Xanthomonas is a genus of Gram-negative γ-proteobacterium, which poses a tremendous threat to a wide range of economically important crops and plants (Bradbury 1986; Bonas et al. 2000; Chan and Goodwin 1999; Williams 1980). The genus comprises 27 species, each with numerous highly host-plant specific pathovars (Burkholder et al. 1957; Jun et al. 2010; Parkinson et al. 2007). Moreover, more than 100 pathovars display tissue specificity upon infection of the host plant (Ryan et al. 2011). Based on recent phylo-taxonogenomic investigations, the taxonomic boundaries of the genus have been revised, revealing that several closely related genera, including Stenotrophomonas, Pseudoxanthomonas, and Xylella, should be reclassified under Xanthomonas (Bansal et al. 2023).
Belonging to the most analyzed species of the genus, Xanthomonas campestris pv. campestris is the causal agent of black rot disease in Brassica oleracea (B. oleracea), resulting in substantial losses in yield and produce quality (He and Zhang 2008; Onsando 1992; Vicente and Holub 2013). As a vascular pathogen, X. campestris pv. campestris initially colonizes the xylem vessels (Ryan et al. 2011). The most typical symptom of the black rot disease is the browning of the leaf midribs and the formation of a V-shaped lesion on the leaf with both vertices directed towards the middle vein. With the progression of the infection, the vein darkens due to the bacterial infection within the vascular system, ultimately leading to necrosis and falling of the affected leaf (Vicente and Holub, 2013).
Xanthomonas lives part of its life cycle saprophytically outside the host plant, for instance, freely in the soil or on plant tissue debris. The life cycle during infection can be generally divided into the epiphytic and endophytic stages. During the epiphytic stage, bacteria are introduced into the host tissue. Upon infiltrating the host plant, X. campestris pv. campestris enters the endophytic stage. Once a high population density is reached, X. campestris pv. campestris re-emerges on the leaf surface and can be transmitted to a new host (Cheng et al. 2019; Timilsina et al. 2020). At each stage, access to nutrients changes, as well as the general environmental conditions to which X. campestris pv. campestris is exposed, differ. Therefore, different regulatory cues and adaptation mechanisms must be employed depending on the current lifestyle.
The bacterial ability to successfully infect a host plant and to multiply efficiently within the invaded tissue depends on a variety of virulence factors and secretion systems that are crucial for bacterial motility, adhesion to biotic and abiotic surfaces, nutrition, stress resistance, and ultimately the suppression of the plant immune response upon infection (Büttner and Bonas, 2010; Dow et al. 2003). Despite the fact that the rich repertoire of virulence factors in Xanthomonas has been studied, the regulatory cues and proteins that control gene transcription in X. campestris pv. campestris still need to be investigated more thoroughly.
To identify novel putative transcriptional regulators that contribute to bacterial infection and growth in planta, Leßmeier (et al. 2016) conducted a pull-down approach based on DNA affinity chromatography. For this purpose, X. campestris pv. campestris cultures were grown on two different minimal media supplemented with either glucose or sucrose. The latter officiated to mimic the conditions during infection by X. campestris pv. campestris, as sucrose is the main carbon source for the pathogen in planta. Among the identified proteins that bound to the promoters of sucrose-related target genes, a small number were annotated as transcriptional regulators. The previously known regulators SuxR and cAMP receptor-like protein (Clp) were identified as sucrose-related. The former was confirmed to control the transcription of the sucrose transport and utilization genes in its genomic proximity (Blainvillain et al. 2007), while the latter was found bound to the putative promoter region of the suxR gene. This finding linked the important Quorum Sensing (QS) regulator Clp to the sucrose scavenging regulator SuxR (Leßmeier et al. 2016). Furthermore, two proteins, encoded by the previously uncharacterized genes xccb100_2791 and xccb100_2861, showed affinity for the putative promoter regions of the sucrose-related target genes. Both are predicted to be members of the LysR family of transcriptional regulators. The LysR-type transcriptional regulators (LTTRs) are the most abundant among DNA-binding proteins in prokaryotes, controlling a broad spectrum of genes, and thus are implicated in regulation of virulence, QS, motility or amino acid and carbohydrate metabolisms (Deziel et al. 2005; Hartmann et al. 2013; Heroven and Dersch 2006; Minezaki et al. 2006; O’Grady et al. 2011; Park et al. 2020; Rashid et al. 2016; Russel et al. 2004; Schulte et al. 2019). Members of this family share a highly conserved protein structure, with an N-terminal helix-turn-helix (HTH) DNA-binding domain and a C-terminal substrate-binding domain (Maddocks and Oyston, 2008). The DNA-binding putative regulator Xccb100_2791 was determined to bind promoter regions of the suxA/suxC, pel3, pglA1, and suxR genes (Leßmeier et al. 2016). It is encoded by a gene located between a putative oligosaccharide ABC exporter gene and an inner-membrane ABC exporter gene.
A recent study investigated the transcription pattern of transcriptional regulators in X. campestris pv. campestris during the nitrogen starvation induced stationary growth stage (Alkhateeb et al. 2018). The analysis revealed that three LTTRs (ampR, xccb100_1534, and xccb100_2665) were up-regulated in their transcription upon entry in the stationary growth stage, whereas the transcription of the xccb100_2791 gene was down-regulated, indicating its involvement in environmental sensing (Alkhateeb et al. 2018).
In this study, we introduce XCCB100_2791 (hereafter named carbohydrate-related transcriptional regulator Crt1, due to its affinity to sucrose-related target genes) as a novel transcriptional regulator and present a comprehensive characterization in terms of its poignant phenotypic features, virulence efficiency, repertoire of differentially expressed target genes, as well as abundance alterations within the cytosolic and membrane-bound proteome. We investigated whether and how LTTR Crt1 modulates the transcription of target genes and affects the abundance of proteins involved in the adaptation to the virulent lifestyle of X. campestris pv. campestris. For this purpose, a deletion of the crt1 gene was constructed. The mutant was characterized with respect to phenotype and pathogenicity. Furthermore, a differential analysis was carried out between the afore-mentioned strain and the wild-type strain Xcc B100, both on the transcriptome and proteome level.
Unless otherwise specified, all molecular biology procedures, including chromosomal DNA purification, DNA amplification, and plasmid transformation, were performed according to standard protocols described in Green and Sambrook (2012). The X. campestris pv. campestris B100 wild-type strain was used as a parental strain for the generation of the deletion mutant and is available from the National Collection of Plant Pathogenic Bacteria (NCPPB) under the accession number NCPPB 4615 (Alkhateeb et al. 2018). The deletion mutant was constructed by disrupting the crt1 gene utilizing the vector pk18mobsacB as the backbone of the cloning vector (Schäfer et al. 1994). Two fragments flanking upstream and downstream of the excision site were amplified by PCR from chromosomal DNA of X. campestris pv. campestris B100 using primers designed according to Gibson assembly cloning method (Gibson et al. 2009). All primers used for this study are listed in Supplementary Table II. Both amplified flanking regions were cloned into the pK18mobsacB vector in one step Gibson assembly procedure. The cloning vector was transformed into the shuttle host, Escherichia coli Dh5α. The isolated cloning vector was then transformed into electrocompetent X. campestris pv. campestris B100 cells via electroporation. Kanamycin-resistant X. campestris pv. campestris B100 colonies that contained the integrated plasmid were selected. Bacterial colonies were transferred in parallel to selective and non-selective media. Double cross-over occurring at both flanking regions led to the directed excision of a 91 nt long sequence within the crt1 gene, causing a frameshift and disrupting the correct folding of the regulator protein. Deletion mutants were selected by colony PCR using the primers as listed in Supplementary Table II. The successful and accurate excision was validated with Sanger sequencing and MiSeq sequencing (Illumina, USA). The complemented mutant was constructed via plasmid-based complementation. The broad-host-range vector pBBR1-MCS5 carrying a gentamycin cassette was used as the backbone, and the full-length crt1 gene, including its upstream regulatory region, was cloned into the vector using a onestep Gibson assembly procedure. The resulting plasmid was introduced into X. campestris pv. campestris B100 Δcrt1 by electroporation, and correct integration was confirmed by Sanger sequencing of the positive clone. All sequencing approaches were carried out by the inhouse sequencing core facility. The detection of SNP was carried out with analysis tool ReadXplorer2 (Hilker et al. 2016).
All bacterial strains and vectors constructed and applied in this work are listed in Supplementary Table II. All cultivations were performed in a parallel shaking flask at 180 rpm and 28°C. Pre-cultures of X. campestris pv. campestris were grown in TY rich media (5 g/l tryptone, 3 g/l yeast extract, 0.7 g/l CaCl2) (Beringer 1974), intermediate and main cultures in minimal XMD media (Schatschneider et al. 2013), supplemented with 10 g/l glucose as sole carbon source and 0.6 g/l KNO3 as nitrate source. Antibiotics were added to the media with respective concentrations: streptomycin 800 μg/ml, kanamycin 50 μg/ml, and gentamicin 30 μg/ml.
Xanthan quantification was performed according to Palaniraj et al. (2011). For this purpose, X. campestris pv. campestris main cultures were grown in minimal XMD media as described above and harvested after 72 h of cultivation. Cells were diluted 1:8 (v/v) in pre-cooled water and subsequently centrifuged for 1 h at 12,500 rpm and 4°C. Afterwards, the supernatant was separated thoroughly from the pellet and added to pre-cooled absolute isopropanol 1:3 v/v. After 30 min of incubation on ice, the sample was centrifuged for 90 min at 9,000 rpm at 4°C. The resulting supernatant was carefully discarded, and the xanthan pellet was dried for 72 h at 60°C and subsequently weighed. For each strain, 5 biological replicates have been considered. For the assessment of statistical significance, an equal-variance t-test was performed.
Viscosity of xanthan harvested from X. campestris pv. campestris cultures was measured utilizing a rotational viscometer (Physica MCR 101, Anton Paar, Austria). For this purpose, dried EPS was diluted to 1% w/v in sterile water. For each measurement, 1.5 ml of the sample was placed on the stationary plate. The measurements were carried out at room temperature, and all calculations were performed with RheoPlus Rheometer Software (Anton Paar, Austria). For each strain, 4 biological replicates have been considered. To investigate the statistical significance of the obtained results, a paired two-sample t-test was performed.
Three independent biological replicates of each strain were cultivated in minimal XMD medium with 10 g/l glucose and harvested at the early stationary growth stage, with an OD600 nm of 1.8. Cultivation of X. campestris pv. campestris in sucrose leads to overproduction of EPS during the early stationary stage, hence preventing to obtain material of high quality and purification grade without distorting the results of the whole-genome analysis.
For the purpose of isolation of total RNA, cells were harvested by centrifugation at 14,000 rpm for 4 minutes. The resulting cell pellet was frozen immediately in liquid nitrogen and stored at -80°C until further treatment. Frozen cell pellets were suspended carefully in 2 ml DNA/RNA Shield provided with the Quick RNA Miniprep Kit (ZymoResearch, USA), and 750 μl of the suspension was transferred to Lysing Matrix Tubes (MP Biomedical, USA), containing ceramic and glass beads, for homogenization. Cells were disrupted by the Ribolyzer (Hybaid, Germany) thrice for 40 sec at speed setting 6.5, and 2-min cooling on ice in between the homogenization steps. After the centrifugation for 5 min at 4°C and 13,500 rpm, supernatant was transferred into new RNAse-free reaction tubes. All subsequent steps of RNA isolation and DNA digestion were carried out with Quick RNA Miniprep Kit and RNA Clean & Concentrator (ZymoResearch, USA) according to the manufacturer’s protocol. To verify complete removal of residual DNA, PCR with primers binding to multiple genomic regions was carried out. Quantity of the RNA was measured with a NanoDrop 1000 ND-1000 spectrophotometer (Thermo Fisher, USA).
All the subsequent experimental procedures, including cDNA synthesis, labelling of cDNA, hybridization, and washing of microarrays were performed according to the protocol Two-Color Microarray-Based Prokaryote Analysis FairPlay III Labeling (Agilent Technologies, USA). Any adjustments to the afore-mentioned protocol are described below. To synthesize cDNA for each strain, three biological replicates were pooled in equal amounts to 5 μg of total RNA, and AffinityScript HC Reverse Transcriptase and aminoallyl dUTPs were used. Purified cDNA was labeled with Cy3- or Cy5-N-hydroxysuccinimidyl ester mono-reactive dyes (GE Healthcare, UK). Subsequent quantification of labelled cDNA, as well as specific activity of mono-reactive dyes, was performed with NanoDrop ND-1000 spectrophotometer (Thermo Fisher, USA) with the method ssDNA-33. Only samples with specific activity of at least 40 pmol of mono-reactive dye per μg of cDNA were considered for the subsequent hybridization.
For the purpose of this study, custom whole-genome oligonucleotide microarrays representing the coding sequence of X. campestris pv. campestris B100 were designed with Agilent eArray and ordered in the 8x15K format (both Agilent Technologies, USA). Each array consists of 4,679 features representing genes and putative small RNAs of Xanthomonas campestris pv. campestris B100. The complete design and dataset of the microarray have been deposited at EMBL Array-Express Archive for Functional Genomics, accession number E-MTAB-12383. For each microarray the mix was prepared with 330 ng of Cy3-labeled cDNA, 330 ng of Cy5-labeled cDNA, and 11 μl gene expression blocking agent. The cDNA blocking mix was filled up to 55 μl with H20 and joined with 55 μl Hi-RPM hybridization buffer. The quantities were adjusted to fit the 8x15K format of the array. For each microarray, 100 μl of the hybridization mix was used. To reduce artefact formation, the washing procedure was modified by increasing the number of wash cycles (both 5 minutes in buffer 1 and 1 minute in buffer 2, as described in the manufacturer’s protocol). All procedures, reagents, and equipment for microarray hybridization and scanning were ordered and used as described in the manufacturer’s protocol referenced above.
Feature extraction was performed according to the manufacturer’s protocol with the Agilent Feature Extraction Software (Agilent Technologies, USA), and the obtained data were further analysed with the web-based software EMMA2 (Dondrup et al. 2009). For this study, data from two arrays, resulting from a two-colour dye-swap approach, were combined to obtain more reliable results. The data were normalized using the LOWESS method, and a subsequent one-sample Holm t-test was performed. Transcripts with the logarithmic signal intensities (A-value) ≥ 8.0, mean values of transcriptome fold-change [log2] (M-value) ≥ 1 or ≤ -1, and the p-value ≤ 0.05, were considered as significantly differentially regulated.
In order to investigate the cytosolic proteome, both X. campestris pv. campestris strains were cultivated as described above for the transcriptome analysis. Sample preparation, tryptic digestion, and SEP Pak purification were carried out according to Wang [et al. 2005] and Russell [et al. 2001]. X. campestris pv. campestris cells were first diluted 1:2 v/v in 0.9% NaCl and promptly centrifuged at 4°C and 13,500 rpm for 2 minutes. Afterwards, the supernatant was completely discarded and the remaining pellets, consisting of cytosolic and periplasmic proteins, were frozen in liquid nitrogen. Frozen cell pellets were lyophilized in order to dispose of the water molecules that would interfere with the sample purification and final measurements. To each lyophilized sample, 100 μl of 100mM ammonium bicarbonate (Ambic), 100 μl 2,2,2-trifluoroethanol (TFE), and 5 μl 200 mM Tris (2-carboxyethyl) phosphine (TCEP) were added. During the incubation at 60°C for 60 minutes, samples were occasionally inverted. Subsequently, 20 μl of 400 mM chloroacetamide was added to each sample, and the samples were further incubated at room temperature in the dark for 90 minutes. Afterwards, 5 μl of TCEP was added to each sample, and the samples were incubated at room temperature for 60 minutes. In the next step, 437 μl Ambic and 437 μl ddH2O were added. Finally, 2 μl of 1 μg μl-1, Trypsin Gold (Mass Spectrometry Grade, Promega, USA) was added and the samples were incubated overnight at room temperature.
After digestion, samples were purified using Sep-Pak C18 Columns (Waters Corporation, USA) as described earlier (Droste et al. 2020). Afterwards, the eluted samples were dried in a vacufuge concentrator (Eppendorf, Germany) and rehydrated with 12 μl Solution A (98% Acetonitril, 2% ddH20, 0.01% TFA). Rehydrated samples were centrifuged at 13,500 rpm for 2 minutes. Protein concentration and purity of the samples were assessed with NanoDrop 2000/2000c (Thermo Fisher, USA) at the wavelength of 280 nm.
Mass spectrometry was performed on a Q Exactive mass spectrometer (Thermo Fisher, USA). The peptides were separated on a 25 cm steel column Acclaim™ PepMap™ C18-LC-column with a particle size of 2 μm and a diameter of 75 μm (Thermo Fisher, USA). For data analysis, ms spectra were searched with SEQUEST against the target-decoy protein sequence database of X. campestris pv. campestris B100. Protein abundances were determined with the Proteome Discoverer 2.3 (Thermo Fisher, USA). Only unique peptides were used for the quantification. An oxidation of methionine (15.99 Da) was allowed up to three times per peptide. As a static modification, a carbamidomethylation of cysteine (57.02 Da) and the dynamic modification of the N-terminus with an acetylation (42.01 Da) were allowed. The mass spectrometry proteomics data have been deposited in the MassIVE via the PRIDE partner depository with the data set identifier (Identifier for Review Process: Dataset MSV000090790).
Brassica oleracea ‘express’ cultivar (Quedlinburger Saatgut, Germany) was used as a host plant for X. campestris pv. campestris strains. All plants were grown on soil in a humidified chamber at 23°C in incandescent light with a 16 h light/8 h dark cycle. X. campestris pv. campestris strains were grown overnight at 28°C on agar plates in TY rich medium supplemented with antibiotic streptomycin 800 μg/ml. The leaves of 4-week-old seedlings were inoculated by infiltrating a single colony of the bacterial strain using a sterile needle. At least 11 plants were infiltrated by either of the X. campestris pv. campestris strains. Additionally, mock inoculation was performed on 5 plants by infiltration with a sterile syringe only in order to exclude mechanical stress upon leaf tissue as the potential cause of occurring disease symptoms. The infection symptoms were observed and photographed every 48 h after inoculation. Both lesion area and entire leaf area at 10 dpi were quantified using FIJI software (Schindelin et al. 2012). Disease was rated by the percentage of the entire leaf area affected by the bacterial strain. For each strain, ≥ 11 replicates have been assessed, and a two-tailed t-test (p < 0.01) was performed for both sample groups.
Gene crt1 is located at the position 3207118 to 3208017 in the genome of X. campestris pv. campestris B100 (Supplementary Fig. 1). It is proposed to be organized as a gene downstream of the gene msbA4 within a single operon, based on RNA sequencing data, partially presented in the genome-wide study of transcription start sites (Alkhateeb et al. 2016). The genes in its immediate vicinity encode a putative lipooligosaccharide ABC exporter and an inner membrane ABC efflux protein (msbA4 and xccb100_2792, respectively), as shown in Supplementary Fig. 2. Investigation of the coding sequence of the ORF xccb100_2791 provided the first insight into the putative role of the newly identified regulator Crt1. The gene product Xccb100_2791 comprises 299 amino acid residues with a predicted molecular mass of 32 kDa. It contains two protein domains, a winged HTH DNA-binding domain (amino acids 8 to 66), and a LysR substrate-binding domain (amino acids 93 to 298). A more detailed analysis based on SWISS-MODEL revealed potential homo-dimer and homo-tetramer formation of the Crt1 protein, similar to that of another LysR-family regulator, OxyR (Bertoni et al. 2017; Guex et al. 2009; Waterhouse et al. 2018).
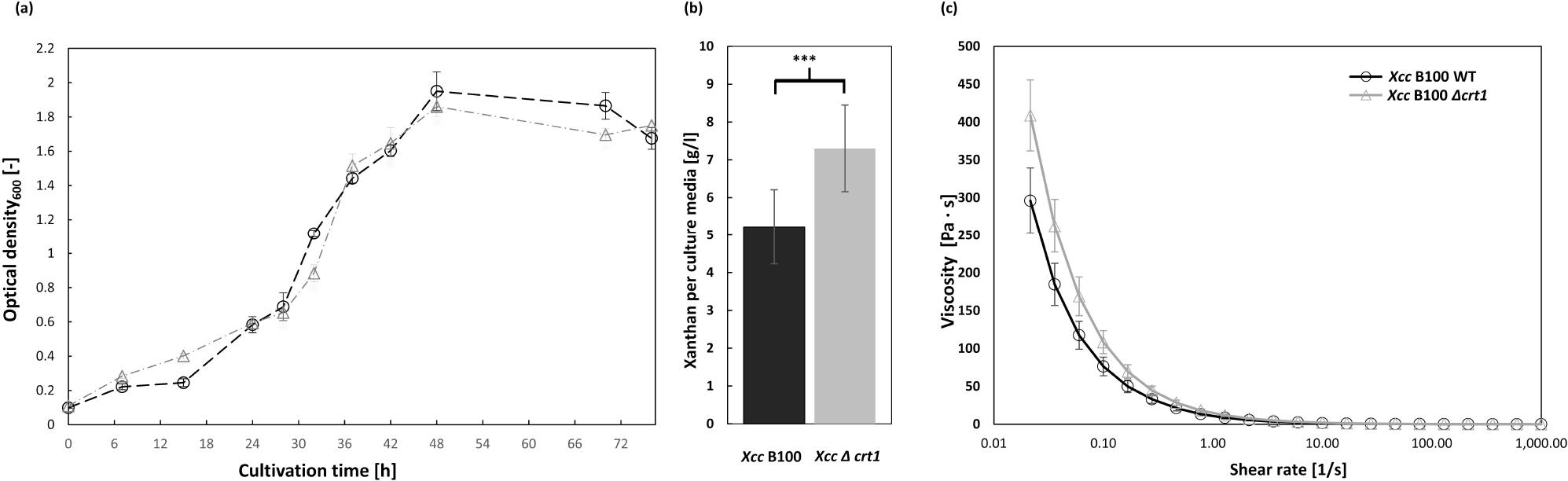
Phenotype characterization of the deletion mutant Xanthomonas campestris pv. campestris B100 Δcrt1 compared to the wild-type strain Xanthomonas campestris pv. campestris B100.
a) Growth behaviour of the wild-type (○) and the deletion mutant (Δ) strains cultivated in XMD minimal medium with 10 g/l glucose as sole carbon source; b) xanthan production was measured after 72 h of cultivation; xanthan was precipitated with isopropanol, dried and weighted relative to culture media; n≥ 5 biological replicates were investigated; two-tailed t-test resulted in significant p-value of 0.027 (***); viscosity of xanthan was assessed for both strains; xanthan was harvested after 72 h of cultivation, isolated and dried; all measurements were performed with 1% w/v xanthan with rheometer at room temperature; n = 4 biological replicates were investigated for each strain.
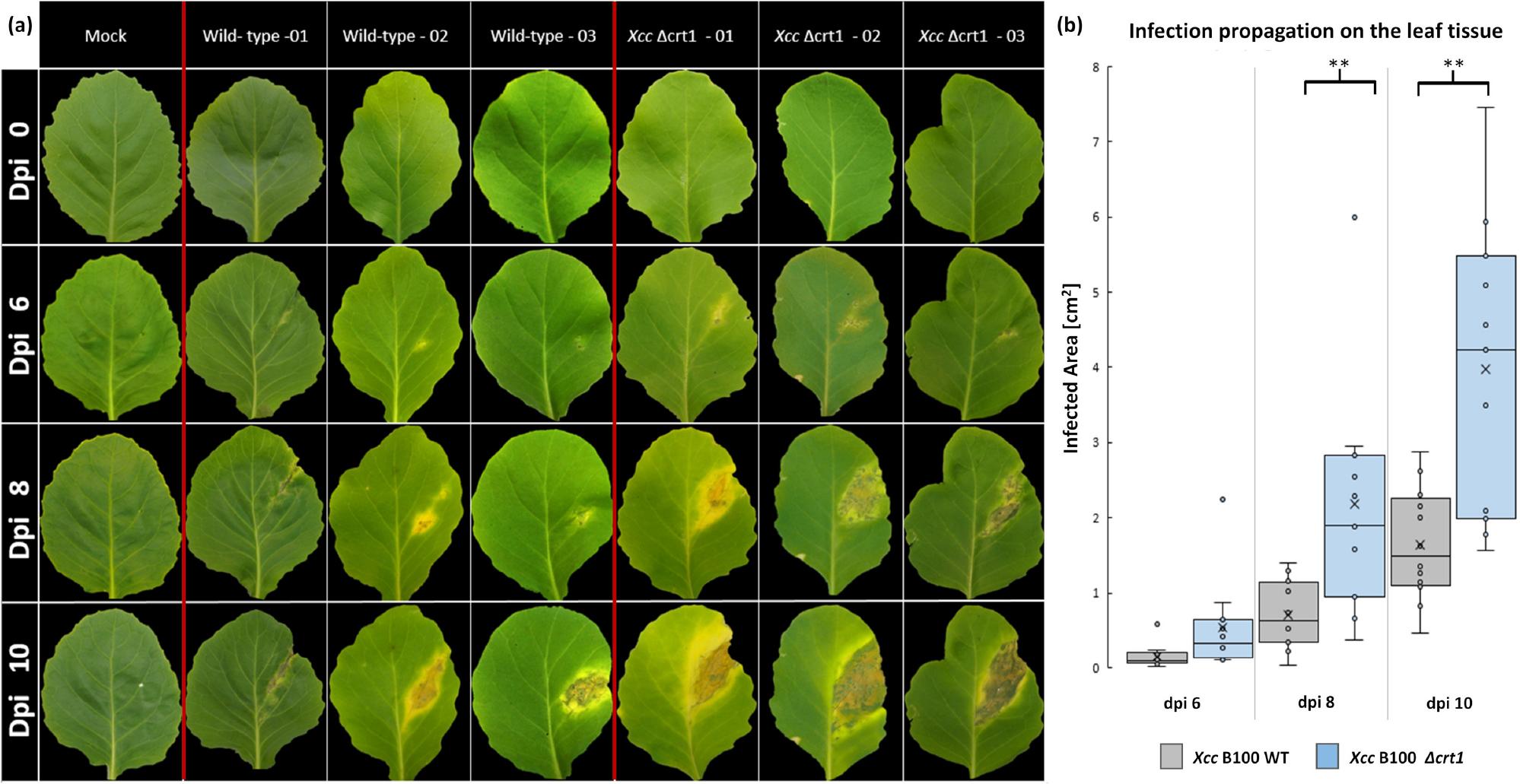
Bacterial virulence against host plant Brassica oleracea.
a)Leaf lesions in Brassica oleracea caused 6,8, and 10 dpi by infection with the X. campestris pv. campestris B100 wild-type (WT), deletion mutant strain X. campestris pv. campestris B100 Δcrt1, and a mock sample; inoculation was conducted with a sterile syringe, infection progress was documented at 6,8, and 10 days post inoculation; a representative set of leaves was chosen for the depiction of symptom propagation; b) analysis of the lesion area of the plants infected with either wild-type or deletion mutant strain, measured at 0, 6, 8, and 10 days post inoculation; depicted are calculated mean values for each condition; statistical relevance was confirmed by t-test (Supplementary Table I), for each bacterial strain n ≥11 samples were considered.
In order to confirm Crt1 as a transcriptional regulator and to gain a first insight into the network of genes under its control, the crt1 gene in the wildtype strain X. campestris pv. campestris B100 was disrupted by a non-in-frame deletion. For this purpose, regions flanking the deletion site upstream and downstream were cloned into the pK18mobsacB vector, and the region of interest was excised from the genome via double cross-over. Successful excision of the fragment within the crt1 coding sequence was confirmed by whole-genome sequencing, which was carried out to identify any single-nucleotide polymorphism (SNP) divergence between the wildtype strain X. campestris pv. campestris B100 and the novel deletion mutant X. campestris pv. campestris B100 Δcrt1. In addition to confirming the directed deletion within the crt1 coding sequence, two SNPs were determined within the coding sequences of fhuA and nuoN. In both cases, the SNPs result in an amino acid exchange: however, in-depth in silico analysis showed no effect on the structure or functionality of the proteins encoded by fhuA and nuoN. For this purpose, the open-source toolboxes within RaptorX and SWISS-MODEL were used (Bertoni et al. 2017; Guex et al. 2009; Wang et al. 2017b; Wang et al. 2018; Waterhouse et al. 2018; Xu et al. 2021). Furthermore, both genes, fhuA and nuoN, and the proteins encoded by them were not identified in the differential analysis of the transcriptome and proteome, respectively, among those with the most significant fold changes.
To characterize the phenotype of the obtained deletion mutant X. campestris pv. campestris B100 Δcrt1, first, its growth was compared with that of the ancestral wild-type strain in parallel shaking flask cultivation with minimal XMD medium supplemented with 10 g/l glucose as the sole carbon source. Disruption of the crt1 gene did not influence the growth, as shown in Fig. 1a. Thus, the transcriptional regulator is not implied to control the growth of the X. campestris pv. campestris at any growth stage under laboratory conditions in minimal medium. However, the measured xanthan output was increased by 39.8% in the X. campestris pv. campestris B100 Δcrt1 culture to 7.3 g/l culture compared to 5.22 g/l of EPS measured for wild-type X. campestris pv. campestris B100 (Fig. 1b).
The significance of the measured difference was confirmed by an equal-variance t-test, yielding a twotailed p-value of 0.011. For each condition, at least 5 biological replicates were considered. Similarly, the detected viscosity of the mutant strain was significantly higher than that of the wild-type strain (Fig. 1c). For each strain, xanthan obtained from four biological replicates has been used, and the statistical analysis using a paired two-sample, two-tailed t-test for means led to a significant p-value of 0.027. Furthermore, EPS quantification was carried out for wild-type, deletion mutant and complemented mutant strains X. campestris pv. campestris B100 Δcrt1::crt1. The xanthan output was measured as 3.92 g/l, 4.99 g/l, and 4.06 g/l of the culture, respectively. The significance between the measurements of the deletion mutant, and both wildtype and complemented mutant was confirmed with a two-tailed t-test with p-values of 0.028 and 0.035, respectively. Two-tailed t-test demonstrated no statistical significance between wild-type X. campestris pv. campestris B100 and the complemented mutant strain (Supplementary Fig. 3).
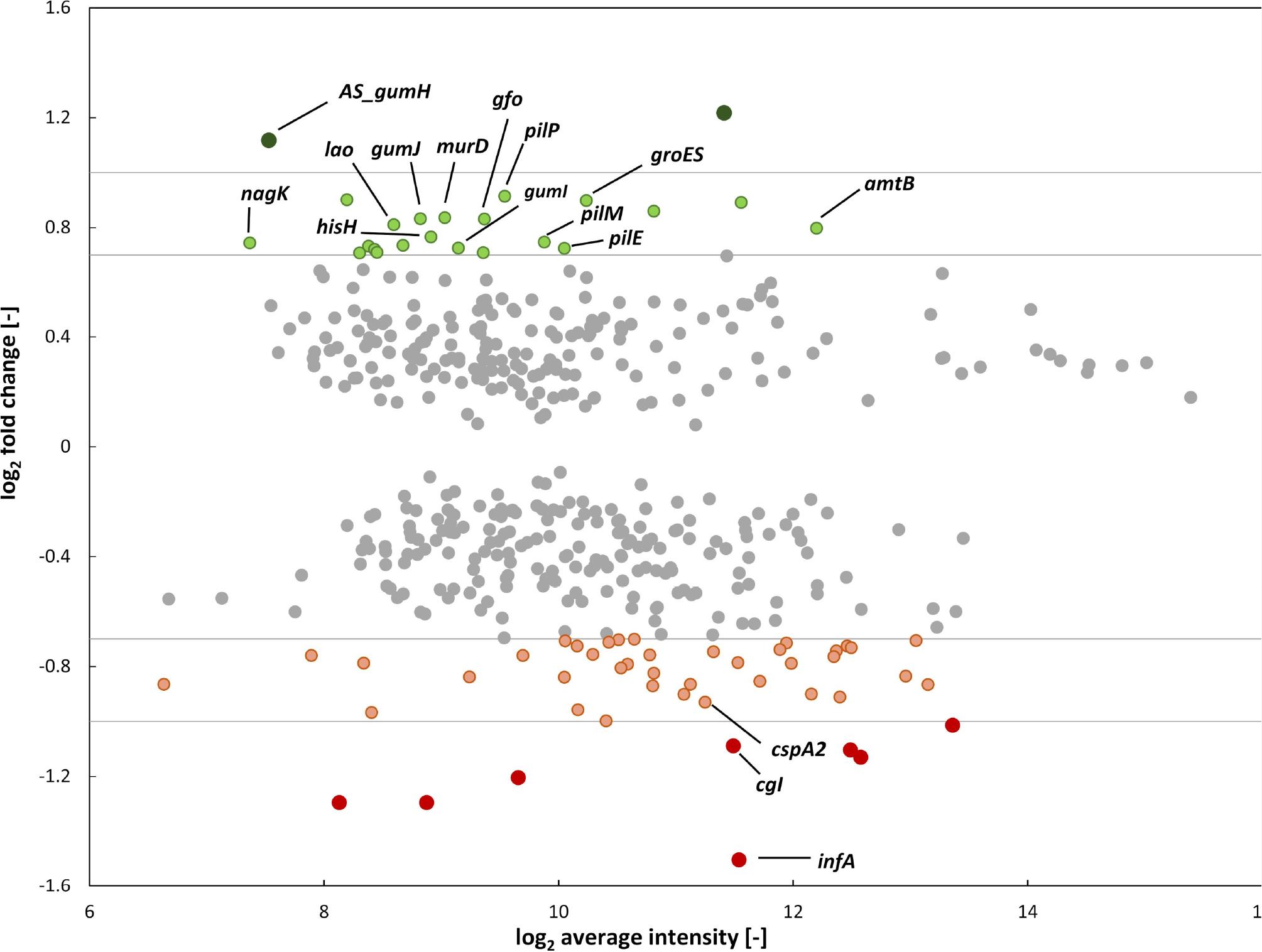
Genome-wide differential transcriptional analysis of the deletion mutant Xanthomonas campestris pv. campestris B100 Δcrt1 compared to the wild-type strain.
Ratio/intensity plot obtained from whole genome microarrays of the strain X. campestris pv. campestris B100 Δcrt1 compared to the wildtype strain X. campestris pv. campestris B100; cultivation was performed in the minimal XMD medium with 10 g/l glucose as a sole carbon source; cells were harvested at an early stationary growth stage at an oD600 nm of 1.8; shown are only targets with confidence level greater than 95%; green and red dots represent target genes with increased and decreased, respectively, transcript levels in the mutant strain X. campestris pv. campestris B100 Δcrt1; genes with transcription unaffected by the deletion of crt1 gene are represented by grey dots; annotated target genes with highest fold-change compared to the wild-type strain marked by gene name.
X. campestris pv. campestris synthesises xanthan under nitrogen limitation during the stationary growth stage, thus the increase in the EPS output has no effect on the biomass formation during exponential bacterial growth. The role of EPS xanthan has been suggested to be beneficial for Xanthomonas during the colonization of host plant tissue (Vojnov et al. 2000; Alvarez et al. 2000; Yun et al. 2006). Other studies have suggested that xanthan biosynthesis is modulated by components of Quorum Sensing (Dow et al. 2003; Crossman and Dow 2004). In light of these findings, the increased xanthan production in the mutant strain may improve the protection of X. campestris pv. campestris against adverse environmental conditions, including plant defense mechanisms.
The putative transcriptional regulator Crt1 was identified in an approach to screen for sucrose-related regulatory proteins and showed affinity for the promoter regions of target genes differentially expressed under sucrose supplementation of the growth media (Leßmeier et al. 2016). As sucrose is abundant in the plant tissues, it serves as a major carbon source for the Xanthomonas scavenged from the host cell. Proteins that bind to the promoters of the sucrose-related target genes are therefore implied to play a role in the pathogenic behaviour of X. campestris pv. campestris.
To investigate the influence Crt1 exerts on bacterial virulence regulation, we conducted in planta experiments. For this purpose, virulence assays were employed to test the effect of both the wildtype strain X. campestris pv. campestris B100 and the deletion mutant X. campestris pv. campestris B100 Δcrt1 on the host plant Brassica oleracea (B. oleracea). Mock inoculation was performed to exclude mechanical stress on the leaf material as a potential cause of the observed symptoms. Six days post inoculation (dpi) of bacterial cells, the first disease symptoms on B. oleracea leaves were noticeably visible on host plants inoculated with either strain, whereas no disease hallmarks were noticed on plant material with only a mock inoculation (Fig. 2a). The average lesion area caused by the inoculation of cabbage leaves with the wild-type strain was 0.15, 0.71 and 1.64 [cm2] at 6, 8 and 10 dpi, respectively (Fig. 2b). Infection with the deletion mutant strain X. campestris pv. campestris B100 Δcrt1 resulted in a remarkable increase in lesion area with a calculated value of 0.54 [cm2] at 6 dpi, 2.18 [cm2] at 8 dpi, and 3.97 [cm2] at 10 dpi. To statistically evaluate these findings, a two-tailed t-test with n ≥ 11 was conducted for each condition at each measurement time point (Supplementary Table I and Fig. 2). The analysis yielded p-values of 0.06 at dpi 6, 0.005 at dpi 8, and 0.0027 at dpi 10. These data show that deletion of the crt1 gene led to a distinct, statistically significant increase in the virulence of X. campestris pv. campestris against B. oleracea. Investigation of infection propagation was additionally performed for both of the aforementioned strains and the complemented mutant X. campestris pv. campestris B100 Δcrt1::crt1. The latter demonstrated significantly decreased infection efficiency compared to the deletion mutant X. campestris pv. campestris B100 Δcrt1. The calculated mean of the infection area was nearly equal to the mean value for leaves infected with wild-type colonies (Supplementary Fig. 4 and Supplementary Table III).
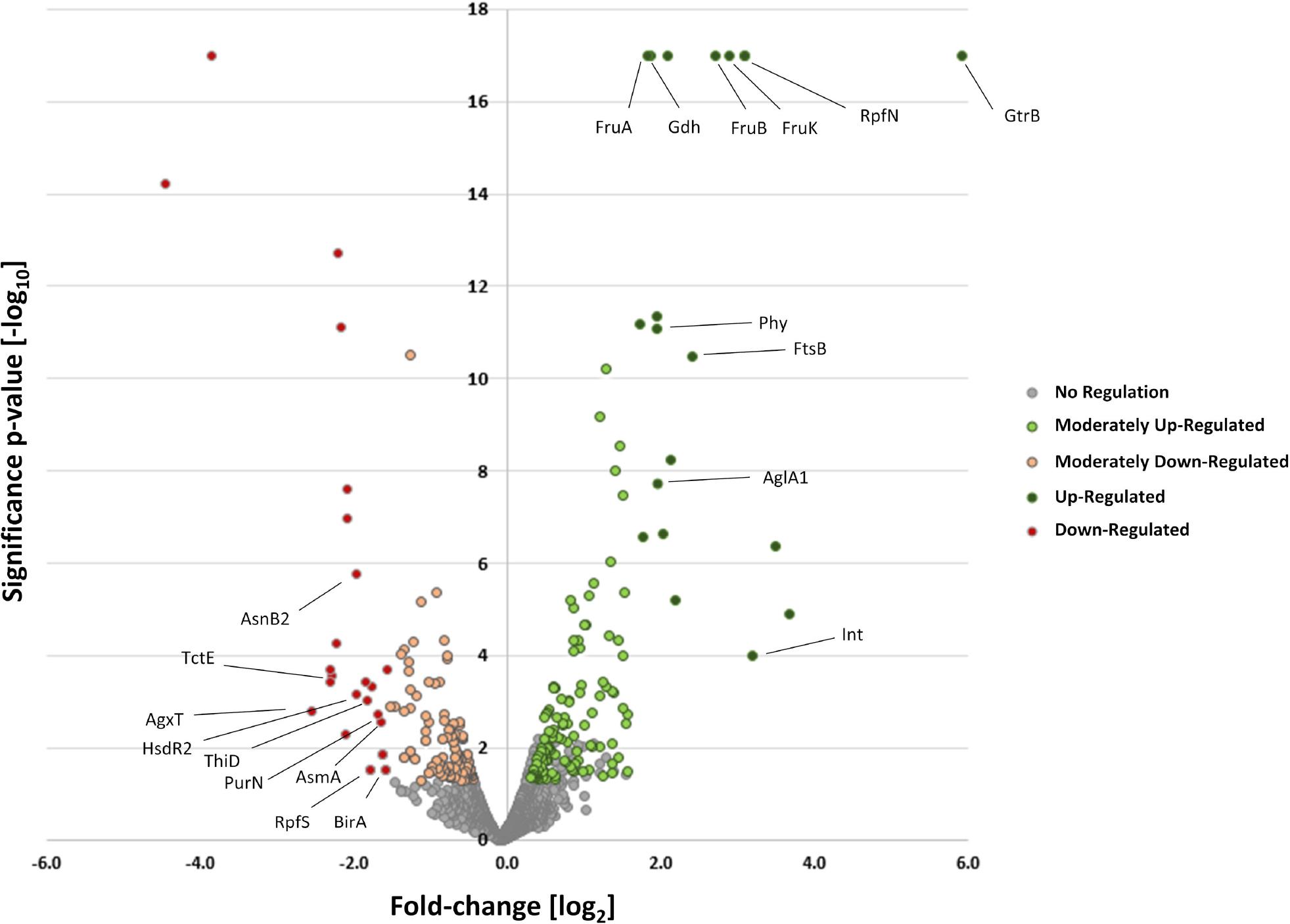
Differential proteome analysis of the deletion mutant Xcc B100 Δcrt1 compared to the wild-type strain.
Volcano plot obtained from proteome investigation of the strain X. campestris pv. campestris B100 Δcrt1 compared to the wild-type strain X. campestris pv. campestris B100; cultivation was performed in the minimal XMD medium with 10 g/l glucose as a sole carbon source; Cells were harvested at an early stationary growth stage at an oD600 nm of 1.8; shown are proteins with identified three unique peptides or more, and with peaks found for all replicates; green and red dots represent proteins with increased and decreased, respectively, abundance in the mutant strain Xcc B100 Δcrt1; cytosolic and membrane-bound proteins with expression unaffected by the deletion of crt1 gene are represented by grey dots; annotated proteins with highest fold-change compared to the wild-type strain marked with their respective protein name.
Effect of the crt1 deletion on X. campestris pv. campestris gene expression.
| Locus | Gene name | Annotated gene product | M-value | A-value | p-Value |
|---|---|---|---|---|---|
| Up-regulated genes | |||||
| XCCB100_2325 | putative filamentous hemagglutinin-related protein | 1.22 | 11.4 | 7.42E-04 | |
| AS_XCCB100_1718 | antisense transcript gumH | 1.12 | 7.5 | 3.49E-03 | |
| XCCB100_0957 | pilP | type IV pilus assembly protein | 0.91 | 9.5 | 9.00E-04 |
| XCCB100_0907 | putative cytochrome c551 | 0.90 | 8.2 | 4.92E-03 | |
| XCCB100_0552 | groES | 10 kDa chaperonin | 0.90 | 10.2 | 1.99E-03 |
| XCCB100_0905 | putative exported protein | 0.89 | 11.6 | 2.65E-03 | |
| XCCB100_3396 | putative exported enzyme | 0.86 | 10.8 | 1.23E-09 | |
| XCCB100_1423 | murD | UDP-N-acetylmuramoylalanine-D-glutamate ligase | 0.84 | 9.0 | 3.95E-03 |
| XCCB100_1720 | gumJ | xanthan repeating unit exporter | 0.83 | 8.8 | 3.25E-02 |
| XCCB100_3538 | gfo | exported glucose-fructose oxidoreductase | 0.83 | 9.4 | 3.07E-02 |
| XCCB100_0906 | lao | L-amino-acid oxidase | 0.81 | 8.6 | 8.41E-03 |
| XCCB100_0207 | amtB | ammonium transporter | 0.80 | 12.2 | 6.28E-04 |
| XCCB100_2099 | hisH | glutamine amidotransferase | 0.77 | 8.9 | 7.83E-03 |
| XCCB100_0954 | pilM | type IV pilus assembly protein | 0.75 | 9.9 | 4.70E-03 |
| XCCB100_1209 | glk2 | glucokinase | 0.74 | 7.4 | 1.86E-02 |
| XCCB100_1615 | conserved hypothetical protein | 0.74 | 8.7 | 1.76E-02 | |
| XCCB100_2311 | hexose O-acetyltransferase | 0.73 | 8.4 | 2.52E-02 | |
| XCCB100_rna061 | putative regulatory sRNA | 0.73 | 19.7 | 1.86E-04 | |
| XCCB100_1719 | gumI | mannosyltransferase | 0.73 | 9.1 | 1.21E-02 |
| XCCB100_1672 | pilE | type IV pilus assembly protein PilE | 0.72 | 10.0 | 6.89E-03 |
| Down-regulated genes | |||||
| XCCB100_2267 | infA | translation initiation factor IF-1 | -1.50 | 11.5 | 1.11E-04 |
| XCCB100_2352 | hypothetical protein | -1.30 | 8.9 | 7.20E-04 | |
| XCCB100_1776 | hypothetical protein | -1.30 | 8.1 | 4.39E-02 | |
| XCCB100_1297 | homoserine O-acetyltransferase-like enzyme | -1.20 | 9.7 | 6.91E-06 | |
| XCCB100_2980 | conserved hypothetical protein | -1.13 | 12.6 | 7.39E-09 | |
| XCCB100_3868 | conserved hypothetical protein | -1.10 | 12.5 | 3.37E-04 | |
| XCCB100_1298 | cgl | cystathionine gamma-lyase | -1.09 | 11.5 | 1.50E-07 |
| XCCB100_4110 | hypothetical protein | -1.01 | 13.4 | 3.72E-03 | |
| XCCB100_3996 | conserved hypothetical protein | -1.00 | 10.4 | 1.31E-08 | |
| XCCB100_2290 | TonB-dependent outer membrane receptor precursor | -0.97 | 8.4 | 2.37E-03 | |
| XCCB100_rna193 | putative regulatory RNA | -0.96 | 10.2 | 2.16E-03 | |
| XCCB100_3017 | cspA2 | cold shock protein | -0.93 | 11.2 | 3.27E-06 |
| XCCB100_3810 | putative secreted protein | -0.91 | 12.4 | 8.14E-03 | |
| XCCB100_1741 | conserved hypothetical protein | -0.90 | 11.1 | 1.29E-02 | |
| XCCB100_3869 | putative manganese-containing catalase | -0.90 | 12.2 | 2.48E-03 | |
| XCCB100_3892 | conserved hypothetical protein | -0.87 | 10.8 | 8.28E-03 | |
| XCCB100_2320 | conserved hypothetical protein | -0.87 | 13.1 | 7.39E-03 | |
| XCCB100_2291 | hypothetical protein | -0.86 | 11.1 | 7.85E-03 | |
| XCCB100_1773 | hypothetical protein predicted | -0.86 | 6.6 | 1.02E-04 | |
| XCCB100_3966 | putative exported protein | -0.85 | 11.7 | 2.14E-10 | |
Target genes with most fold-change of transcription under deletion of crt1 gene as compared to the wild-type strain X. campestris pv. campestris B100; M-value represents the fold-change [log2] of transcript in mutant strain compared to wild-type strain; A-value denotes the intensity of the given signal; p-value has been calculated with significance test with Holm method.
Strikingly, the deletion of the crt1 did not affect the growth of X. campestris pv. campestris in vitro, however, it greatly affected the infection propagation within the host plant tissue compared to the wild-type strain. Since in silico analysis predicted that Crt1 contains an HTH DNA-binding domain, we decided to investigate the regulatory network of the regulator Crt1 in more detail.
To investigate the effect of the deletion of the transcriptional regulator gene crt1 on the transcriptome of X. campestris pv. campestris, comparative genome-wide analysis was conducted with microarrays. For this purpose, three independent biological replicates of each strain were cultivated in minimal XMD medium with 10 g/l glucose and harvested at the early stationary growth stage with an OD600 nm of 1.8.
For this study, only target genes with p-value ≤0.05 were considered for further analysis (Fig. 3). Statistical analysis of the whole transcriptome resulted in the identification of 11 differentially expressed genes with M-values ≥ 1 or ≤ -1, manifesting that the novel transcriptional regulator Crt1 exerts its regulatory function merely as an adaptation under particular conditions. Upon deletion of crt1, only two target genes showed increased transcription (M-values ≥ 1). Significant upregulation of transcription was observed for the antisense transcript within the coding sequence of gumH. Furthermore, among 20 targets with the highest up-regulation gumI and gumJ were identified (Table I). This finding coincides with the examined phenotype of the deletion mutant, which showed significantly increased production of the EPS xanthan (Fig. 1b), and indicates that modulation of transcription of genes downstream of the primary TSS gumH significantly impacts xanthan biosynthesis, possibly serving as a bottleneck in xanthan generation.
Noteworthy is moderate upregulation of transcription of several motility-related genes coding for type IV pilus assembly proteins pilP, pilM, and pilE, with M-values of 0.91, 0.75, and 0.72, respectively. pilP and pilM are located in close proximity on the genome, whereas pilE belongs to the operon situated afar from the pil gene cluster (Alkhateeb et al. 2016; Vorhölter et al. 2008). Among down-regulated genes, a target coding for sRNA denoted by its ORF xccb100_rna193 was identified; it remains to be annotated and its biological functionality determined. However, small regulatory RNAs can modulate gene expression at the post-transcriptional level, thus complementing transcriptional regulators in their role in adapting bacteria to ever-changing environmental cues (Richards and Vanderpool, 2011; Storz et al. 2011). sRNA regulators were investigated in Xanthomonas campestris pv. vesicatoria (recently reclassified as Xanthomonas euvesicatoria pv. euvesicatoria), and further highly conserved sRNAs in X. campestris pv. campestris spp. were identified. Four novel regulatory units in X. campestris pv. campestris included sRNAs and are proposed to play an important post-transcriptional regulatory role in X. campestris pv. campestris spp. (Schmidtke et al. 2013). Interestingly, the small RNA sX13, also present in X. campestris pv. campestris, was shown to affect expression of the hrp regulon and type IV pilus biosynthesis, contributing to the regulation of bacterial virulence.
Differential transcriptional analysis revealed downregulation of several targets that code for enzymes implicated in methionine utilization. However, methionine-related proteins do not act as virulence factors; they affect xanthan production. The “methionine effect” on xanthan biosynthesis has been investigated by various research groups. Supplementation of the culture media with 5 mM methionine resulted in a 40% decrease in xanthan biosynthesis (Pielken et al. 1987; Pielken et al. 1988). The findings could be reproduced in a more recent study of yet another LysR-family transcriptional regulator, CysB (Schulte et al. 2019). Although direct evidence could not be presented in our study so far, the fact of differentially expressed methionine-related targets strongly suggests links between the novel regulator and the “methionine effect”, and further implies a putative link between methionine metabolism and virulence.
In bacteria, transcriptional regulators commonly cause a co-regulation of target genes, thus binding site motifs are usually conserved upstream of the transcription start sites of the respective genes. For the purpose of determining binding sites of novel regulator Crt1, sequences upstream of the transcription start sites (TSSs) of the considered target genes were analysed with the web-based tool MEME (Bailey et al. 2009). However, no such conserved motif could be found among the target genes identified in our study. We observed upregulation of transcripts within the gum cluster, which encodes proteins of the machinery necessary for xanthan production. Remarkably, among targets with the most significant fold-change as compared to the wildtype strain were those coding for factors essential for the virulent lifestyle of X. campestris pv. campestris, namely, T4P assembly proteins. Furthermore, the transcription of several methionine-related genes and sRNA with unknown functions was altered.
To shed light on the more systemic effects of altered transcriptional regulation caused by the deletion of the crt1 gene, a comparative analysis of the cytosolic and membrane proteomes has been carried out. X. campestris pv. campestris was cultivated under identical conditions as those used for the whole-genome cDNA microarray experiments to correlate the results of both approaches. Mass spectrometry measurements of peptide samples were performed with a nanoLC coupled to a Q Exactive mass spectrometer (Thermo Fischer, USA). In total, 2871 proteins were identified representing 64% of all proteins encoded within X. campestris pv. campestris genome. The comparative investigation of the cytosolic and membrane proteomes was completed using a background-based, nested ANOVA test. This resulted in the identification of 63 proteins with fold-change (log2) of abundance compared to the wildtype strain of ≥ 1, and 58 proteins with fold-change of ≤ - 1, at a confidence level of at least 99% (Fig. 4). Identified proteins with the highest fold-change in abundance in the deletion mutant are presented in Table II, and the list of all proteins with statistically significant fold-change is presented in Supplementary Table V.
| Locus | Gene name | Function | Abundance ratio | p-Value | Unique peptides |
|---|---|---|---|---|---|
| Up-regulated proteins | |||||
| XCCB100_1291 | gtrB | putative bactoprenol glucosyltransferase | 5.92 | 1.00E-17 | 3 |
| XCCB100_3258 | conserved hypothetical protein | 3.67 | 1.30E-05 | 4 | |
| XCCB100_3905 | putative carboxypeptidase | 3.50 | 4.47E-07 | 3 | |
| XCCB100_2818 | int | phage-related integrase | 3.19 | 1.05E-04 | 3 |
| XCCB100_1799 | rpfN | carbohydrate-selective porin | 3.08 | 1.00E-17 | 20 |
| XCCB100_3640 | conserved hypothetical protein | 3.08 | 1.00E-17 | 4 | |
| XCCB100_1801 | fruK | 1-phosphofructokinase | 2.89 | 1.00E-17 | 7 |
| XCCB100_1802 | fruB | PTS fructose porter | 2.71 | 1.00E-17 | 35 |
| XCCB100_2558 | ftsB | septum formation initiator protein FtsB | 2.41 | 3.35E-11 | 3 |
| XCCB100_2909 | conserved hypothetical protein | 2.19 | 6.21E-06 | 3 | |
| XCCB100_1450 | pirin-related protein | 2.14 | 5.65E-09 | 7 | |
| XCCB100_0445 | hypothetical protein | 2.08 | 1.00E-17 | 4 | |
| XCCB100_2092 | transcriptional regulator; Crp/Fnr family | 2.02 | 2.26E-07 | 3 | |
| XCCB100_1788 | phy | putative exported phytase | 1.96 | 8.66E-12 | 3 |
| XCCB100_0128 | TonB-dependent outer membrane receptor precursor | 1.95 | 4.39E-12 | 18 | |
| XCCB100_1697 | aglA1 | alpha-glucosidase | 1.95 | 1.81E-08 | 11 |
| XCCB100_1807 | gdh | glutamate dehydrogenase | 1.87 | 1.00E-17 | 124 |
| XCCB100_1800 | fruA | PTS fructose porter | 1.82 | 1.00E-17 | 12 |
| XCCB100_1894 | conserved hypothetical protein | 1.77 | 2.80E-07 | 7 | |
| XCCB100_1761 | conserved hypothetical protein | 1.74 | 6.40E-12 | 3 | |
| Down-regulated proteins | |||||
| XCCB100_0308 | agxT | aminotransferase | -2.54 | 1.66E-03 | 4 |
| XCCB100_3852 | putative aromatic ring-opening dioxygenase | -2.31 | 2.09E-04 | 3 | |
| XCCB100_0796 | putative nuclease / phosphatase | -2.31 | 3.84E-04 | 3 | |
| XCCB100_0846 | tctE | two-component system sensor histidine kinase | -2.28 | 2.85E-04 | 5 |
| XCCB100_2019 | short chain dehydrogenase | -2.22 | 5.37E-05 | 6 | |
| XCCB100_3906 | putative transcriptional regulator; LysR family | -2.21 | 1.93E-13 | 6 | |
| XCCB100_4189 | conserved hypothetical protein | -2.17 | 7.94E-12 | 3 | |
| XCCB100_1528 | ABC transporter permease and ATP-binding protein | -2.09 | 5.28E-03 | 5 | |
| XCCB100_1398 | DJ-1/PfpI family protein | -2.09 | 1.12E-07 | 8 | |
| XCCB100_2784 | conserved hypothetical protein | -2.08 | 2.49E-08 | 7 | |
| XCCB100_3277 | hsdR2 | Type I site-specific deoxyribonuclease (restriction subunit) | -1.96 | 7.02E-04 | 8 |
| XCCB100_2866 | asnB2 | asparagine synthase (glutamine-hydrolysing) | -1.95 | 1.68E-06 | 10 |
| XCCB100_0546 | conserved hypothetical protein | -1.85 | 3.77E-04 | 3 | |
| XCCB100_2526 | thiD | phosphomethylpyrimidine kinase | -1.83 | 9.26E-04 | 6 |
| XCCB100_2607 | rpfS | DSF-sensing histidine kinase/response regulator hybrid | -1.79 | 3.06E-02 | 3 |
| XCCB100_0601 | membrane-located putative protein phosphatase | -1.76 | 4.57E-04 | 5 | |
| XCCB100_1293 | asmA | AsmA family membrane protein | -1.64 | 2.84E-03 | 7 |
| XCCB100_2313 | NDP-hexulose epimerase | -1.62 | 1.40E-02 | 3 | |
| XCCB100_4124 | birA | bifunctional biotin repressor / co-repressor biosynthesis enzyme | -1.58 | 3.02E-02 | 3 |
| XCCB100_2292 | short chain dehydrogenase | -1.56 | 1.98E-04 | 6 | |
Proteins with most significant abundance fold-change under deletion of crt1 gene as compared to the wild-type strain X. campestris pv. campestris B100; Abundance ratio represents the fold-change [log2] measured for replicates of deletion mutant relative to replicates of wild-type; Abundance ratio p-value has been calculated with significance background-based, nested ANOVA test, with n = 3; unique peptides describe number of peptides that occur exclusively in the given target protein were employed for its identification.
Among intracellular and membrane-bound proteins with enhanced abundance upon deletion of the crt1 gene were RpfN, FruAB, and FruK, encoded by genes within a single operon. FruA and FruB form an active phosphotransferase system (PTS) fructose importer. 1-phosphofructokinase FruK transfers phosphate to the imported fructose moiety (Dills et al. 1980). One of the factors contributing to the successful X. campestris pv. campestris colonization of the host plant relies on the ability to efficiently use nutrients provided by the invaded host cell. Degradation of the plant cell wall produces a set of sugars, including fructose. Bacteria use PTS systems for the uptake of the degraded cell wall compounds (Moreira et al. 2004), as the Fru-AB PTS-importer mentioned above. The pathogenicity factor RpfN is a highly conserved regulator throughout the genus Xanthomonas and has been proposed as an outer-membrane porin or a glucose-sensitive porin.
In the deletion mutant X. campestris pv. campestris B100 Δcrt1, we measured remarkably elevated protein abundance of another protein involved in carbohydrate utilization, α-glucosidase AglA1, with a fold-change (log2) value of 1.87. As part of the sucrose and starch metabolism, it catalyses the hydrolysis of the terminal 1,4-linked αD-glucose residues from sucrose to form D-fructose. Significantly increased level of GtrB, a glucosyl transferase responsible for the transfer of sugar moieties onto the O-antigen component of the lipo-polysaccharide was observed. The O-antigen polysaccharide side chain, together with the core oligosaccharide, forms an important surface-associated virulence factor, the lipopolysaccharide (LPS) (Newman et al. 2001; Vorhölter et al. 2001). It is involved in adhesion processes and protects bacteria from the environmental conditions (Braun et al. 2005; Desaki et al. 2006; Meyer et al. 2001).
We detected a decrease in the abundance of the two-component system sensor histidine kinase, TctE, in X. campestris pv. campestris B100 Δcrt1. TctE, together with the corresponding response regulator TctD, controlled the expression of citH gene in Xanthomonas campestris pv. vesicatoria (recently reclassified as Xanthomonas euvesicatoria pv. euvesicatoria) in a culture as well as in planta and affected bacterial growth on citrate (Tamir-Ariel et al. 2011). Two-component signal transduction systems function in bacteria as part of the adaptation mechanism to ever-changing environmental stimuli (Hoch, 2000; Stock et al. 2000). Noteworthy is the significantly attenuated abundance of further histidine sensor kinase RpfS, which contains a C-terminal receiver (REC) domain, and an N-terminal Per/Arnt/Sim (PAS) domain with affinity to the diffusible signal factor (DSF), a key player of QS (An et al. 2014). DSF signalling is implicated in regulation of factors involved in pathogenicity (Ryan et al. 2011). More thoroughly investigated, DSF signal perception and transduction involve a two-component system, RpfC-RpfG. Transcriptional analysis of both RpfS and tandem RpfC/RpfG revealed a divergent subset of regulated target genes, with RpfS controlling the expression of genes involved in Type IV secretion and chemotaxis (An et al. 2013; An et al. 2014; O’Connell et al. 2013; Ryan et al. 2011). Mutations in RpfS decreased virulence of Xanthomonas solely upon spraying the bacteria onto leaf tissue of the host plant but had no effect on infection symptoms after introducing the bacteria directly into the leaf vascular system. The former requires motility for X. campestris pv. campestris to move towards the hydathodes, hence showing involvement of RpfS in DSF-dependent regulation of chemotaxis and motility (An et al. 2014). Another protein regulated by the DSF, which demonstrated significantly altered abundance in the mutant X. campestris pv. campestris B100 Δcrt1 was asparagine synthase AsnB2. The measured fold-change (log2) compared to the wild-type strain X. campestris pv. campestris B100 was -1.95. In Xanthomonas oryzae, AsnB was shown to be regulated by both DSF and the global regulator Clp, and AsnB was found to be involved not only in aspartate metabolism but also in resistance to oxidative stress, virulence, and the regulation of several virulence-related genes (Qian et al. 2013).
Remarkably, among proteins with the highest fold-change increase upon deletion of the crt1 gene, was putative phytase Phy (ORF xccb100_1788). Phytases are capable of dephosphorylating myo-inositol-hexakisphosphate (InsP6, phytate), which is the major phosphate storage in plant seeds, and is involved in defense mechanisms of the host plant against pathogens (Raboy, et al. 2003). In Xanthomonas, the Type III effector XopH was shown to have phytase activity that interferes with plant hormone pathways (Blüher et al. 2017). Chatterjee et al. (2003) suggest that the phytase PhyA contributes to scavenging more phosphate from plant tissue, thus improving the nutritional status of Xanthomonas.
X. campestris pv. campestris B100 Δcrt1 mutant demonstrated a significant increase in the abundance of the septum formation initiator protein FtsB. FtsB, together with FtsL and FtsQ, form a macromolecular complex that facilitates coordinated invagination of the plasma membrane and peptidoglycan wall (Villanelo et al. 2011). In a recent study, Ferreira et al. (2017) conducted a screening of a mutant library of Xanthomonas citri to identify novel genes involved in adaptation and virulence that had not previously been implicated in the virulence of Xanthomonas. Mutants with a disrupted ftsB gene were unable to induce hyperplasia and showed reduced necrosis in the infected host plant.
Among identified targets with increased abundance in the deletion mutant compared to the wildtype strain was glutamate dehydrogenase Gdh (ORF XCCB100_1807), which is involved in glutamate metabolism. L-glutamate is fundamental for growth in Xanthomonas and plays a central role in nitrogen metabolism, an important factor in xanthan production (Rojas et al. 2013; Soares et al. 2010).
Noteworthy as well is the significantly altered expression of two putative transcriptional regulators. Deletion of crt1 resulted in increased abundance of a yet non-characterized Crp-family transcriptional regulator encoded by the gene xccb100_2092. Significantly decreased levels were observed for the non-annotated LysR family transcriptional regulator (XCCB100_3906). Interestingly, the latter (XCCB100_3906) contains a conserved PucR-like DNA-binding domain and a substrate-binding domain of CysB type that depends on intermediates of the sulphur metabolism (Beier et al. 2002; Tyrrell et al. 1997). Nevertheless, the function of regulators with the most significant fold-change in X. campestris pv. campestris B100 Δcrt1 can only be speculated on at the present state of knowledge.
Considering the cytosolic and membrane proteins with the most significant fold-change of abundance, we inspected the respective transcriptional differences between the deletion mutant and the wild-type strain. We observed that genes coding for the above-mentioned proteins were not among the targets with the highest transcript fold changes.
This investigation revealed altered expression of various factors involved in the virulent lifestyle of X. campestris pv. campestris or linked to the pathogenicity (Table III). In the study, transcriptional upregulation of the motility-related genes pilP, pilM, and pilE was demonstrated. This observation indicates that Crt1 officiates as a repressor of pilus assembly genes. Dugé de Bernonville (et al. 2014) conducted cDNA microarray-based transcriptome analysis of X. campestris pv. campestris cultivated in xylem sap. The study showed transcriptional reprogramming of genes encoding TonB-dependent transporters and upregulation of twitching motility-related genes, including type IV pilus assembly protein-encoding genes. Upon colonization of the Brassicaceae, phytopathogens like X. campestris spp. initially penetrate xylem elements of the vascular systems of the host plant or intercellular spaces of the mesophyll parenchyma (Ryan et al. 2011). Hence, identified target genes with altered transcription upon crt1 deletion are necessary for X. campestris pv. campestris to adapt to the novel environment and host-derived metabolites and defense mechanisms.
Effect of the regulator gene crt1 deletion on transcriptome and proteome of X. campestris pv. campestris.
| Locus | Gene name | Function | COG | Fold-change | Transcript/Protein abundance change | Studies in Xanthomonas |
|---|---|---|---|---|---|---|
| Cell motility | ||||||
| XCCB100_0954 | pilM | type IV pilus assembly protein | NU | 0.748 | T | Dugé de Bernonville et al, 2014 |
| XCCB100_0957 | pilP | Tfp pilus assembly protein | NU | 0.914 | T | Dugé de Bernonville et al, 2014 |
| XCCB100_1672 | pilE | Tfp pilus assembly protein | NU | 0.725 | T | Dugé de Bernonville et al, 2014 |
| Carbohydrate transport and metabolism | ||||||
| XCCB100_1697 | aglA1 | alpha-glucosidase | G | 1.950 | P | Blanvillain et al, 2007; Fatima and Kumar, 2015; Eom et al. 2015; Ruan et al. 2014; Watt et al. 2009 |
| XCCB100_1799 | rpfN | carbohydrate-selective porin | G | 3.080 | P | Moreira et al. 2004 |
| XCCB100_1800 | fruA | PTS fructose porter | G | 1.820 | P | Moreira et al. 2004 |
| XCCB100_1801 | fruK | 1-phosphofructokinase | G | 2.890 | P | Li et al. 2019 |
| XCCB100_1802 | fruB | PTS fructose porter | G | -0.507 / 2.71 | T/P | Moreira et al. 2004 |
| Signal transduction mechanisms and information processing | ||||||
| XCCB100_0846 | tctE | two-component system sensor histidine kinase | T | -2.280 | P | Timilsina et al. 2020; Wang et al. 2016; Wang et al. 2017b |
| XCCB100_2607 | rpfS | DSF-sensing histidine kinase/response regulator | T | -1.790 | P | An et al. 2014; O’Connell et al. 2013; Ryan et al. 2007 |
| Amino acid transport and metabolism | ||||||
| XCCB100_1297 | homoserine O-acetyltransferase-like enzyme | -1.204 | T | Pielken et al. 1987; Pielken et al. 1988; O’Connell et al. 2013 | ||
| XCCB100_1298 | cgl | cystathionine gamma-lyase | -1.089 | T | Pielken et al. 1987; Pielken et al. 1988; O’Connell et al. 2013 | |
| XCCB100_2099 | hisH | glutamine amidotransferase | 0.766 | T | Li et al. 2019 | |
| XCCB100_2866 | asnB2 | asparagine synthase (glutamine-hydrolysing) | E | -1.950 | P | Qian et al. 2013 |
| EPS biosynthesis | ||||||
| AS_ XCCB100_1718** | antisense transcript gumH | 1.118 | T | Chan et al. 1999; Kiraly et al. 1997; Vojnov et al. 2001; Rigano et al. 2007; Alvarez et al. 2000; Yun et al. 2006 | ||
| XCCB100_1719 | gumI | mannosyltransferase | R | 0.726 | T | Chan et al. 1999, Kiraly et al. 1997; Vojnov et al. 2001; Rigano et al. 2007; Alvarez et al. 2000; Yun et al. 2006 |
| XCCB100_1720 | gumJ | xanthan repeating unit exporter | M | 0.833 | T | Chan et al. 1999, Kiraly et al. 1997; Vojnov et al. 2001; Rigano et al. 2007; Alvarez et al. 2000; Yun et al. 2006 |
| Lipid transport and metabolism | ||||||
| XCCB100_1788 | phy | putative exported phytase | I | 1.960 | P | Raboy 2003; Blüher et al, 2017; Chatterjee et al. 2003 |
| Cell wall, membrane, envelope biogenesis | ||||||
| XCCB100_1291 | gtrB | putative bactoprenol glucosyl-transferase | 5.92 | P | Newman et al. 2001; Vorhölter et al. 2001; Li et al. 2019 | |
| Other functions | ||||||
| XCCB100_2558 | ftsB | septum formation initiator protein FtsB | D | 2.41 | P | Ferreira et al. 2017 |
Target genes, and cytosolic and membrane-bound proteins, involved or linked to the virulent life-style of X. campestris pv. campestris, with most significant abundance fold-change under deletion of crt1 gene as compared to the wild-type strain X. campestris pv. campestris B100; Abundance ratio represents the fold-change [log2] measured for replicates of deletion mutant relative to replicates of wild-type; Abundance ratio p-value has been calculated with significance background-based, nested ANOVA test, with n = 3; unique peptides describe how many peptides unique for a single target protein were employed for its identification.
Most bacteria utilize two-component signal transduction systems as part of their adaptation to ever-changing environmental stimuli (Hoch, 2000; Stock et al. 2000). Extraordinarily, decreased abundance of two separate two-component system sensor histidine kinases, namely TctE and RpfS, was observed in this study. Histidine kinases (HK) of X. campestris pv. campestris are usually membrane-bound and paired with a cytosolic response regulator (RR) that conducts downstream regulation (Bourret and Silversmith 2010). Within the genome of X. campestris pv. campestris there are 53 genes coding for HKs, classified in three groups according to their membrane topology (Qian et al. 2008a; Qian et al. 2008b). So far, only several HKs have been investigated in X. campestris pv. campestris, among others RpfC, VgrS (ColS), PcrK, RavS or HpaS. All of the aforementioned have been implicated in the modulation of virulence, thus facilitating coordinated bacterial lifestyle transition between free swimming and virulence (Büttner and Bonas 2010; Cheng et al. 2019; Qian et al. 2008a; Tang et al. 1991; Timilsina et al. 2020; Wang et al. 2016; Wang et al. 2017a). The HK RpfC coupled with RR RpfG are encoded within the regulation of pathogenicity factors (rpf) gene cluster (Crossman and Dow 2004; Dow et al. 2003). Upon DSF recognition, RpfG hydrolyses the second messenger cyclic di-GMP, thereby averting the repression of another important QS regulator, namely Clp. Both cyclicdi-GMP and Clp contribute to the pathogenicity of X. campestris pv. campestris, as they regulate various virulence factors (Büttner and Bonas, 2010; McCarthy et al. 2008; Ryan et al. 2007; Simm et al. 2004; Tischler and Camilli 2004). In this study, the sensor histidine kinase RpfS was found to be upregulated upon deletion of the regulator gene crt1. RpfS was identified to be involved in DSF-dependent signalling, alongside RpfC-RpfG, with two-component systems controlling two separate subsets of target genes. RpfS regulated the expression of genes involved in T4 secretion and chemotaxis (An et al. 2013; An et al. 2014; O’Connell et al. 2013; Ryan et al. 2011). Furthermore, mutations in the rpfS gene resulted in decreased virulence of Xanthomonas, as intact motility was required to enable entry into the hydathodes. This leads to the conclusion that Crt1 conducts more sophisticated, complex regulation, suggesting that the novel regulator might be responsible for fine-tuning within the regulatory network of virulence factors. The biological importance of TctE was shown in a study on Xanthomonas campestris pv. vesicatoria (recently reclassified as Xanthomonas euvesicatoria pv. euvesicatoria). The two-component system TctE-TctD controlled the expression of the citH gene in a culture as well as in planta and affected bacterial growth on citrate (Tamir-Ariel et al. 2011).
X. campestris pv. campestris employs the type III secretion system (T3SS), encoded by genes within the hrp (hypersensitive response and pathogenicity) cluster, to translocate type III effector proteins (T3Es) into the host plant cell cytoplasm. A distinct family of T3Es, known as transcription activation-like effectors (TALEs) modulate the transcription of host immunity-related genes, as well as genes encoding for sugar biosynthesis and exporter systems (SWEET) of the plant, as a strategy to acquire nutrients from the host tissue (Boch and Bonas 2010; Cohn et al. 2014; Cox et al. 2017; Fatima and Kumar 2015; da Silva et al. 2002; Timilsina et al. 2020). TALEs of other Xanthomonas species were demonstrated to induce the expression of SWEET11 and SWEET13 transporter systems in rice, which promote the efflux of glucose and sucrose, respectively (Chen et al. 2010; Eom et al. 2015; Zhou et al. 2015). Sucrose is regarded as a major transport carbohydrate in planta,% essential for the growth of Xanthomonas upon colonization (Blanvillain et al. 2007; Eom et al. 2015; Fatima and Kumar 2015; Ruan 2014; Watt et al. 2009).
Considering the fact that Crt1 has been identified as a sucrose-related transcriptional regulator (Leßmeier et al. 2016), it is strongly suggested that the Crt1-directed repression is a part of the aforementioned plant-affiliated transcriptional reprogramming.
The limited overlap observed between transcriptomic and proteomic datasets is likely due to post-transcriptional and post-translational regulation, including differences in mRNA stability, translation efficiency, and protein turnover. Additionally, technical factors such as the dynamic range of protein detection and sampling may contribute to the discrepancies observed at both levels of investigation. Despite these limitations, the proteomic data provide valuable insights into the functional consequences of transcriptional regulator Crt1 deletion. We detected significantly increased protein abundance of α-glucosidase AglA1 in the deletion mutant X. campestris pv. campestris B100 Δcrt1. AglA1 hydrolyses sucrose to D-fructose as part of sucrose and starch metabolism, hence it is involved in the utilization of plant-based carbohydrates. Furthermore, we measured elevated levels of the RpfN, FruAB, and FruK proteins, encoded by a single operon and involved in fructose import and intracellular metabolism. Moreover, we detected elevated levels of putative phytase, annotated as Phy (XCCB100_1788), in the deletion mutant X. campestris pv. campestris B100 Δcrt1. The enzyme can dephosphorylate a major phosphate storage in plant seeds, phytate, and has been speculated to be involved in scavenging plant-based phosphate (Chatterjee et al. 2003; Raboy et al. 2003). Another study revealed that essential T3E XopH exerts its role through phytase activity and interferes with plant hormone pathways (Blüher et al. 2017). This reinforces the findings of this study that regulator Crt1 is involved in the re-programming of the X. campestris pv. campestris lifestyle to ensure a more efficient reclaim of nutrients from the invaded host plant tissue.
X. campestris pv. campestris produces the exopolysaccharide (EPS) xanthan, which, due to its physicochemical features, contributes to protection against environmental stresses, such as desiccation or damage by reactive oxygen species (ROS) (Kiraly et al. 1997; Vojnov et al. 2001). EPS biosynthesis in Xanthomonas was correlated with the attachment to host plant tissue and biofilm formation (Rigano et al. 2007). EPS is implied to prevent host plants from recognizing X. campestris pv. campestris and thus facilitates the successful colonization of the plant tissue (Alvarez, et al. 2000). It was also shown that xanthan causes wilting of host plants by blocking water flow in xylem vessels (Chan and Goodwin, 1999). EPS has also been implicated in suppressing callose deposition in the plant cell wall, a basal plant defense response, by binding extracellular calcium ions (Yun et al. 2006).
In this study, the generation of EPS xanthan was almost 40% higher in the deletion mutant than in the wild-type strain X. campestris pv. campestris B100. Moreover, a significant upregulation of the antisense gumH transcript abundance, as well as of the downstream genes, was observed in the mutant strain. The further downstream of gumH, the less significant was the fold-change of transcript levels between the deletion mutant and the wildtype strain. Proteins required for xanthan production and the export machinery are encoded by twelve genes contained within the gum cluster (Katzen et al. 1996; Katzen et al. 1998; Vojnov et al. 2001; Vorhölter et al. 2008). The expression of the gum operon was proposed to be directed by two promoter regions – one located upstream of gumB, and the second weak promoter located upstream of gumK (Katzen et al. 1996). Investigation of the TSSs of X. campestris pv. campestris B100 led to the identification of additional transcriptional initiation sites within the operon – additional active primary TSS were identified upstream of gumD and gumH (Alkhateeb et al. 2016). Furthermore, the study led to the detection of five antisense TSSs on the opposite strand of gumB, gumD, gumE, gumH, and gumK. We observed only moderate up-regulation of six of the twelve gum genes - gumE, gumI, gumJ, gumK, gumL, and gumM in the deletion mutant strain, with the highest fold-change identified within the transcript of the antisense TSS of gumH (Table I and Supplementary Table IV).
Among the surface-associated virulence factors are lipopolysaccharides (LPS), consisting of membrane-anchored lipid A, a core oligosaccharide, and O-antigen polysaccharide side chains (Newman et al. 2001; Vorhölter et al. 2001). The O-antigen polysaccharide is involved in adhesion processes and protection from adverse environmental conditions and thus favors the virulence of X. campestris pv. campestris. A significant increase in the abundance of GtrB, a glucosyl transferase involved in the synthesis of O-antigen was detected in the Crt1 mutant. The ties between GtrB and Quorum Sensing were demonstrated by in planta DSF/Rpf-mediated upregulation of gene expression of gtrb gene in Xca during early stages of host infection (Li et al.2019). As described previously, DSF and RpfC-Rp-fG component systems are involved in QS, regulating the expression of diverse traits in Xanthomonas spp., including those responsible for the virulent behavior of bacteria (Li et al, 2019). The schematic network of factors regulated by Crt1 and their interactions with virulence-secretion systems is depicted in Fig. 5.
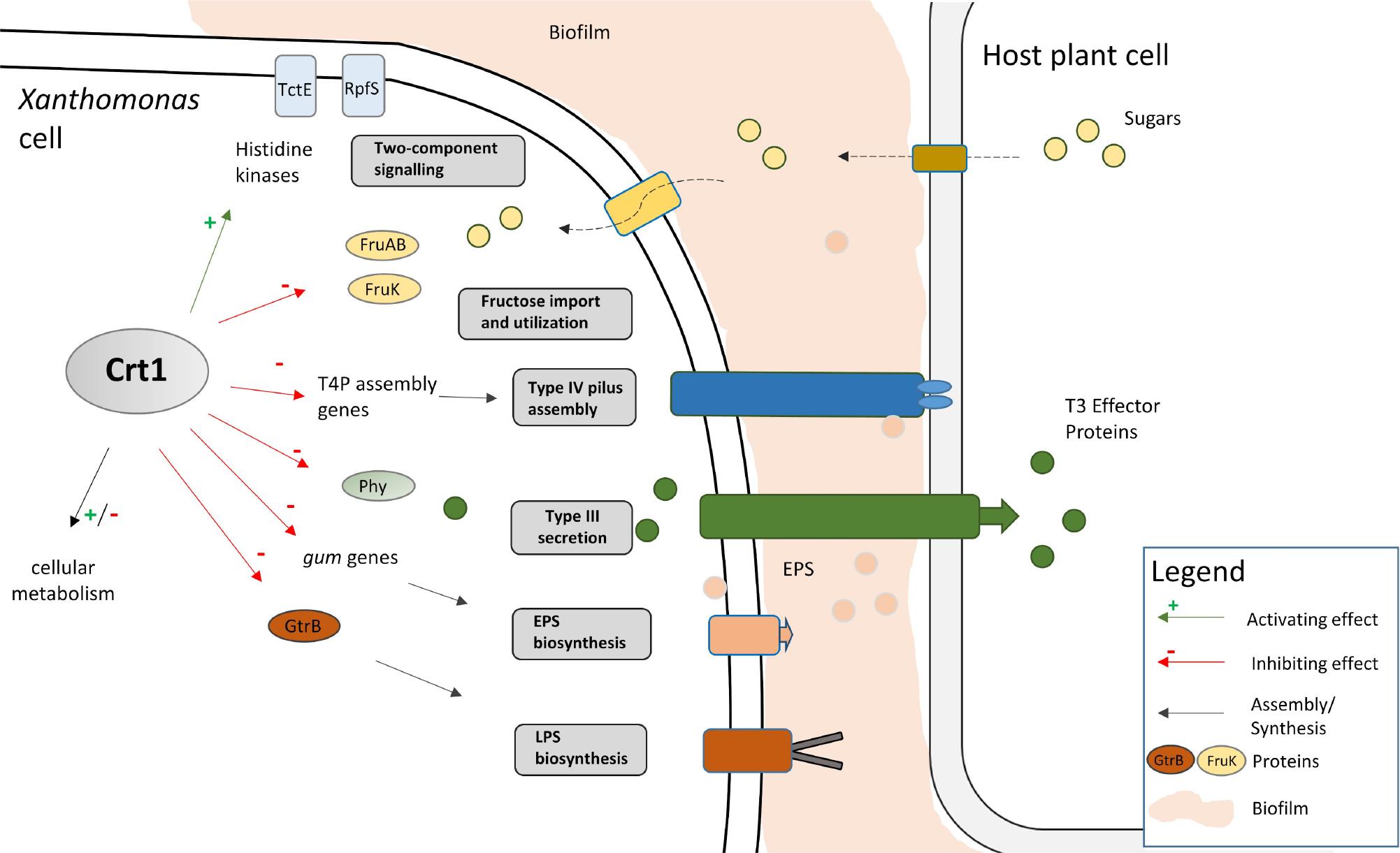
Regulatory network of the carbohydrate related transcription factor Crt1 in Xanthomonas campestris pv. campestris.
Depicted is the schematic view of the regulatory network that is influenced by the novel regulator Crt1, as well as different virulence factors in X. campestris pv. campestris. Activating effect of Crt1 is denoted by green arrows with “+”; inhibiting effect is denoted by red arrows with “-” Virulence of X. campestris pv. campestris depends on T3S system, adhesion provided by T4 pilus, EPS, and LPS. T3S system translocates T3 effector proteins into the host plant cell, where they can modulate host gene expression. T3 effectors may also activate plant defense response cues. The adaptation of X. campestris pv. campestris lifestyle during infection includes regulation of import and utilization of plant-derived nutrients. Crt1 regulates virulence factors involved in T4P assembly, EPS and LPS biosynthesis, signaling transduction and utilization of plant-related nutrients. Furthermore, novel regulator affects enzymes implicated in methionine metabolism.
In summary, with this work, we introduced the transcriptional regulator Crt1 and demonstrated in a multi-omics approach that it regulates a variety of virulence factors, including those involved in T4P assembly, EPS and LPS biosynthesis, signalling, and utilization of plant-related nutrients. The findings have been confirmed by the pathogenicity assay conducted on Br. oleracea. Infection of the host plant tissue caused by the deletion mutant X. campestris pv. campestris B100 Δcrt1 was significantly more severe than by the wildtype strain. Hence, the transcriptional regulator Crt1 we introduce and characterize in this study is involved in the modulation of virulence factors and influences the pathogenic lifestyle of X. campestris pv. campestris. However, under the deletion of the crt1 gene, merely 11 genes were significantly altered in their transcription (fold-change[log2] |M-value| of ≥ 1), and 118 proteins were differentially expressed (|abundance fold-change (log2) | ≥1). This finding underscores that Crt1 does not act as a global regulator yet controls the expression of numerous targets. Therefore, it is predicted to be a mid-tier regulator involved in re-programming during the early stage of bacterial infection.