Fig. 1.
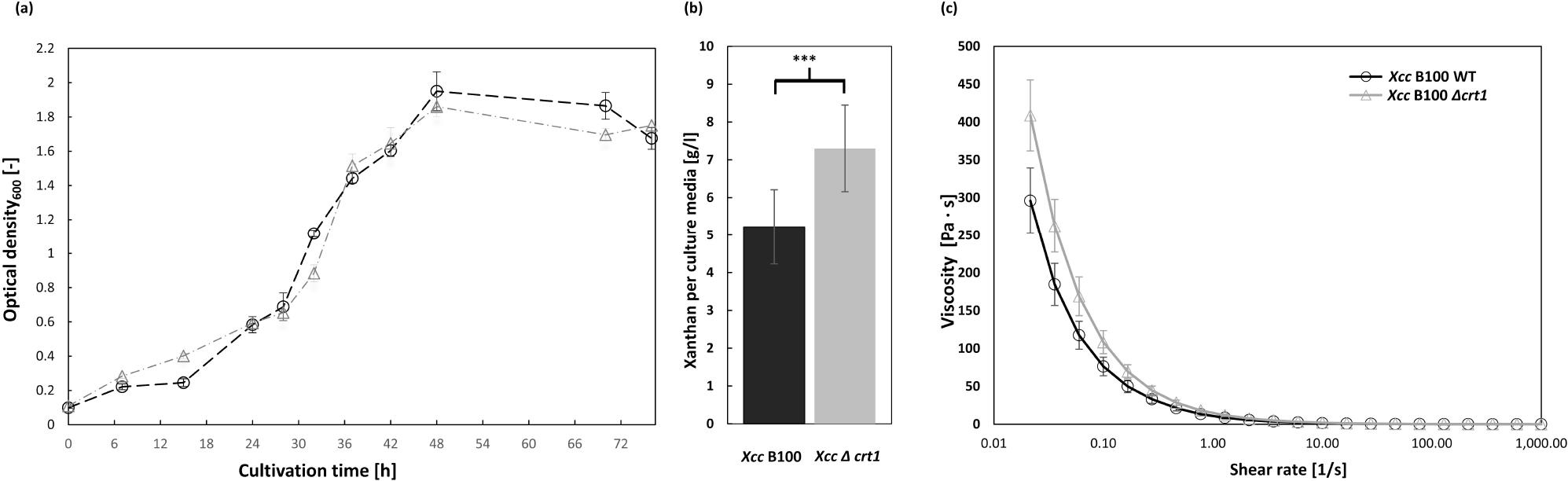
Fig. 2.
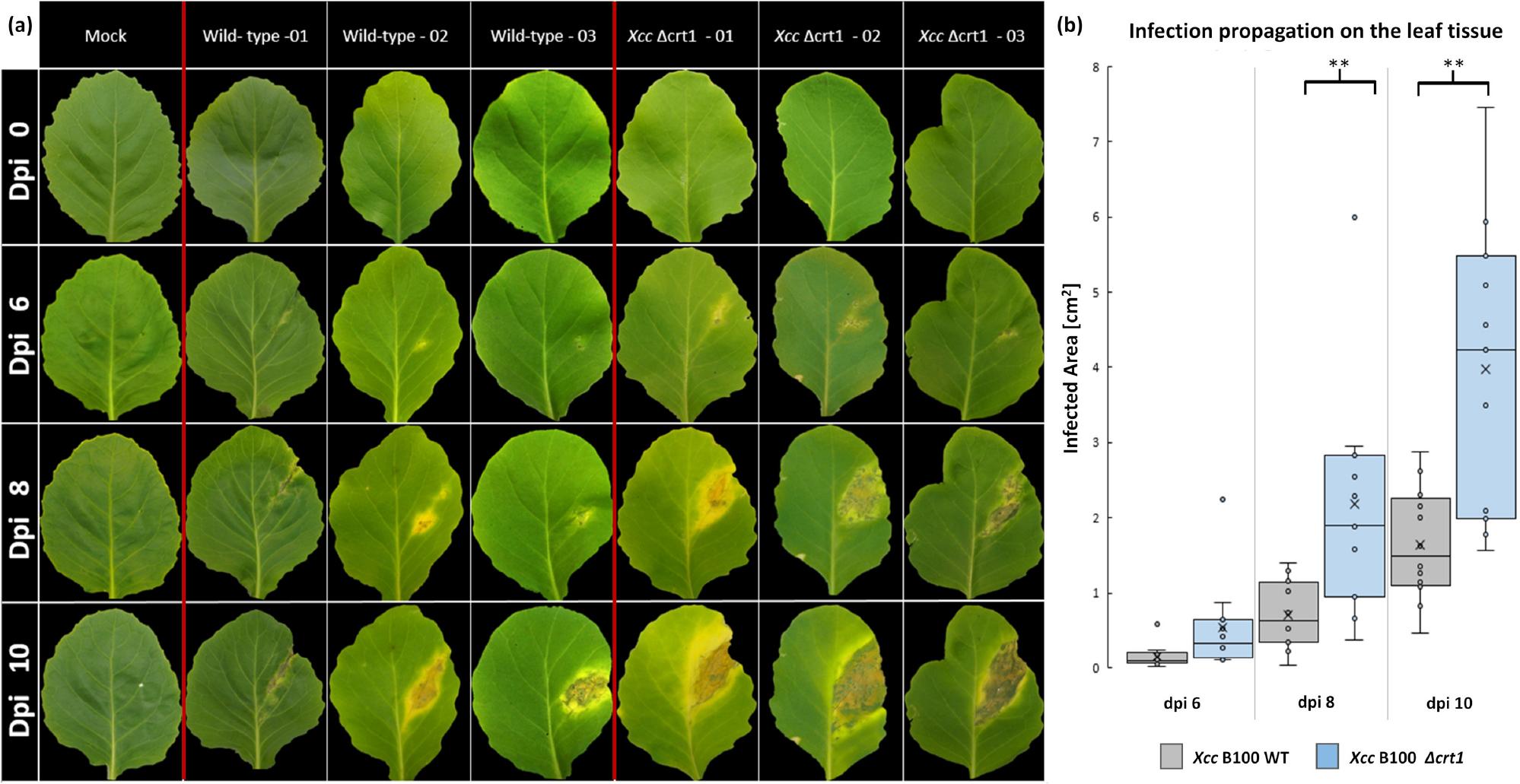
Fig. 3.
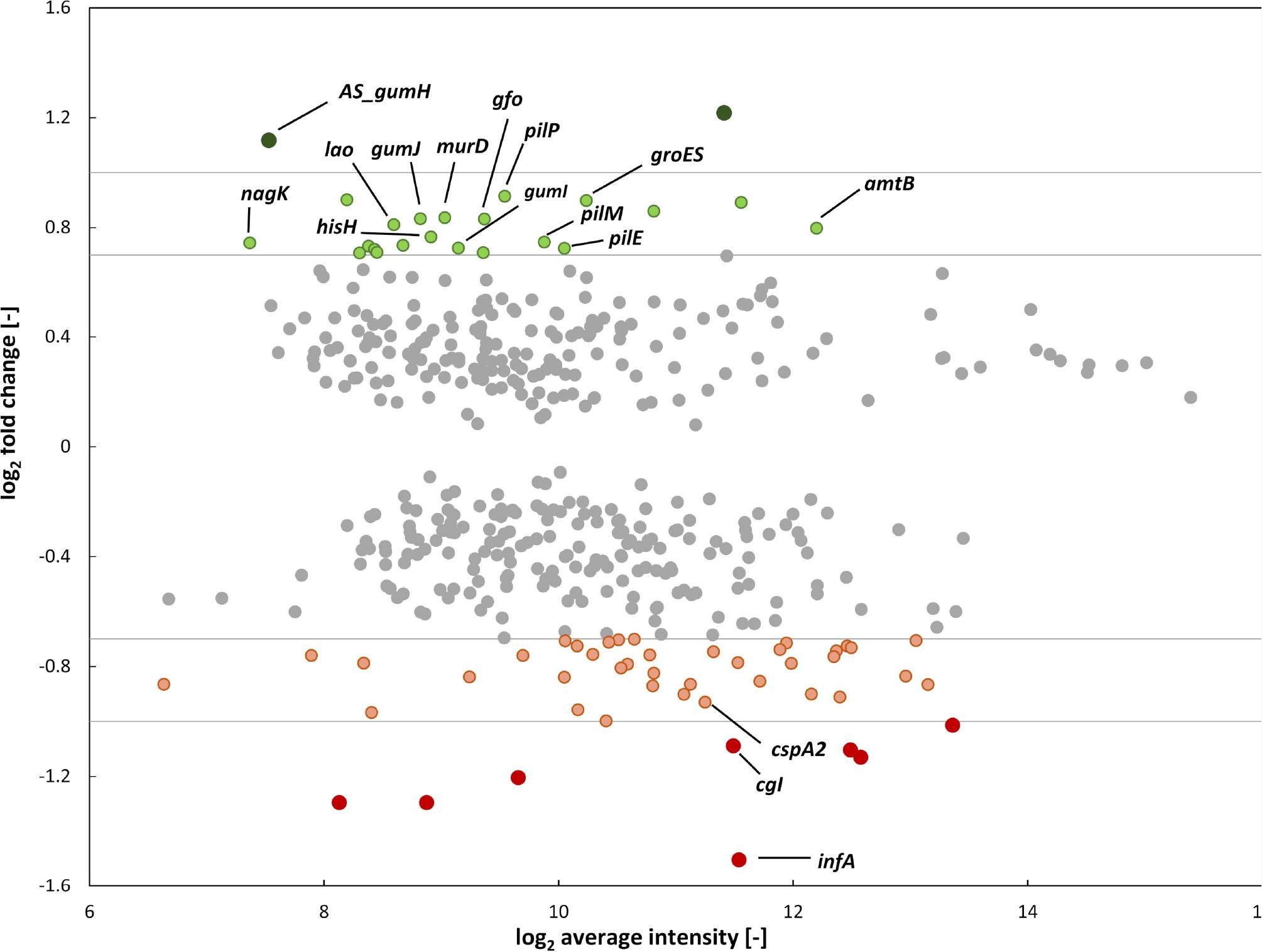
Fig. 4.
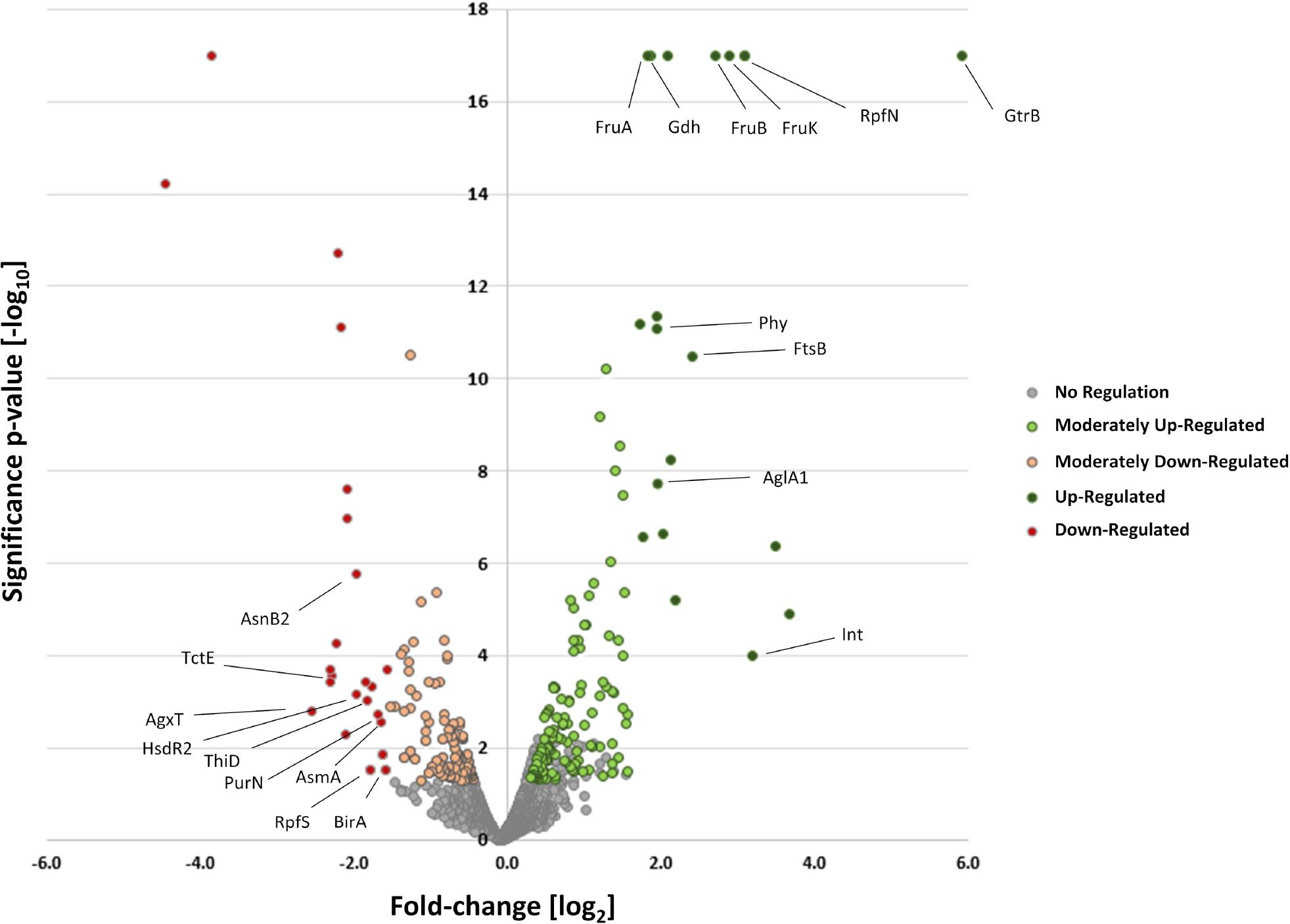
Fig. 5.
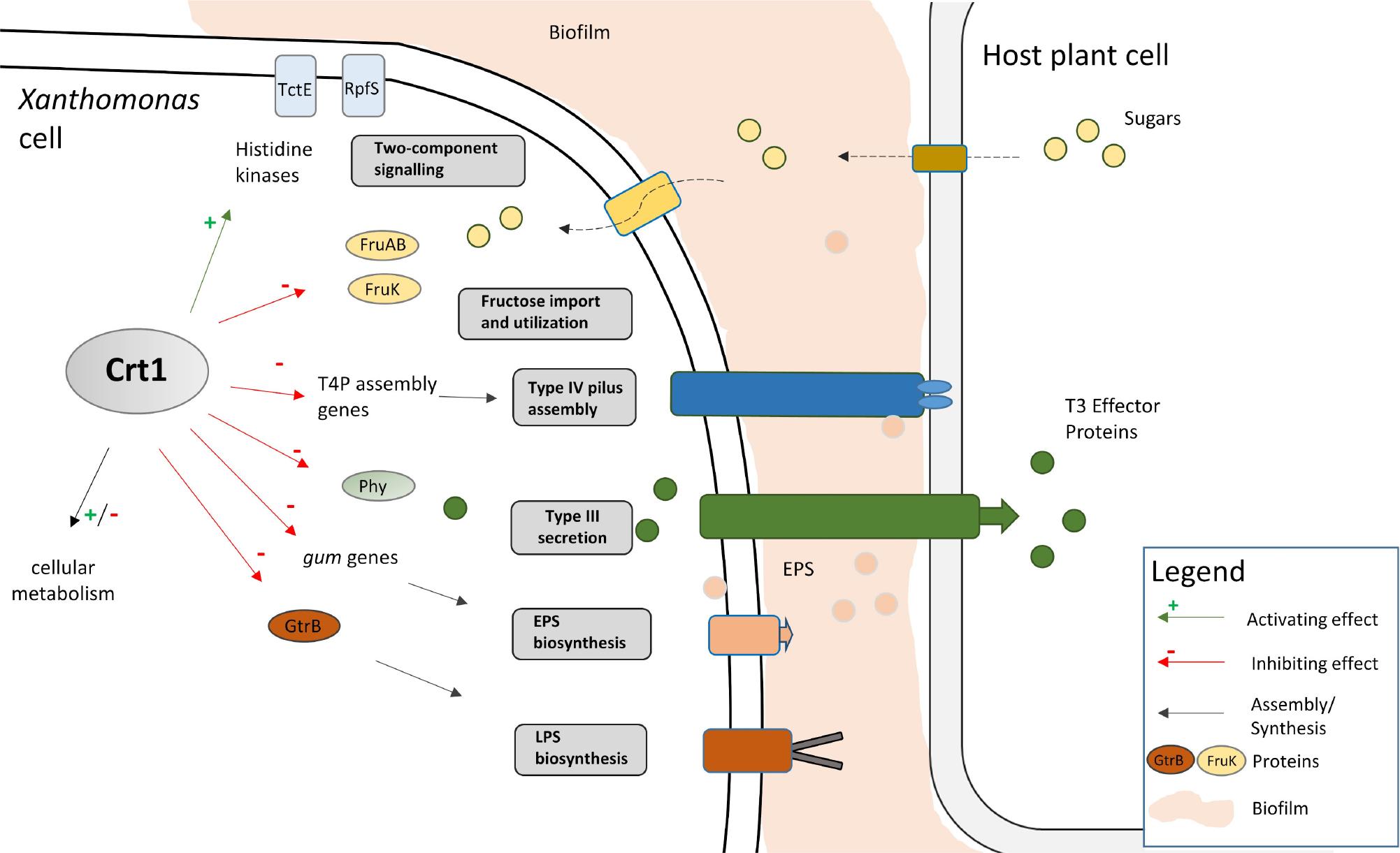
Effect of the crt1 deletion on the cytosolic and membrane-bound proteome of X_ campestris pv_ campestris during early stationary growth stage_
| Locus | Gene name | Function | Abundance ratio | p-Value | Unique peptides |
|---|---|---|---|---|---|
| Up-regulated proteins | |||||
| XCCB100_1291 | gtrB | putative bactoprenol glucosyltransferase | 5.92 | 1.00E-17 | 3 |
| XCCB100_3258 | conserved hypothetical protein | 3.67 | 1.30E-05 | 4 | |
| XCCB100_3905 | putative carboxypeptidase | 3.50 | 4.47E-07 | 3 | |
| XCCB100_2818 | int | phage-related integrase | 3.19 | 1.05E-04 | 3 |
| XCCB100_1799 | rpfN | carbohydrate-selective porin | 3.08 | 1.00E-17 | 20 |
| XCCB100_3640 | conserved hypothetical protein | 3.08 | 1.00E-17 | 4 | |
| XCCB100_1801 | fruK | 1-phosphofructokinase | 2.89 | 1.00E-17 | 7 |
| XCCB100_1802 | fruB | PTS fructose porter | 2.71 | 1.00E-17 | 35 |
| XCCB100_2558 | ftsB | septum formation initiator protein FtsB | 2.41 | 3.35E-11 | 3 |
| XCCB100_2909 | conserved hypothetical protein | 2.19 | 6.21E-06 | 3 | |
| XCCB100_1450 | pirin-related protein | 2.14 | 5.65E-09 | 7 | |
| XCCB100_0445 | hypothetical protein | 2.08 | 1.00E-17 | 4 | |
| XCCB100_2092 | transcriptional regulator; Crp/Fnr family | 2.02 | 2.26E-07 | 3 | |
| XCCB100_1788 | phy | putative exported phytase | 1.96 | 8.66E-12 | 3 |
| XCCB100_0128 | TonB-dependent outer membrane receptor precursor | 1.95 | 4.39E-12 | 18 | |
| XCCB100_1697 | aglA1 | alpha-glucosidase | 1.95 | 1.81E-08 | 11 |
| XCCB100_1807 | gdh | glutamate dehydrogenase | 1.87 | 1.00E-17 | 124 |
| XCCB100_1800 | fruA | PTS fructose porter | 1.82 | 1.00E-17 | 12 |
| XCCB100_1894 | conserved hypothetical protein | 1.77 | 2.80E-07 | 7 | |
| XCCB100_1761 | conserved hypothetical protein | 1.74 | 6.40E-12 | 3 | |
| Down-regulated proteins | |||||
| XCCB100_0308 | agxT | aminotransferase | -2.54 | 1.66E-03 | 4 |
| XCCB100_3852 | putative aromatic ring-opening dioxygenase | -2.31 | 2.09E-04 | 3 | |
| XCCB100_0796 | putative nuclease / phosphatase | -2.31 | 3.84E-04 | 3 | |
| XCCB100_0846 | tctE | two-component system sensor histidine kinase | -2.28 | 2.85E-04 | 5 |
| XCCB100_2019 | short chain dehydrogenase | -2.22 | 5.37E-05 | 6 | |
| XCCB100_3906 | putative transcriptional regulator; LysR family | -2.21 | 1.93E-13 | 6 | |
| XCCB100_4189 | conserved hypothetical protein | -2.17 | 7.94E-12 | 3 | |
| XCCB100_1528 | ABC transporter permease and ATP-binding protein | -2.09 | 5.28E-03 | 5 | |
| XCCB100_1398 | DJ-1/PfpI family protein | -2.09 | 1.12E-07 | 8 | |
| XCCB100_2784 | conserved hypothetical protein | -2.08 | 2.49E-08 | 7 | |
| XCCB100_3277 | hsdR2 | Type I site-specific deoxyribonuclease (restriction subunit) | -1.96 | 7.02E-04 | 8 |
| XCCB100_2866 | asnB2 | asparagine synthase (glutamine-hydrolysing) | -1.95 | 1.68E-06 | 10 |
| XCCB100_0546 | conserved hypothetical protein | -1.85 | 3.77E-04 | 3 | |
| XCCB100_2526 | thiD | phosphomethylpyrimidine kinase | -1.83 | 9.26E-04 | 6 |
| XCCB100_2607 | rpfS | DSF-sensing histidine kinase/response regulator hybrid | -1.79 | 3.06E-02 | 3 |
| XCCB100_0601 | membrane-located putative protein phosphatase | -1.76 | 4.57E-04 | 5 | |
| XCCB100_1293 | asmA | AsmA family membrane protein | -1.64 | 2.84E-03 | 7 |
| XCCB100_2313 | NDP-hexulose epimerase | -1.62 | 1.40E-02 | 3 | |
| XCCB100_4124 | birA | bifunctional biotin repressor / co-repressor biosynthesis enzyme | -1.58 | 3.02E-02 | 3 |
| XCCB100_2292 | short chain dehydrogenase | -1.56 | 1.98E-04 | 6 | |
Effect of the crt1 deletion on X_ campestris pv_ campestris gene expression_
| Locus | Gene name | Annotated gene product | M-value | A-value | p-Value |
|---|---|---|---|---|---|
| Up-regulated genes | |||||
| XCCB100_2325 | putative filamentous hemagglutinin-related protein | 1.22 | 11.4 | 7.42E-04 | |
| AS_XCCB100_1718 | antisense transcript gumH | 1.12 | 7.5 | 3.49E-03 | |
| XCCB100_0957 | pilP | type IV pilus assembly protein | 0.91 | 9.5 | 9.00E-04 |
| XCCB100_0907 | putative cytochrome c551 | 0.90 | 8.2 | 4.92E-03 | |
| XCCB100_0552 | groES | 10 kDa chaperonin | 0.90 | 10.2 | 1.99E-03 |
| XCCB100_0905 | putative exported protein | 0.89 | 11.6 | 2.65E-03 | |
| XCCB100_3396 | putative exported enzyme | 0.86 | 10.8 | 1.23E-09 | |
| XCCB100_1423 | murD | UDP-N-acetylmuramoylalanine-D-glutamate ligase | 0.84 | 9.0 | 3.95E-03 |
| XCCB100_1720 | gumJ | xanthan repeating unit exporter | 0.83 | 8.8 | 3.25E-02 |
| XCCB100_3538 | gfo | exported glucose-fructose oxidoreductase | 0.83 | 9.4 | 3.07E-02 |
| XCCB100_0906 | lao | L-amino-acid oxidase | 0.81 | 8.6 | 8.41E-03 |
| XCCB100_0207 | amtB | ammonium transporter | 0.80 | 12.2 | 6.28E-04 |
| XCCB100_2099 | hisH | glutamine amidotransferase | 0.77 | 8.9 | 7.83E-03 |
| XCCB100_0954 | pilM | type IV pilus assembly protein | 0.75 | 9.9 | 4.70E-03 |
| XCCB100_1209 | glk2 | glucokinase | 0.74 | 7.4 | 1.86E-02 |
| XCCB100_1615 | conserved hypothetical protein | 0.74 | 8.7 | 1.76E-02 | |
| XCCB100_2311 | hexose O-acetyltransferase | 0.73 | 8.4 | 2.52E-02 | |
| XCCB100_rna061 | putative regulatory sRNA | 0.73 | 19.7 | 1.86E-04 | |
| XCCB100_1719 | gumI | mannosyltransferase | 0.73 | 9.1 | 1.21E-02 |
| XCCB100_1672 | pilE | type IV pilus assembly protein PilE | 0.72 | 10.0 | 6.89E-03 |
| Down-regulated genes | |||||
| XCCB100_2267 | infA | translation initiation factor IF-1 | -1.50 | 11.5 | 1.11E-04 |
| XCCB100_2352 | hypothetical protein | -1.30 | 8.9 | 7.20E-04 | |
| XCCB100_1776 | hypothetical protein | -1.30 | 8.1 | 4.39E-02 | |
| XCCB100_1297 | homoserine O-acetyltransferase-like enzyme | -1.20 | 9.7 | 6.91E-06 | |
| XCCB100_2980 | conserved hypothetical protein | -1.13 | 12.6 | 7.39E-09 | |
| XCCB100_3868 | conserved hypothetical protein | -1.10 | 12.5 | 3.37E-04 | |
| XCCB100_1298 | cgl | cystathionine gamma-lyase | -1.09 | 11.5 | 1.50E-07 |
| XCCB100_4110 | hypothetical protein | -1.01 | 13.4 | 3.72E-03 | |
| XCCB100_3996 | conserved hypothetical protein | -1.00 | 10.4 | 1.31E-08 | |
| XCCB100_2290 | TonB-dependent outer membrane receptor precursor | -0.97 | 8.4 | 2.37E-03 | |
| XCCB100_rna193 | putative regulatory RNA | -0.96 | 10.2 | 2.16E-03 | |
| XCCB100_3017 | cspA2 | cold shock protein | -0.93 | 11.2 | 3.27E-06 |
| XCCB100_3810 | putative secreted protein | -0.91 | 12.4 | 8.14E-03 | |
| XCCB100_1741 | conserved hypothetical protein | -0.90 | 11.1 | 1.29E-02 | |
| XCCB100_3869 | putative manganese-containing catalase | -0.90 | 12.2 | 2.48E-03 | |
| XCCB100_3892 | conserved hypothetical protein | -0.87 | 10.8 | 8.28E-03 | |
| XCCB100_2320 | conserved hypothetical protein | -0.87 | 13.1 | 7.39E-03 | |
| XCCB100_2291 | hypothetical protein | -0.86 | 11.1 | 7.85E-03 | |
| XCCB100_1773 | hypothetical protein predicted | -0.86 | 6.6 | 1.02E-04 | |
| XCCB100_3966 | putative exported protein | -0.85 | 11.7 | 2.14E-10 | |
Effect of the regulator gene crt1 deletion on transcriptome and proteome of X_ campestris pv_ campestris_
| Locus | Gene name | Function | COG | Fold-change | Transcript/Protein abundance change | Studies in Xanthomonas |
|---|---|---|---|---|---|---|
| Cell motility | ||||||
| XCCB100_0954 | pilM | type IV pilus assembly protein | NU | 0.748 | T | Dugé de Bernonville et al, 2014 |
| XCCB100_0957 | pilP | Tfp pilus assembly protein | NU | 0.914 | T | Dugé de Bernonville et al, 2014 |
| XCCB100_1672 | pilE | Tfp pilus assembly protein | NU | 0.725 | T | Dugé de Bernonville et al, 2014 |
| Carbohydrate transport and metabolism | ||||||
| XCCB100_1697 | aglA1 | alpha-glucosidase | G | 1.950 | P | Blanvillain et al, 2007; Fatima and Kumar, 2015; Eom et al. 2015; Ruan et al. 2014; Watt et al. 2009 |
| XCCB100_1799 | rpfN | carbohydrate-selective porin | G | 3.080 | P | Moreira et al. 2004 |
| XCCB100_1800 | fruA | PTS fructose porter | G | 1.820 | P | Moreira et al. 2004 |
| XCCB100_1801 | fruK | 1-phosphofructokinase | G | 2.890 | P | Li et al. 2019 |
| XCCB100_1802 | fruB | PTS fructose porter | G | -0.507 / 2.71 | T/P | Moreira et al. 2004 |
| Signal transduction mechanisms and information processing | ||||||
| XCCB100_0846 | tctE | two-component system sensor histidine kinase | T | -2.280 | P | Timilsina et al. 2020; Wang et al. 2016; Wang et al. 2017b |
| XCCB100_2607 | rpfS | DSF-sensing histidine kinase/response regulator | T | -1.790 | P | An et al. 2014; O’Connell et al. 2013; Ryan et al. 2007 |
| Amino acid transport and metabolism | ||||||
| XCCB100_1297 | homoserine O-acetyltransferase-like enzyme | -1.204 | T | Pielken et al. 1987; Pielken et al. 1988; O’Connell et al. 2013 | ||
| XCCB100_1298 | cgl | cystathionine gamma-lyase | -1.089 | T | Pielken et al. 1987; Pielken et al. 1988; O’Connell et al. 2013 | |
| XCCB100_2099 | hisH | glutamine amidotransferase | 0.766 | T | Li et al. 2019 | |
| XCCB100_2866 | asnB2 | asparagine synthase (glutamine-hydrolysing) | E | -1.950 | P | Qian et al. 2013 |
| EPS biosynthesis | ||||||
| AS_ XCCB100_1718** | antisense transcript gumH | 1.118 | T | Chan et al. 1999; Kiraly et al. 1997; Vojnov et al. 2001; Rigano et al. 2007; Alvarez et al. 2000; Yun et al. 2006 | ||
| XCCB100_1719 | gumI | mannosyltransferase | R | 0.726 | T | Chan et al. 1999, Kiraly et al. 1997; Vojnov et al. 2001; Rigano et al. 2007; Alvarez et al. 2000; Yun et al. 2006 |
| XCCB100_1720 | gumJ | xanthan repeating unit exporter | M | 0.833 | T | Chan et al. 1999, Kiraly et al. 1997; Vojnov et al. 2001; Rigano et al. 2007; Alvarez et al. 2000; Yun et al. 2006 |
| Lipid transport and metabolism | ||||||
| XCCB100_1788 | phy | putative exported phytase | I | 1.960 | P | Raboy 2003; Blüher et al, 2017; Chatterjee et al. 2003 |
| Cell wall, membrane, envelope biogenesis | ||||||
| XCCB100_1291 | gtrB | putative bactoprenol glucosyl-transferase | 5.92 | P | Newman et al. 2001; Vorhölter et al. 2001; Li et al. 2019 | |
| Other functions | ||||||
| XCCB100_2558 | ftsB | septum formation initiator protein FtsB | D | 2.41 | P | Ferreira et al. 2017 |