variability, meaning false negatives may occur and negative results require confirmatory molecular testing | (Florio et al. 2020) |
| Extended-spectrum class C beta-lactamase | Acinetobacter baumannii | Peak at 40,279 ± 87 m/z contributing to Acinetobacter derived cephalosporinases (ADC) family | Not stated | Not stated | High sensitivity (96%) and specificity (73%) in comparison to the microdilution imipenem susceptibility testing | Not stated | (Florio et al. 2020) |
| Methicilin-resistant Staphylococcus aureus (MRSA) | Staphylococcus aureus | Peak at 2415 ± 2 m/z corresponding to phenol-soluble protein toxin (PSM-mec) | PSM-mec can predict carriage of mecA – gene coding penicillin binding protein 2A (PBP2A) responsible for MRSA phenotype | High specificity for PSM-mec correlating with MRSA but the protein isn’t present in every isolate | Specificity for PSM-mec close to 100% | The majority of coagulase-negative staphylococci (CoNS) doesn’t produce PSM-mec thus MALDI-TOF MS has limited usefulness there | (Florio et al. 2020) |
| Methicilin-resistant Staphylococcus aureus (MRSA) | Staphylococcus aureus | Peak at 4594 m/z contributing to 50S ribosomal protein L28 | mecA gene | Distinguishing between MRSA and MSSA based on spectrum pattern, without additional preparation | High specificity of 96.8% to detect MRSA isolates | Not stated | (Flores-Treviño et al. 2019) |
| Ribotype 027 | Clostridioides difficile | Peak at 6654 m/z corresponding to 30S ribosomal protein S20 | 30S ribosomal protein binding to 16S rRNA | Differentiation of ribotype 027 from non-027 ribotypes; association with hypervirulent strains | High sensitivity, specificity and AUC values for each peak (0.96 and 0.99) | Peak at 6712 m/z is present in multiple ribotypes, complicating differentiation between 027 and 176 | (Flores-Treviño et al. 2019) |
| Carbapenemase | Acinetobacter baumannii | Peaks at 6304 and 6332 m/z corresponding to NADH-quinone oxidoreductase subunit K | blaOXA-24 and blaOXA -58 | Differentiation of MDR A. baumannii and discrimination between blaOXA-58- and blaOXA-24-positive isolates | High AUC values (0,99), specificity and sensitivity | Limited prior evidence and comparison with indirect methods. Biomarkers identified in MDR isolates; previous studies relied on carbapenem hydrolysis assays rather than direct MALDI-TOF MS | (Flores-Treviño et al. 2019) |
| MDR/non-MDR strains | Pseudomonas aeruginosa | Loss of signals at 2726 and 5455 m/z in MDR strains | Presumably UPF0270 protein Pfl01_4103 (2726 m/z) and UPF0391 membrane protein Patl_1732 (5455 m/z) | Differentiation between MDR and non-MDR isolates based on signal loss | Moderate to low effectiveness (AUC 0.81–0.84; combined sensitivity 75%, specificity 74.1%) | Signal loss is non-specific, may be due to non-genetic factors, and cannot be used as a reliable biomarker | (Flores-Treviño et al. 2019) |
One of the earliest methods for detecting antibiotic resistance using MALDI-TOF involves measuring the hydrolysis of β-lactam antibiotics. Bacteria that produce β-lactamases can cause the breakdown of the β-lactam ring of the antibiotic. This is visible in the mass spectra as a decreasing peak that corresponds to the intact antibiotic and increasing peaks representing its hydrolysis products (Florio et al. 2020; Elbehiry et al. 2022). Extended-spectrum β-lactamase (ESBL)-producing bacteria can be detected after 30 minutes of incubation (reaction time of one hour is recommended for cefotaxime and cefpodoxime). Ceftazidime, not recommended for ESBL identification due to its low susceptibility to hydrolysis, has been demonstrated to reduce the effectiveness of MALDI-TOF MS analysis. In such cases, the incubation period should be extended incubation period (Oviaño and Rodríguez-Sánchez 2021). It was employed to evaluate the presence of β-lactam resistance, a process which is initiated by the detection of β-lactamase activity. In the event of a negative result, it is imperative to exclude the possibility of alternative resistance mechanisms (Florio et al. 2020).
In the case of Gram-positive bacteria, particular attention was paid to methicillin resistance in Staphylococcus aureus and S. epidermidis, and to glycopeptide resistance in Enterococcus faecium. This was usually in relation to individual classes of antibiotics, such as oxacillin and cephalosporins for methicillin resistance, and vancomycin and teicoplanin for glycopeptide resistance, and to a limited number of strains representing selected epidemic clones. Relatively little data is available for other drug groups such as aminoglycosides, fluoroquinolones or macrolides. Analyses covering broader antibiotic panels and diverse species outside the ESCAPE group remain scarce, which makes it difficult to generalise the results and compare the effectiveness of different approaches in real clinical settings. Previous studies on the detection of enzymes responsible for the ESBL and AmpC mechanisms using MALDI-TOF MS have shown that the diagnostic parameters achieved differ depending on the antibiotic or chemotherapeutic agent used. When ceftriaxone was analysed using automated software (e.g. MBT STAR-BL), sensitivity and specificity of 100% were achieved after 30 minutes of incubation, establishing it as one of the most reactive and selective markers of β-lactam hydrolysis (Astudillo et al. 2024). Cefotaxime and cefpodoxime showed significantly lower sensitivity. In some studies, ceftriaxone produced approximately 70% and 20% more positive results than these cephalosporins while maintaining high specificity. Ceftazidime proved particularly problematic as visible hydrolysis effects required incubation times of often ≥120 minutes and shortening the incubation time led to false negative results in strains with weaker β-lactamase activity. By contrast, cefepime demonstrated very high sensitivity (up to 100%) in detecting AmpC cephalosporinase activity. However, this came at the cost of low specificity (approximately 27%), due to unexpected hydrolysis by various β-lactamases. This limits its usefulness as a selective marker. The wide variation in parameters for individual cephalosporins emphasises the importance of choosing the right antibiotic for the MALDI-TOF test, and this choice must be optimised and validated for each specific strain population and expected resistance mechanisms (Burckhardt and Zimmermann 2018).
The detection of biomarkers associated with antimicrobial resistance using MALDI-TOF MS involves identifying specific mass peaks (m/z) in the obtained proteomic spectra that correlate with resistance enzymes, structural modifications or characteristic protein profiles.
For example, one potential methodology for the identification of carbapenem resistance in bacteria is predicated upon the detection of plasmids harbouring carbapenemase genes. Johansson et al. (2014) conducted a study on Bacteroides fragilis, anaerobic pathogen that may harbour the cfiA gene, which encodes a metallo-β-lactamases (MBL). The presence of this gene has been demonstrated to be associated with resistance to nearly all β-lactam antibiotics. The employment of MALDI-TOF MS facilitated the differentiation of B. fragilis strains that were positive and negative for the cfiA gene, based on a characteristic spectral shift in the 4000–5500 m/z range (Johansson et al. 2014; Yoon and Jeong 2021). The identification of carbapenemase-producing bacteria and the subsequent reporting of such findings to the relevant authorities is of paramount importance within the domain of clinical diagnostics. This approach is instrumental in the prevention of the dissemination of resistance. The utilisation of contemporary methodologies, such as MALDI-TOF MS technique, has been demonstrated to markedly expedite the process, thereby facilitating the expeditious implementation of therapeutic and epidemiological interventions.
Another example is the detection of methicillin resistance in staphylococci, which is mainly based on identifying the PSM-mec biomarker (m/z ≈ 2,415 Da). This peptide is produced as part of the SCCmec cassette carrying the mecA gene and can be analysed using classification algorithms to identify diverse spectral patterns. PSM-mec is characterised by very high specificity (approximately 100%) for MRSA and MRSE strains, while allowing rapid differentiation from methicillin-sensitive S. aureus (MSSA) and S. epidermidis (MSSE) strains. However, the sensitivity of this biomarker is estimated to be 40–60%, since not all MRSA isolates express it in sufficient quantities. Furthermore, the analysis of complex spectral profiles in the 2–7 kDa range (e.g., the m/z 4594 Da doublet) using machine learning (ML) algorithms achieves 83% sensitivity and 97% specificity for MRSA. However, this method depends on local epidemic clones and requires periodic database updates (Flores-Treviño et al. 2019; Florio et al. 2020; Santos et al. 2025).
A similar approach is used for E. faecium to distinguish between susceptible strains (VSE) and vancomycin-resistant strains (VRE) that exhibit the VanA or VanB phenotype. This method uses spectral patterns containing cellular proteins that differ in peak shifts associated with cell wall remodelling, which depends on VanA (strong resistance to vancomycin and teicoplanin) versus VanB (vancomycin only). Although studies have demonstrated the good specificity (90–95%) of these methods, the sensitivity is moderate (70–85%) and depends heavily on the genetic background of the population and the quality of the classification algorithms. The absence of a single universal peak for all Van clusters has been demonstrated, which may limit the universality of this approach (Vrioni et al. 2018; Andreadi et al. 2025).
These two examples demonstrate the potential of MALDI-TOF for the rapid detection and differentiation of resistance phenotypes. However, they also highlight the need for it to be used in conjunction with phenotypic or genetic methods to ensure full diagnostic reliability.
Unlike molecular methods, which are based on the direct amplification and/or detection of point mutations in known resistance genes (e.g., mecA and blaKPC), the MALDI-TOF MS peak-based approach identifies indirect proteomic biomarkers, i.e., specific mass peaks that correlate with protein changes associated with resistance. For instance, the PSM-mec peak (m/z 2415 Da) suggests a high likelihood of detecting an MRSA strain with good specificity, albeit with only around 60% sensitivity. In contrast, PCR on the mecA gene achieves over 95% effectiveness, regardless of gene expression levels. The limitation of the MALDI-TOF MS technique is that it depends on actual protein synthesis, and there is spectral variability between clones. PCR, on the other hand, can produce false positive results in isolates with an unexpressed gene. The two approaches are therefore complementary: MALDI provides faster differentiation of detected resistance phenotypes, while PCR-based reactions provide greater certainty of the result (Flores-Treviño et al. 2019; Candela et al. 2022; Santos et al. 2025).
Modification of the drug binding site by bacteria may block the action of the antimicrobial agent, making the bacteria resistant to drugs from that class. An example of this type that is of great clinical importance is Lipid A in Gram-negative bacteria. It is the target of polymyxins, and when lipid A is modified, bacteria become resistant to colistin, the last-resort antibiotic. The accredited method for determining colistin resistance is the broth dilution method, but this technique requires several days to obtain results (Yoon and Jeong 2021).
A technique called
Despite the fact that the addition of colistin during the preparatory stage is a time-consuming process, it can be a relatively expeditious method that requires less than five hours. One of the disadvantages of this method is that it is only possible to test a small number of strains; however, the results obtained in terms of sensitivity could be very encouraging (Furniss et al. 2019; Jeannot et al. 2021; Yoon and Jeong 2021; Rubens et al. 2024).
An alternative lipid-based MALDI-TOF MS approach for the detection of colistin resistance is the fast lipid analysis technique (FLAT). FLAT is based on direct on-plate extraction of microbial lipids on a heated stainless steel MALDI target, which minimizes sample handling and eliminates the need for centrifugation. The entire workflow can be completed in less than one hour and is highly parallelizable, making the method amenable to automation in a clinical laboratory setting. FLAT generates reproducible lipid barcode spectra that allow species-level identification of bacteria and fungi, while simultaneously enabling the detection of selected antimicrobial resistance-associated lipid modifications. The method can readily detect lipid A modifications associated with colistin resistance, including both intrinsic resistance and plasmid-mediated resistance conferred by mcr-1 in Gram-negative bacteria. Although FLAT does not detect all resistance mechanisms, it enables combined microbial identification and resistance detection within a single MALDI-TOF MS assay (Furniss et al. 2019; Sorensen et al. 2020) current lipid extraction methods require significant hands-on time and are thus not suited for direct adoption as a clinical assay for microbial identification. Here, we have developed a method for lipid extraction directly on the surface of stainless-steel matrix-assisted laser desorption/ionization time-of-flight mass spectrometry (MALDI-TOF MS. The MALDIxin and FLAT systems are two complementary approaches for detecting colistin resistance through the analysis of lipid A modifications in Gram-negative bacteria (mainly Pseudomonas spp., Acinetobacter spp., Klebsiella spp.). The MALDIxin approach involves acid hydrolysis and lipid extraction, achieving baseline sensitivity of 57% (up to 90% with colistin induction after 4 hours), whereas the FLAT approach offers faster (<60 minutes) and simpler preparation without extraction, enabling simultaneous species identification and achieving sensitivity of over 95%. While MALDIxin is better documented in the clinical setting for P. aeruginosa, FLAT shows greater potential for automation. Nevertheless, both solutions are currently limited to detecting colistin-resistant strains and require further multicentre validation.
MALDI Biotyper-Antibiotic Susceptibility Test Rapid Assay (MBT-ASTRA) is an antibiotic resistance semi-quantitative detection method based on calculating and comparing the area under the curve (AUC) of the mass spectra of microorganisms incubated with and without antimicrobials, using MALDI-TOF MS software (Florio et al. 2020). In the study Sauget et al. (2018), a rapid method for Escherichia coli isolates from blood cultures detecting resistance to amoxicillin and cefotaxime was performed. The MBT-ASTRA software was utilised for the classification of isolates as either susceptible or resistant to both antimicrobials, achieving 95% and 84% accuracy for AMX and CTX, respectively, when compared to the reference agar dilution method. The use of MALDI-TOF MS has been shown to facilitate the acquisition of results within a brief timeframe following the establishment of a positive blood culture. This offers a substantial advantage in the management of sepsis (Sauget et al. 2018).
Stable Isotope-Labelling-Based Assays for Peak Shift Detection (MBT-RESIST) is based on the use of isotopically labelled nutrients, most commonly amino acids containing 13C. This enables the evaluation of antimicrobial activity through mass shift spectrum analysis. A comparative analysis of the spectra obtained under both conditions (with and without the drug) was undertaken in order to ascertain the microorganism’s susceptibility to the tested antimicrobial agent (Yoon and Jeong 2021). It was performed to discriminate MSSA from MRSA strains with some misclassification incidents (e.g., for oxacillin or cefoxitin) (Sparbier et al. 2013) the fast and cost-efficient detection of microbial resistance is an important clinical requirement for optimal therapeutic intervention. Current routine assays take at least 5 h, but in most cases an overnight incubation is necessary to identify resistant isolates. The usage of matrix-assisted laser desorption ionization–time of flight mass spectrometry (MALDI-TOF MS or for carbapenem, aminoglicosides and fluorochinolone resistance distinction in P. aeruginosa strains (Jung et al. 2014). The above applications suggest that this method should be considered primarily as a research or validation tool.
A novel approach to identifying antibiotic resistance is the direct-on-target microdroplet growth assay (DOT-MGA), based on MALDI-TOF MS. In the study conducted by Idelevich et al. (2018), evaluated the effectiveness of the detection of meropenem resistance in K. pneumoniae and P. aeruginosa. The test demonstrated 100% and 83.3% sensitivity and specificity (for K. pneumoniae and P. aeruginosa, respectively). The DOT-MGA method can be considered as rapid, accurate, and user-friendly technique for the detection of carbapenem resistance in Gram-negative bacteria. In a subsequent study (Idelevich et al. 2018) we investigated a direct application of this method on positive blood cultures (BCs), this method was also applied for the diagnosis of sepsis, testing blood samples infected with both meropenem-susceptible and -resistant Enterobacterales isolates. The accuracy of the method was reported to 96.3% be, sensitivity 91.7%, and specificity 100%. In addition, it was shown that this method allows for the detection of resistance and enables simultaneous species identification (Idelevich et al. 2018)we investigated a direct application of this method on positive blood cultures (BCs. As demonstrated by Idelevich et al. (2021), the DOT-MGA method, proved to be an effective means of rapid analysis of various samples.
As previously stated, conventional methods for assessing antimicrobial resistance are not without their limitations. MALDI-TOF may offer a solution to these challenges. This method supports universal phenotypic AST, may allow the rapid microbial identification and distinction between resistant and susceptible strains (irrespectively of species, antibiotic type, or resistance mechanism). Even though this method may appear to be rather complex, it is, in fact, the precise opposite.
Firstly, the processes of sample preparation and data processing are both uncomplicated. Nonetheless, the composition of the culture medium exerts a significant influence on the outcomes of MALDI-TOF MS measurements. This solution has the capacity to eliminate the necessity for labour-intensive and time-consuming centrifugation and wash steps, thereby streamlining the process. Furthermore, it facilitates the implementation of automation, thus enhancing efficiency and reducing labour costs. The duration required for the attainment of results is considerably reduced. MALDI-TOF MS facilitates optimal laboratory workflow, enabling the acquisition of all clinically relevant information on the same day. This, in turn, results in faster drug admission and enhanced patient management.
Secondly, MALDI-TOF MS has a wide range of applications. One of the objectives is the development of clinically efficient rapid tests for antimicrobial resistance, with a view to supporting the selection of suitable antibiotics in each therapeutic context. Another potential application of MALDI-TOF MS is the creation of panels comprising multiple antibiotics targeting the same biological target. This process can be automated, integrating pipetting and incubation within a single instrument. Furthermore, MALDI-TOF MS targets can be prepared in advance with antimicrobial substances, for example, in lyophilised form. Furthermore, studies have reported on the efficacy of MALDI-TOF in detecting antimicrobial resistance, even in cases where minimal inhibitory concentrations (MICs) are borderline elevated. Furthermore, the efficacy of MALDI-TOF MS in assessing resistance to last-resort antibiotics is a noteworthy consideration.
In summary, MALDI-TOF MS can offer a rapid, precise and user-friendly assay, suitable for both manual execution and potential automation with multi-antibiotic capability. The method is distinguished by its simplicity and efficiency in sample preparation, as well as its reliance on a reliable and user-friendly data analysis algorithm. It has been demonstrated that MALDI-TOF MS can be a very useful instrument for the purpose of antimicrobial resistance testing in comparison with the standard methods that are currently in use (Idelevich et al. 2018).
Although pilot studies have produced promising results, the use of MALDI-TOF MS for diagnosing antibiotic resistance is hindered by many significant limitations, which have so far prevented its routine implementation in clinical practice (Figure 1).
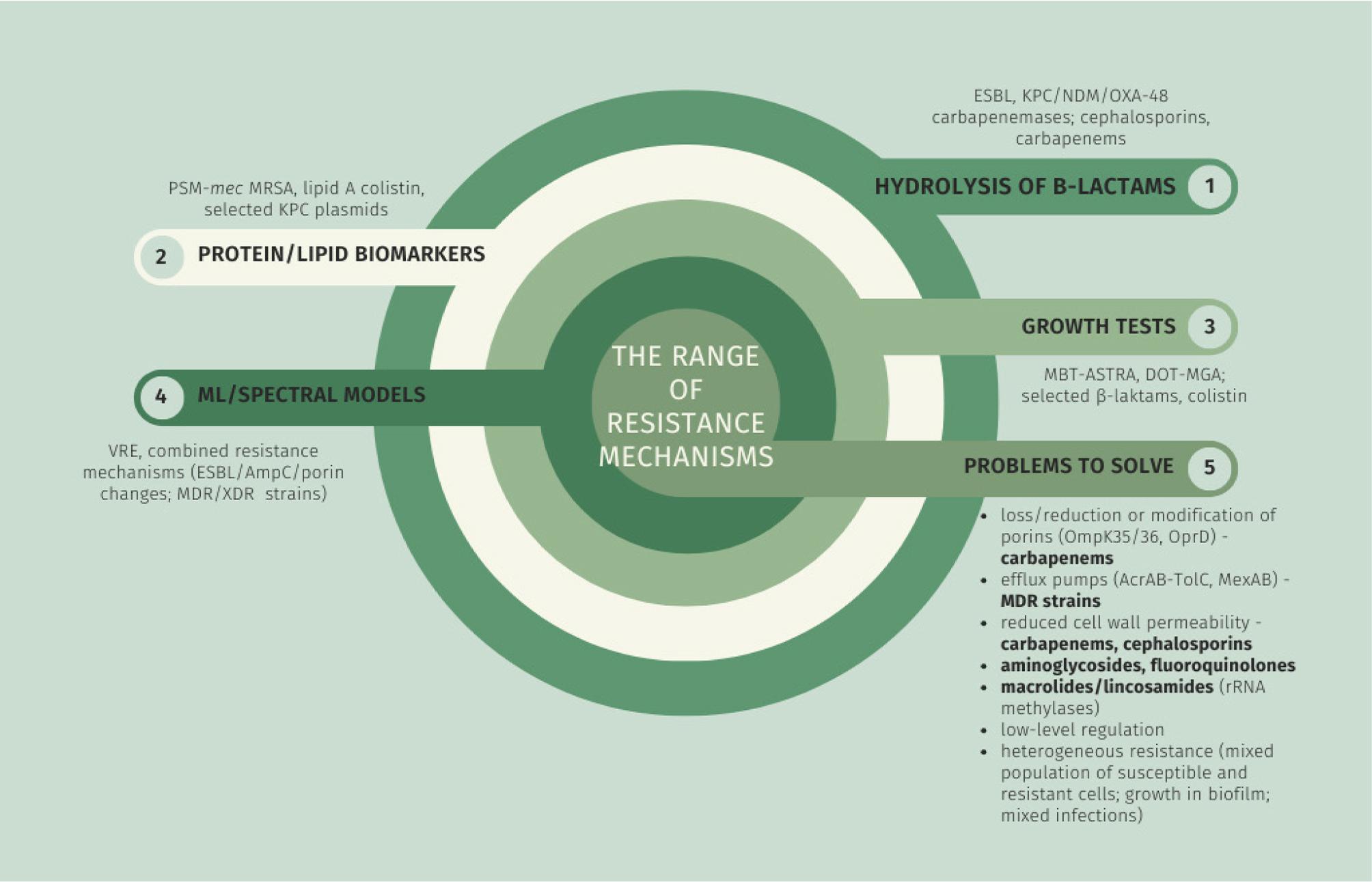
Examples of limitations of MALDI-TOF MS in detecting different classes of antibiotic resistance mechanisms
The first and most significant problem is the limited range of resistance mechanisms that can be detected. Hydrolysis-based methods only detect mechanisms that lead to rapid degradation of antibiotics in the environment. These methods cannot detect resistance resulting from transport mechanisms, such as porin modification (reduction in number or change in structure; e.g., OmpK35/36 for K. pneumoniae, OprD for P. aeruginosa), reduced permeability of cell membranes (to carbapenems or cephalosporins), efflux mechanisms via protein membrane pumps or the overexpression of penicillin-binding proteins (PBPs in MRSA; subtle mutations in PBP in penicillin-resistant pneumococci; overexpression of AcrAB-TolC efflux pumps in Enterobacteriaceae, MexAB in P. aeruginosa). Biomarkers in the spectrum (whether protein or lipid peaks) are currently only available for certain resistance mechanisms, such as specific carbapenemases, PSM-mec in some MRSA strains, and particular P. aeruginosa clones. In this context, information on resistance is highly fragmented and depends heavily on the species or clone of microorganism being studied (Vrioni et al. 2018; Cordovana et al. 2019; Florio et al. 2020).
Another issue is the dependence on test conditions and the absence of interlaboratory standardization. Most available protocols do not have any of the universally accepted interpretation thresholds/cut-off-values that align with EUCAST (European Committee on Antimicrobial Susceptibility Testing) or CLSI (Clinical and Laboratory Standards Institute) guidelines. Numerous studies have shown that isolates with MICs close to the breakpoint are most frequently misclassified (Anderson 2016; Cordovana et al. 2019; Florio et al. 2020). Methods such as MBT-ASTRA tend to produce false-positive results for sensitivity or resistance in the subinhibitory/sublethal concentration range because the differences in growth and spectra are minimal. Similarly, combined mechanisms (e.g., ESBL + AmpC + porin changes; typical MDR/XDR A. baumannii or P. aeruginosa) or heterogeneous resistance, where a portion of the cell population grows differently to the rest (e.g., in biofilms population or mixed infections), can directly distort the average signal in the spectrum, making it difficult to distinguish between phenotypes. All this makes it difficult to compare results between laboratories and means that classic Antimicrobial Susceptibility Testing (AST) methods, as recommended internationally, continue to be used. Furthermore, diagnostic parameters, including sensitivity and specificity, depend on the specific antimicrobial agent, its concentration, the incubation time and the medium composition. Therefore, they require optimization for each species-drug combination. This makes it impossible to compare results between centers and standardize reporting (Hosoda et al. 2025).
A third significant limitation of the MALDI-TOF MS method, particularly when used in conjunction with a ‘subtyping’ or machine learning (ML) approach, is that it recognizes spectral patterns associated with resistance rather than directly detecting the gene product (e.g. protein or enzyme) that encodes the resistance trait (Cordovana et al. 2019; Astudillo et al. 2024; Elbehiry and Abalkhail 2025). This model applies to resistance to aminoglycosides and fluoroquinolones, which is mainly caused by efflux and rRNA modifications. The same applies to resistance to glycopeptides (e.g., vancomycin). Strains with the VRE phenotype typically exhibit nucleotide mutations within the van gene. Analysis of these strains may reveal a lack of specific peaks, although some ML models attempt to predict this indirectly (Elbehiry and Abalkhail 2025). Therefore, the effectiveness of these solutions may decrease when new clones or strains emerge with mechanisms that are not included in the training set of the protocol.
Despite promising performance in pilot studies, MALDI-TOF MS-based assays or technologies for antimicrobial resistance are still only partially standardized and have been validated on relatively small, single-centre collections of isolates, often focusing on selected species-antibiotic combinations (Sauget et al. 2018; Florio et al. 2020; Elbehiry and Abalkhail 2025). Cut-off values, incubation conditions, and interpretation criteria are not harmonized across laboratories, which limits comparability of results and hinders the inclusion of these methods in guideline-driven workflows. For most pathogens and drug classes, MIC determination by standardized phenotypic methods remains the reference, and MALDI-TOF MS can currently only complement, rather than replace, conventional AST. Despite everything it has the potential to complement routine microbiological diagnostics and clinical care, provided several key challenges are overcome (Elbehiry and Abalkhail 2025).
First and foremost, the implementation of AST based on MALDI-TOF MS requires access to a general mass spectrometry platform, which is still unavailable in many smaller laboratories. It also necessitates the development of specialised software modules, particularly with regard to the enzymatic or molecular basis of resistance mechanisms (Florio et al. 2018; Sauget et al. 2018; Rychert 2019; Florio et al. 2020; Gajic et al. 2022; Weis et al. 2022; Calderaro and Chezzi 2024; Hosoda et al. 2025).
Some tests based on MALDI-TOF MS technology require additional incubation steps and sample processing. They also require highly specialised knowledge for data analysis. These factors can only partially offset the theoretical time savings compared to rapid phenotypic or molecular tests. The effectiveness of these techniques also depends on the quality and completeness of the spectrum library. Rare species, new taxa and environmental strains are often poorly represented in system libraries, which can result in incorrect species identification or failure to identify. Therefore, gradually expanding the microorganism library (including anaerobic bacteria and fungi) may enable the development of effective, rapid, comprehensive solutions (identification and drug sensitivity testing) in the future, which could be important for selecting therapy and interpreting resistance (Astudillo et al. 2024; Tejan et al. 2025). Training advanced ML models on large, multi-centre mass spectrometry datasets may significantly improve the sensitivity and specificity with which complex resistance phenotypes are recognised (Astudillo et al. 2024; Lin et al. 2025). In the long term, multicentre, international research projects aimed at harmonising cut-offs, validating antibiotic panels and creating EUCAST/CLSI guidelines for MALDI-AST appear to be necessary. This step could be crucial in enabling MALDI-based techniques to compete with, or even surpass, classical AST methods in routine microbiological diagnostics (Florio et al. 2020). Another challenge will be identifying new biomarkers for transport (e.g. efflux and porin loss) and regulatory resistance. Identifying these biomarkers using a combination of MALDI proteomics and genomics methods (e.g. NGS, multi-label models and gene expression profiling) could provide a more comprehensive view of resistance mechanisms within a single workflow (Astudillo et al. 2024; Elbehiry and Abalkhail 2025; Hosoda et al. 2025; Lin et al. 2025).
MALDI-TOF MS has been instrumental in revolutionizing the field of microbial diagnostics by facilitating rapid and accurate identification of pathogens from minimal biological material. More recently, its application has been extended to the detection of antimicrobial resistance through methods such as the MALDIxin test, MBT-ASTRA, MBT-RESIST, and DOT-MGA. These techniques enable the assessment of resistance by detecting enzymatic activity, resistance-associated biomarkers, or drug target modifications, delivering clinically actionable results within hours. The versatility, speed, and simplicity of MALDI-TOF MS, in conjunction with its capacity for automation and the detection of multidrug resistance in microorganisms, underscores its promising role as an alternative or complementary approach to traditional AST. However, challenges such as standardization of protocols, increased robustness across diverse bacterial species, and integration into routine clinical workflows remain to be addressed.
In conclusion, MALDI-TOF MS holds considerable promise in advancing the rapid diagnosis of antimicrobial resistance. It offers a new possibility for early targeted therapy of infections and better containment of resistance spread, provided that further technological and clinical validations are achieved. Over the next 5–10 years, MALDI-TOF MS enhanced with ML algorithms and available in automated systems may become a standard component of antibiotic resistance routine diagnostics (Burckhardt and Zimmermann 2018; Tejan et al. 2025). This technology would be able to support traditional AST methods and molecular tests, particularly in reference laboratories and highly specialized centers. This would aid both clinical decisions regarding the selection of effective antibiotic therapy and the surveillance of the spread of resistance among microorganisms.