Fig. 1.
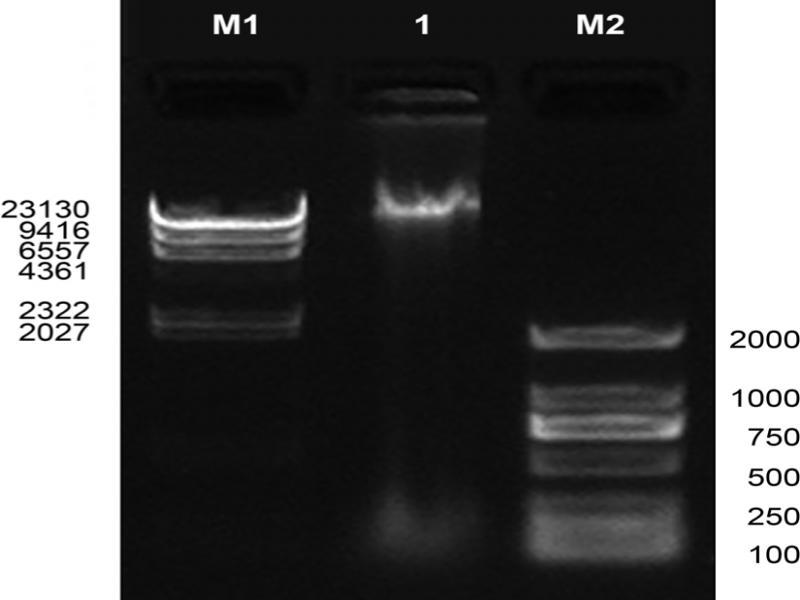
Fig. 2.
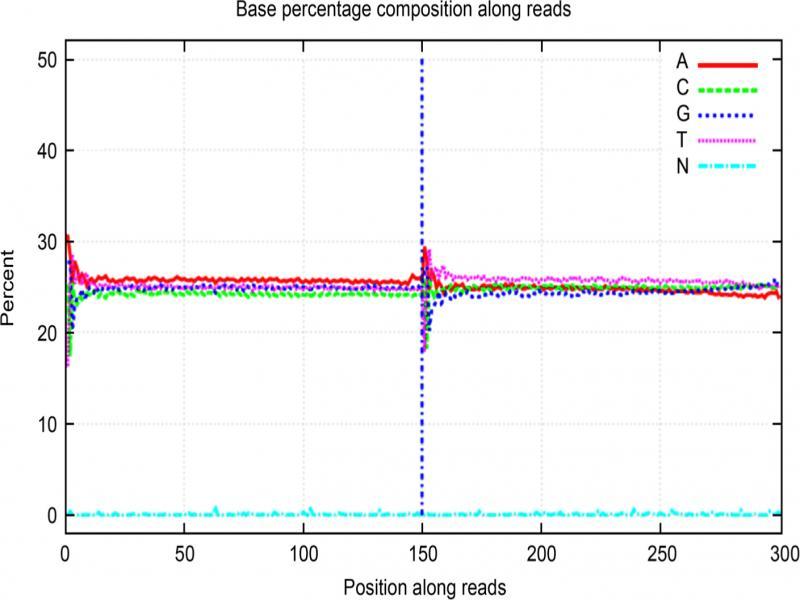
Fig. 3.
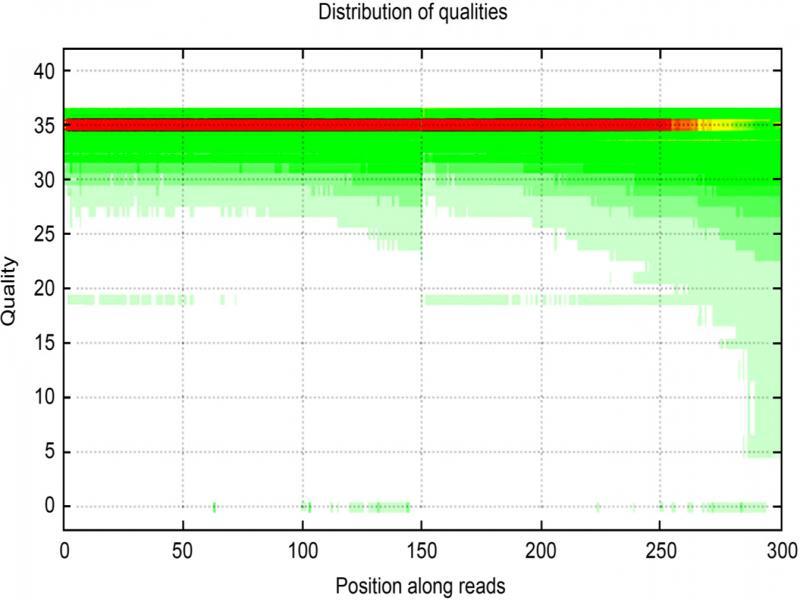
Fig. 4.
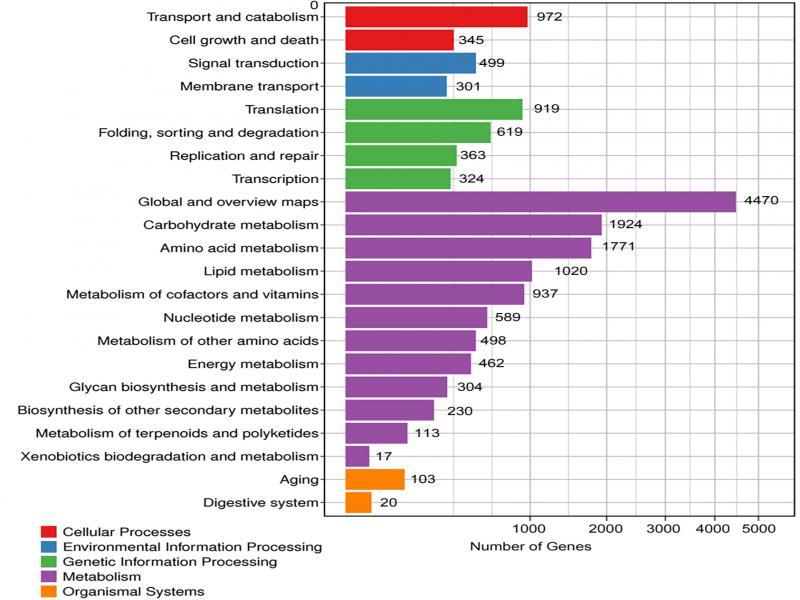
Fig. 5.
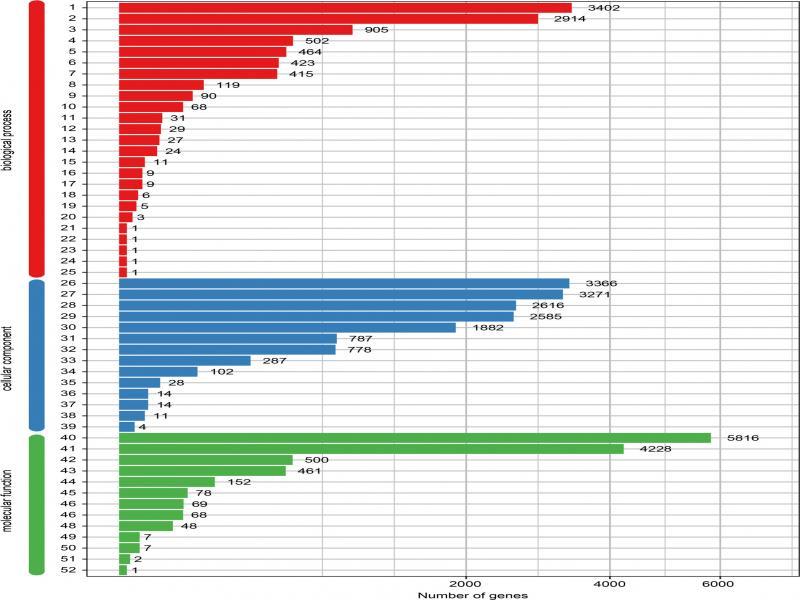
Fig. 6.
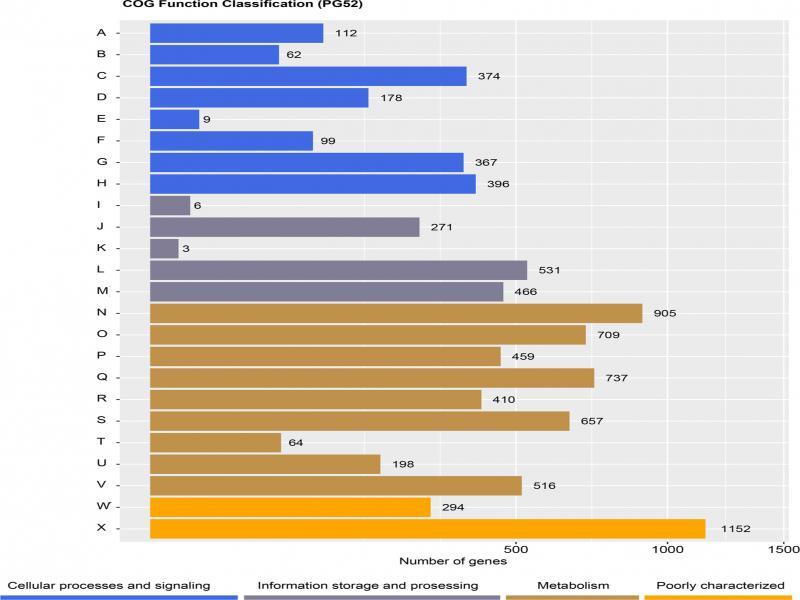
Glycosyl hydrolase families related to mycoparasitic in mycoparasites_
| Species | GH18 | GH75 | GH17 | GH55 | GH64 | GH81 |
|---|---|---|---|---|---|---|
| Pestalotiopsis sp. PG52 | 3 | 6 | 7 | 19 | 3 | 2 |
| Trichoderma harzianum | 20 | 4 | 4 | 5 | 3 | 2 |
| Trichoderma atroviride | 29 | 5 | 5 | 8 | 3 | 2 |
| Trichoderma virens | 36 | 5 | 4 | 10 | 3 | 1 |
The number of polyketide synthases and nonribosomal peptide synthetases of P_ fici, Pestalotiopsis sp_ NC0098 and mycoparasites_
| Secondary metabolites | Pestalotiopsis sp. PG52 | P. fici | Pestalotiopsis sp. NC0098 | T. harzianum | T. atroviride | T. virens |
|---|---|---|---|---|---|---|
| NRPS | 13 | 12 | 12 | 17 | 16 | 28 |
| PKS | 102 | 27 | 21 | 27 | 18 | 18 |
| Total | 115 | 39 | 33 | 44 | 34 | 46 |
The comparison of Pestalotiopsis genome sequences_
| PG52 | FICI | NC0098 | |
|---|---|---|---|
| Assembly size (Mb) | 55 | 52 | 46.61 |
| Scaffold N50 (Mb) | 6.6 | 4.0 | 5 |
| Coverage (fold) | 335.0 | 24.5 | 24 |
| GC content (%) | 53.30 | 48.73 | 51.28 |
| Protein-coding genes | 20,023 | 15,413 | 15,180 |
| Gene density (genes per Mb) | 345.22 | 296.90 | 327.08 |
| Exons per gene | 3.13 | 2.76 | 2.83 |
The genome is compared with the transcription group_
| Genes | Transcription groups | Genome | Expression rate |
|---|---|---|---|
| PKS | 82 | 102 | 80.39% |
| NRPS | 10 | 13 | 76.92% |
| Protease | 137 | 175 | 78.29% |
| Cytochrome P450 | 245 | 317 | 77.29% |
| Zn2Cys6 transcription factor | 1 | 4 | 25.00% |
Numbers of P450, protease and Zn2/Cys6 transcription factor genes of mycoparasites_
| Pestalotiopsis sp. PG52 | T. harzianum | T. atroviride | T. virens | |
|---|---|---|---|---|
| Cytochrome P450 | 317 | 50 | 15 | 40 |
| Zn2/Cys6 transcription factor | 4 | 7 | 69 | 95 |
| Protease | 175 | 53 | 23 | 28 |