FIGURE 1.
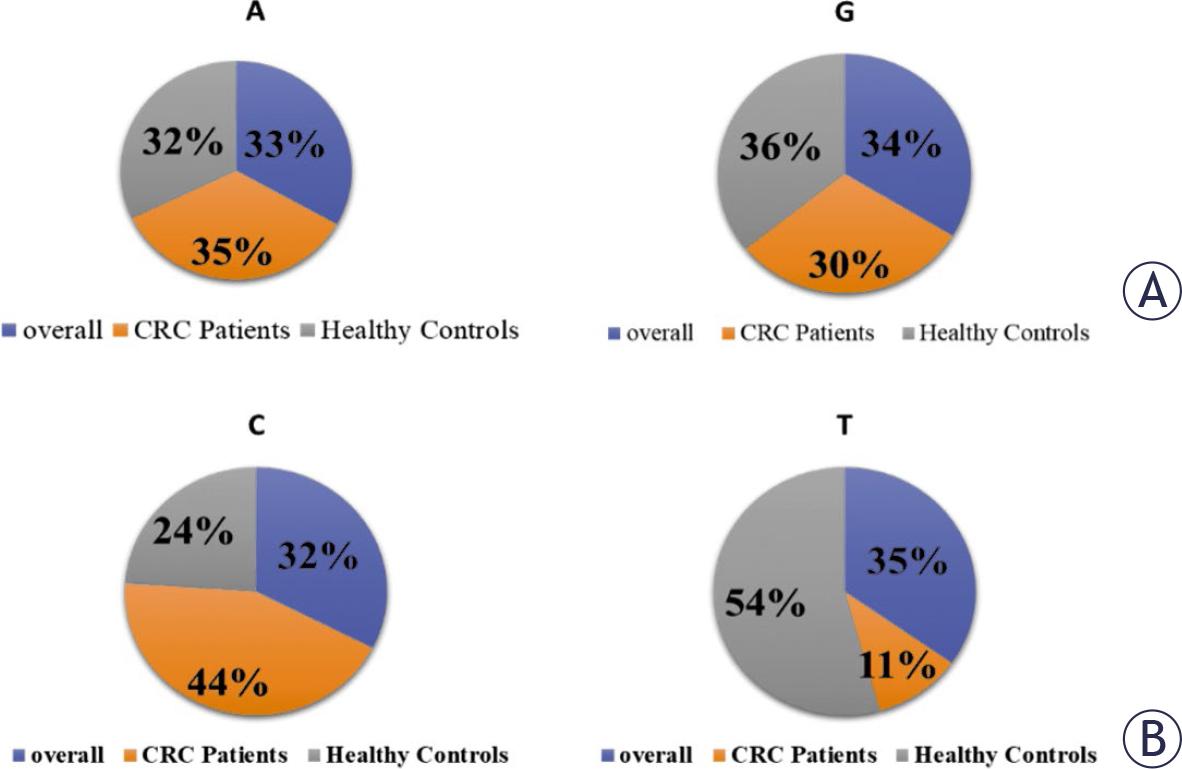
FIGURE 2.
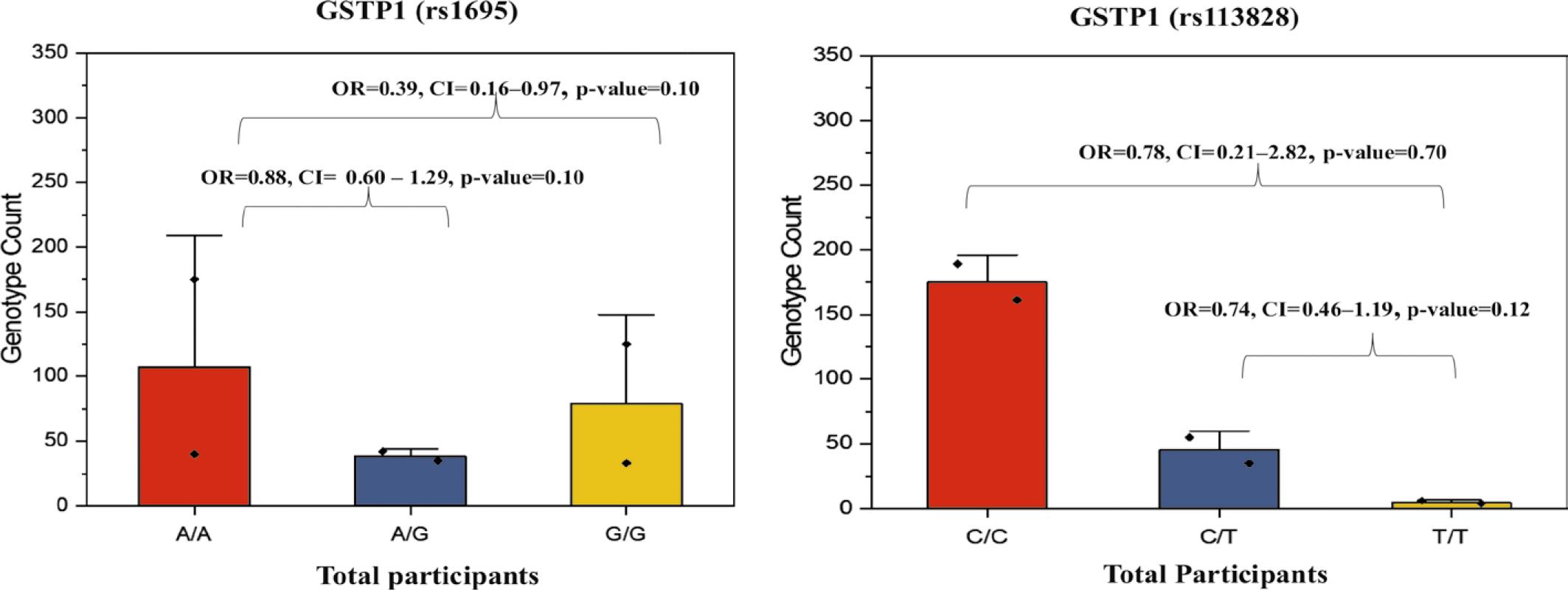
FIGURE 4.
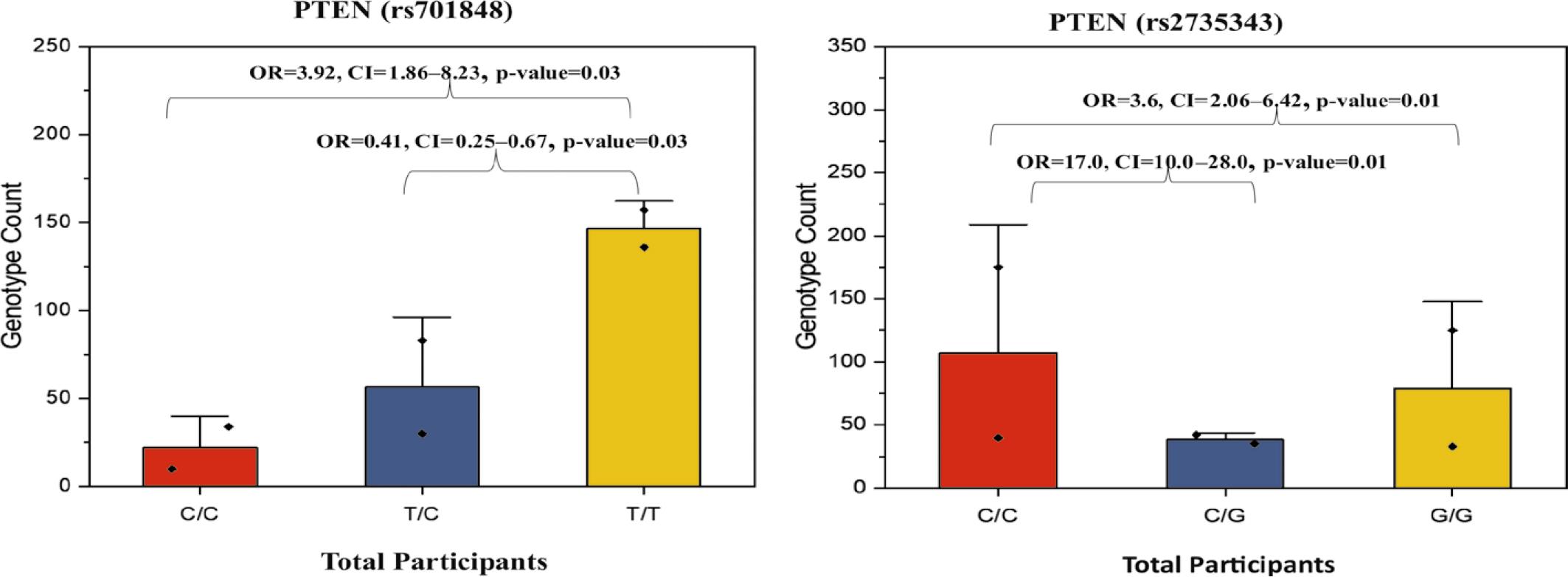
FIGURE 3.
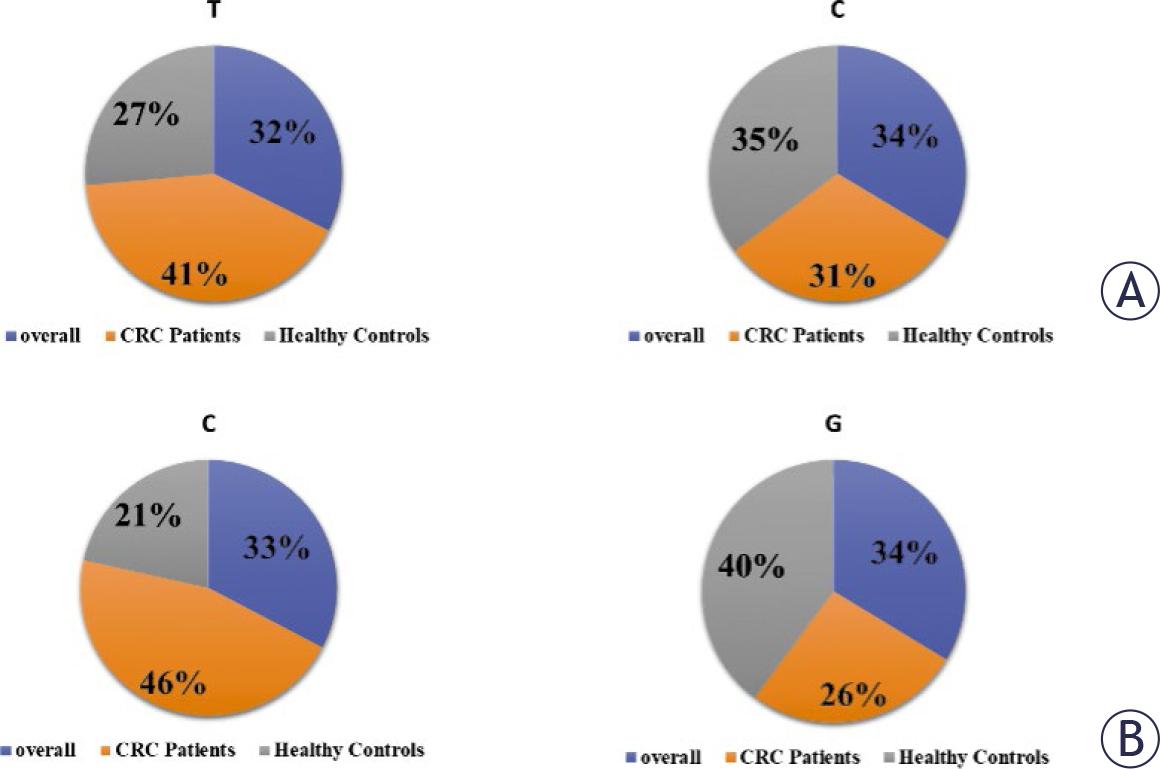
Frequency of PTEN (rs2735343) polymorphism and its association with colorectal cancer (CRC) risk
| Models/Genotype | CRC Patients +Healthy Controls n (%) | CRC Patients n (%) | Healthy Controls n (%) | OR | P Value | 95% CI | RR |
|---|---|---|---|---|---|---|---|
| Overall Subjects | |||||||
| Codominant Model | |||||||
| C/C | 215 (48) | 40 (20) | 175 (70) | Referent | _ | ||
| C/G | 77 (17) | 35 (18) | 42 (17) | 3.6 | < 0.01* | 2.06–6.42 | 0.2 |
| G/G | 158 (35) | 125 (62) | 33 (13) | 17.0 | < 0.01* | 10.0–28.0 | 0.1 |
| Dominant | |||||||
| C/G+G/G | 235 (52) | 160 (80) | 75 (30) | 9.5 | < 0.01* | 6.10–14.7 | 0.4 |
| Recessive Model (C/C+C/G) | 292 | 75 | 217 | 0.09 | < 0.01* | 0.05–0.14 | - |
| Over dominant Model | |||||||
| (C/G) | 77 (17) | 35 (18) | 42 (17) | 1.09 | 0.71 | 0.67–1.79 | - |
| Male | |||||||
| C/C | 139 (48) | 24 (20) | 115 (68) | Referent | _ | ||
| C/G | 47 (16) | 16 (13) | 31 (18) | 2.47 | < 0.01* | 1.17–5.22 | 0.2 |
| G/G | 105 (36) | 82 (67) | 23 (14) | 17.08 | < 0.01* | 9.02–32.3 | 0.2 |
| Dominant | |||||||
| C/G+G/G | 152 (52) | 98 (80) | 54 (32) | 8.70 | < 0.01* | 5.01–15.0 | 0.4 |
| Female | |||||||
| C/C | 76 (48) | 16 (21) | 60 (74) | Referent | _ | ||
| C/G | 30 (19) | 19 (24) | 11 (14) | 6.48 | < 0.01* | 2.57–16.3 | 0.1 |
| G/G | 53 (33) | 43 (55) | 10 (12) | 16.12 | < 0.01* | 6.68–38.9 | 0.1 |
| Dominant | |||||||
| C/G+G/G | 83 (52) | 62(79) | 21 (26) | 11.07 | < 0.01* | 5.28–23.3 | 0.3 |
| HWE (Genotype Frequencies) | |||||||
| C2 | 0.12 | 0.24 | 0.05 | - | - | - | |
| 2CG | 0.45 | 0.5 | 0.35 | - | - | - | |
| G2 | 0.42 | 0.26 | 0.59 | - | - | - | |
| χ2total = 1 | |||||||
Frequency of PTEN (rs701848) polymorphism and its association with colorectal cancer (CRC) risk
| Models/Genotype | CRC Patients +Healthy Controls n (%) | CRC Patients n (%) | Healthy Controls n (%) | OR | P Value | 95% CI | RR |
|---|---|---|---|---|---|---|---|
| Overall Subjects | |||||||
| Codominant Model | |||||||
| T/T | 293 (65) | 136 (68) | 157 (63) | Referent | _ | ||
| T/C | 113 (25) | 30 (15) | 83 (33) | 0.41 | 0.03* | 0.25–0.67 | 0.5 |
| C/C | 44 (10) | 34 (17) | 10 (4) | 3.92 | 0.03* | 1.86– 8.23 | 0.06 |
| Dominant | |||||||
| T/C+C/C | 157 (35) | 64 (32) | 93 (37) | 0.79 | 0.25 | 0.53–1.17 | 0.5 |
| Recessive Model (T/T+T/C) | 406 | 166 | 240 | 0.20 | < 0.01* | 0.09–0.42 | - |
| Over dominant Model | |||||||
| (T/C) | 113 (25) | 30 (15) | 83 (33) | 0.35 | < 0.01* | 0.22–0.56 | - |
| Male | |||||||
| T/T | 197 (68) | 86 (70) | 111 (66) | Referent | |||
| T/C | 71 (24) | 17 (14) | 54 (32) | 0.40 | 0.04* | 0.21–0.75 | _ |
| C/C | 23 (8) | 19 (16) | 04 (2) | 8.02 | 0.01* | 2.29-28.01 | 0.04 |
| Dominant | |||||||
| T/C+C/C | 94 (32) | 36 (30) | 58 (34) | 0.81 | 0.42 | 0.49–1.35 | 0.5 |
| Female | |||||||
| T/T | 96 (60) | 50 (64) | 46 (57) | Referent | _ | ||
| T/C | 42 (27) | 13 (17) | 29 (36) | 0.41 | 0.02* | 0.19–0.89 | 0.6 |
| C/C | 21 (13) | 15 (19) | 06 (7) | 2.05 | 0.14 | 0.77-5.48 | 0.1 |
| Dominant T/C+C/C | 63 (40) | 28 (36) | 35 (43) | 0.72 | 0.32 | 0.38–1.36 | 0.7 |
| HWE (Genotype Frequencies) | |||||||
| T2 | 0.04 | 0.04 | 0.03 | - | - | - | |
| 2TC | 0.34 | 0.32 | 0.29 | - | - | - | |
| C2 | 0.60 | 0.64 | 0.67 | - | - | - | |
| χ2total = 1 | |||||||
Association of GSTP1 and PTEN polymorphism with colon and rectum cancer cases
| Gene/rs | Genotype | Colon n = 102 (%) | Rectum n = 98 (49%) | P Value |
|---|---|---|---|---|
| GSTP1 rs1695 | AA | 62 (61.11) | 26 (27.00) | Referent |
| AG | 34 (33.33) | 72 (73.33) | < 0.01* | |
| GG | 06 (5.65) | - | 0.25 | |
| GSTP1 rs1138272 | CC | 91 (89) | 69 (69.5) | Referent |
| CT | 13 (13.0) | 17 (17.3) | < 0.01* | |
| TT | 17 (17.0) | 17 (17.3) | 0.27 | |
| PTEN rs701848 | TT | 72 (70.0) | 64 (65.4) | Referent |
| TC | 13 (13.0) | 17 (17.3) | < 0.01* | |
| CC | 17 (17.0) | 17 (17.3) | 0.02 | |
| PTEN rs2735343 | CC | 20 (19.6) | 20 (20.4) | Referent |
| CG | 19 (18.6) | 16 (16.3) | 0.71 | |
| GG | 63(61.8) | 62 (63.3) | 0.96 |
Primer sequences and amplification conditions for GSTP1 and PTEN polymorphisms
| Gene | Primer sequence | PCR conditions | Amplicon length (bp) | Restriction enzyme | Length of digest products (bp) | Enzyme specificity |
|---|---|---|---|---|---|---|
| GSTP1 (rs1695) | F:5′GGCTCTATGGGAAGGACCAGCAGG-3′ R:5′GCACCTCCATCCAGAAACTGGCG3′ | 30 cycles of 1 min at 94°C,1 min at 66°C and 2 min at 72°C. | 445 | Alw261 | 330+115+270 | 5’GTCTC(N)1↓3’ |
| GSTP1 (rs1138272) | F:5′CAGCAGAGGCAGCGTGTGTGC-3′ R:5′CCCACAATGAAGGTCTTGCCTCC-3′ | 30 cycles of 1 min at 94°C, 1 min at 64°C and 2 min at 72°C. | 565 | AciI | 365+120+485+80 | 5’C↓CG↑C’3 3’G↓GC↑G’5 |
| PTEN (rs701848) | F:5’-GTGCTTTATTGATTTGCT-3’ R:5’AGTAGTTGTACTCCGCTT-3’ | 5 min at 94°C, 35 cycles of 30 s at 94°C, 30 s at 55°C, 30 s at 72°C, and | 199 | HaeIII | 199+81+118 | 5’GG↑CC3’ |
| PTEN (rs2735343) | F:5’-CTCTTCCTGTTCTCCATCGTG-3’ R:5’-TTCTCCAGGATTTCGTCTGC-3’ | 5 min at 94°C, 35 cycles of 30 s at 94°C, 30 s at 63°C, 30 s at 72°C and 10 min at 72°C. | 272 | HhaI | 272+72+200 | 5’G↑CG↑C3’ 3’C↑GC↑G’5 |
Frequency of GSTP1 (rs1695) polymorphism and its association with colorectal cancer (CRC) risk
| Models/Genotype | CRC Patients + Healthy Controls n (%) | CRC Patients n (%) | Healthy Controls n (%) | OR | P Value | 95% CI | RR |
|---|---|---|---|---|---|---|---|
| Overall Subjects | |||||||
| Codominant Model | |||||||
| A/A | 192 (43) | 91 (46) | 101 (40) | Referent | _ | ||
| A/G | 231 (51) | 102 (51) | 129 (52) | 0.88 | 0.10 | 0.60–1.29 | 1.2 |
| G/G | 27 (6) | 07 (4) | 20 (8) | 0.39 | 0.10 | 0.16–0.97 | 0.2 |
| Dominant Model (A/G+G/G) | 258 (57) | 109 (54) | 149 (60) | 0.81 | 0.28 | 0.56–1.19 | 1.4 |
| Recessive Model (A/A+A/G) | 423 | 193 | 230 | 2.39 | 0.05* | 0.99–5.79 | - |
| Over dominant Model (A/G) | 231 (51) | 102 (51) | 129 (52) | 1.02 | 0.89 | 0.70–1.48 | - |
| Male | |||||||
| A/A | 123 (42) | 58 (48) | 65 (38) | Referent | _ | ||
| A/G | 150 (52) | 63 (52) | 87 (51) | 0.81 | < 0.01* | 0.50–1.31 | 1.3 |
| G/G | 18 (6) | 01 (0.1) | 17 (1) | 0.07 | < 0.01* | 0.01–0.51 | 0.2 |
| Dominant Model (A/G+G/G) | 168 (58) | 64 (52) | 104 (52) | 0.69 | 0.12 | 0.43–1.11 | 1.5 |
| Female | |||||||
| A/A | 69 (43) | 33 (42) | 36 (44) | Referent | _ | ||
| A/G | 81 (51) | 39 (50) | 42 (52) | 1.01 | 0.55 | 0.53–1.93 | 1.1 |
| G/G | 9 (6) | 06 (8) | 03 (4) | 2.18 | 0.55 | 0.50–9.43 | 0.1 |
| Dominant Model (A/G+G/G) | 90 (57) | 45 (58) | 45 (56) | 1.09 | 0.79 | 0.58–2.04 | 1.2 |
| HWE (Genotype Frequencies) | |||||||
| A2 | 0.46 | 0.50 | 0.436 | - | - | - | |
| 2AG | 0.43 | 0.41 | 0.449 | - | - | - | |
| G2 | 0.10 | 0.08 | 0.116 | - | - | - | |
| χ2total = 1 |
Demographic information and risk factors in colorectal cancer (CRC) cases and control
| Variable | Patients N = 200(%) | Control n = 250(%) | P value |
|---|---|---|---|
| *Age (Years) | |||
| ≥ 40 | 132 (66.0) | 151 (60.4) | 0.222 |
| < 40 | 68 (34.0) | 99 (39.6) | |
| Range | 10-60 | 11-60 | - |
| Median | 46 | 32 | - |
| *Gender | |||
| Male | 122 (61.1) | 169(68) | 0.273 |
| Female | 78 (39.0) | 81(32) | |
| **Smoking status | |||
| Never | 165 (82.3) | 22 (8.8) | < 0.010 |
| Ever | 35 (17.6) | 228 (91.1) | |
| *Site of tumor | |||
| Colon | 102(51.00) | 0.984 | |
| Rectum | 98(49.00) | ||
Frequency of GSTP1(rs1138272) polymorphism and its association with colorectal cancer (CRC) risk
| Models/Genotype | CRC Patients +Healthy Controls n (%) | CRC Patients n (%) | Healthy Controls n (%) | OR | P Value | 95% CI | RR |
|---|---|---|---|---|---|---|---|
| Overall Subjects | |||||||
| Codominant Model | |||||||
| C/C | 350 (78) | 161(80) | 189(76) | Referent | _ | ||
| C/T | 90 (20) | 35(18) | 55(22) | 0.74 | 0.12 | 0.46-1.19 | 0.2 |
| T/T | 10 (2) | 4(2) | 6 (2) | 0.78 | 0.70 | 0.21-2.82 | 0.03 |
| Dominant Model C/T+T/T | 100 (22) | 39 (19) | 61 (24) | 0.75 | 0.21 | 0.47-1.18 | 0.3 |
| Recessive Model (C/C+C/T) | 440 | 196 | 244 | 1.20 | 0.77 | 0.33-4.32 | - |
| Over dominant Model | |||||||
| (C/T) | 90 (20) | 35(18) | 55(22) | 0.75 | 0.23 | 0.46-1.20 | - |
| Male | |||||||
| C/C | 229(79) | 98(80) | 131(78) | Referent | - | ||
| C/T | 56(19) | 20(17) | 36(21) | 0.74 | 0.33 | 0.40-1.36 | 0.2 |
| T/T | 6(2) | 4(3) | 02(1) | 2.67 | 0.26 | 0.47-14.89 | 0.02 |
| Dominant Model C/T+T/T | 62(21) | 24(20) | 38(22) | 0.84 | 0.56 | 0.47-1.49 | 0.2 |
| Female | |||||||
| C/C | 121(76) | 61(78) | 60(74) | Referent | _ | ||
| C/T | 34(21) | 15(19) | 19(24) | 0.77 | 0.51 | 0.36-1.66 | 0.3 |
| T/T | 4(3) | 2(3) | 02(2) | 0.98 | 0.98 | 0.13-7.21 | 0.04 |
| Dominant Model C/T+T/T | 38(24) | 17(22) | 21(26) | 0.88 | 0.74 | 0.43-1.82 | 0.3 |
| HWE (Genotype Frequencies) | |||||||
| C2 | 0.462 | 0.81 | 0.25 | - | - | - | - |
| 2CT | 0.435 | 0.18 | 0.5 | - | - | - | - |
| T2 | 0.102 | 0.01 | 0.25 | - | - | - | - |
| χ2total = 1 | |||||||