Fig. 1.
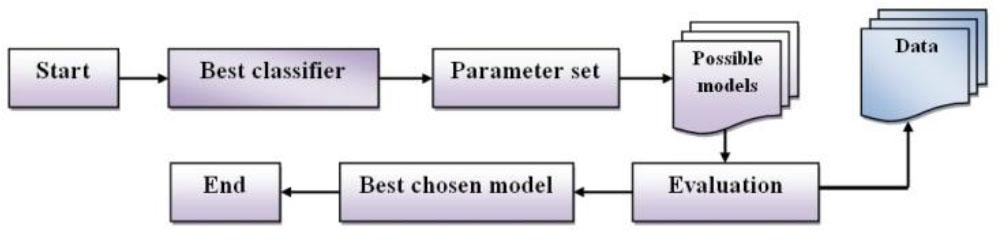
Fig. 2.
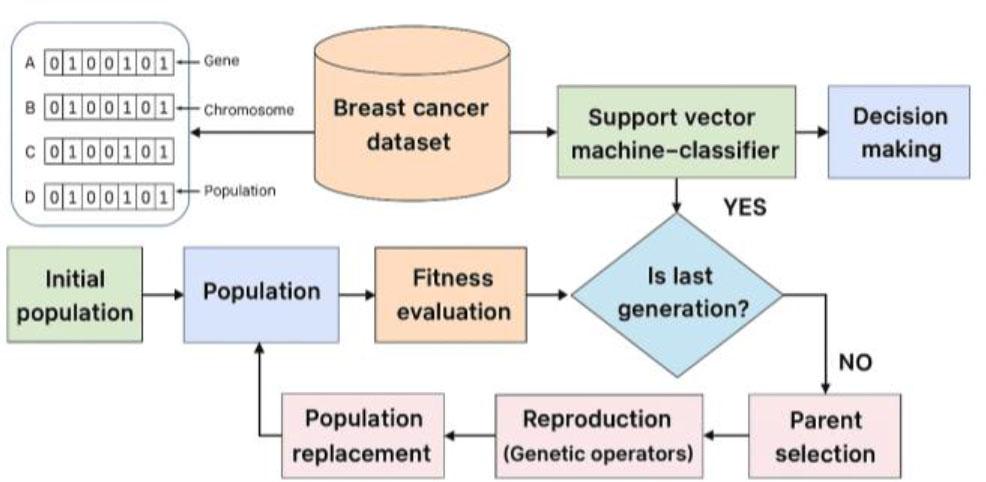
Fig. 3.
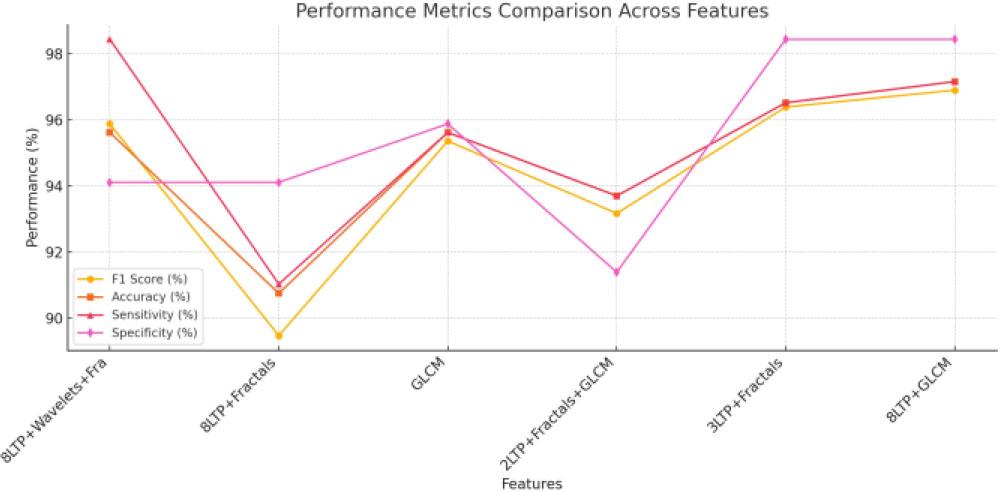
Fig. 4.
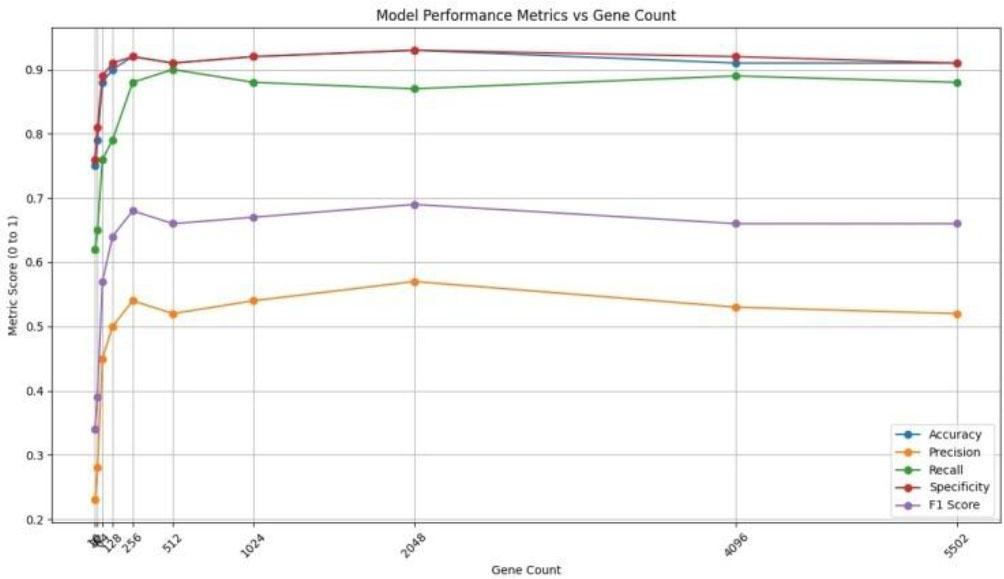
Fig. 5.
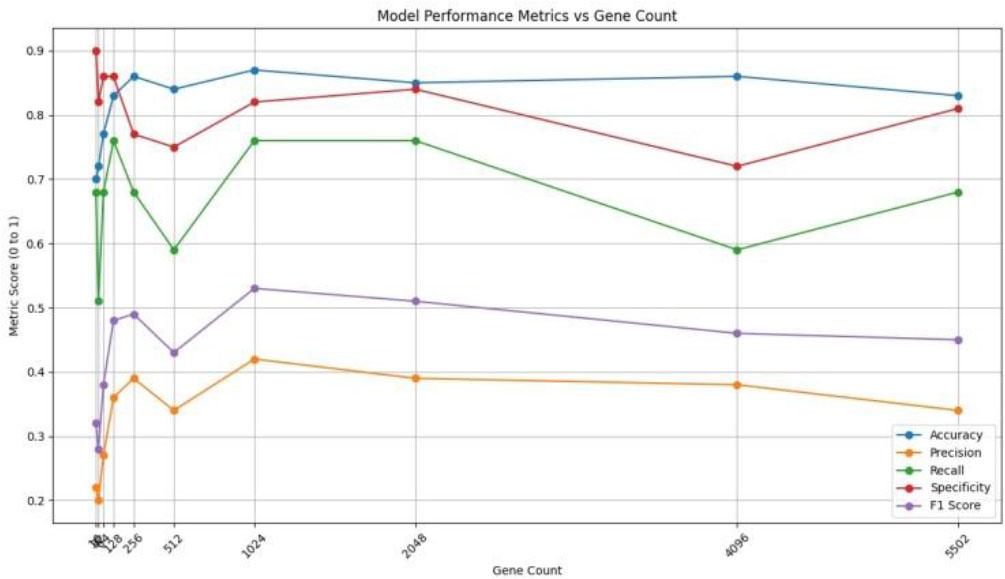
Fig. 6.
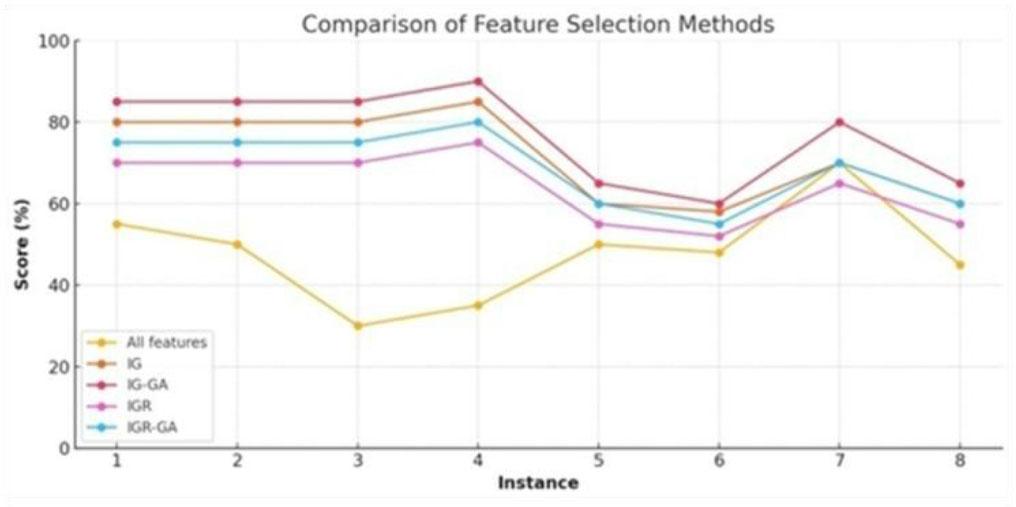
Fig. 7.
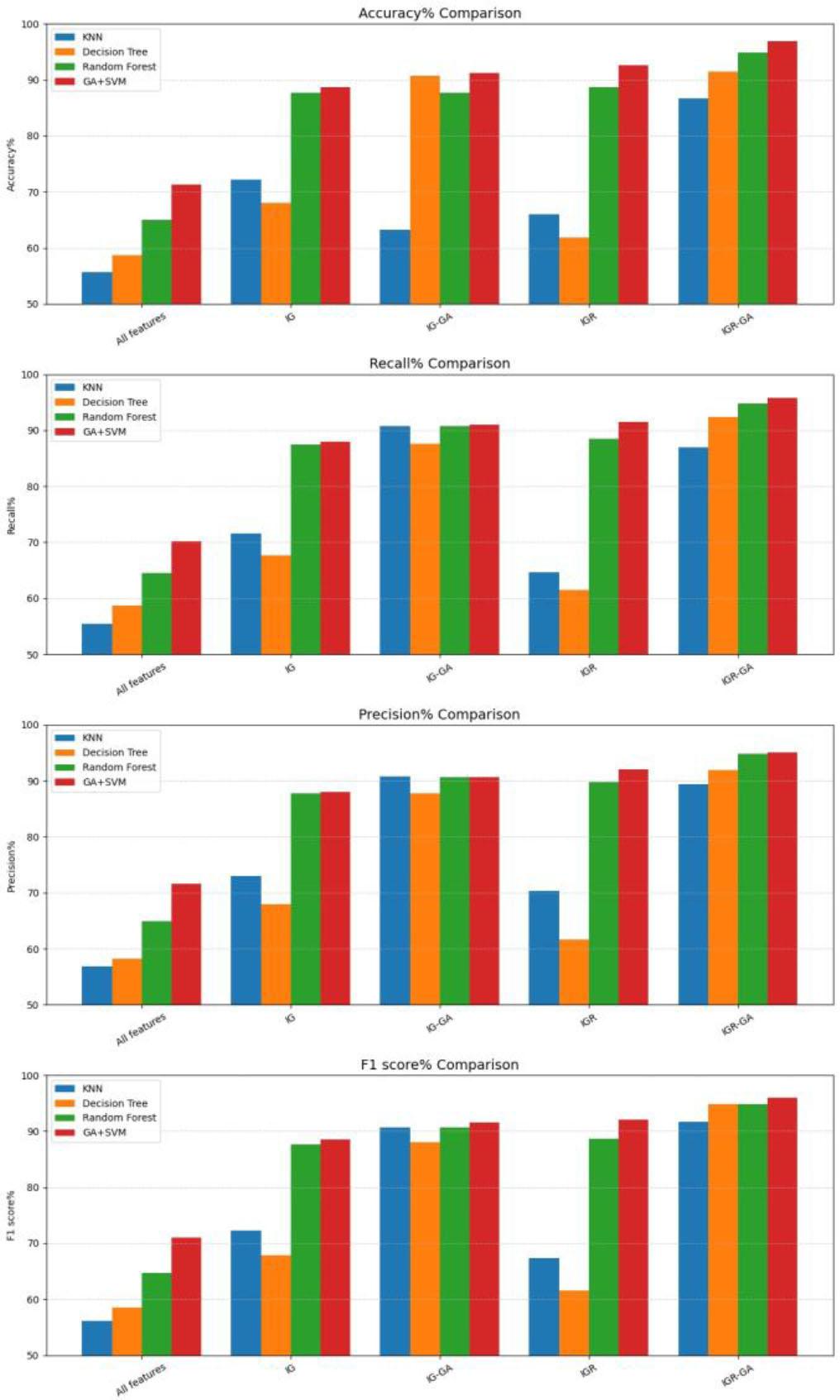
Training dataset performance evaluation based on the number of genes, (Scale: 0-1)_
| Gene count | Accuracy | Precision | Recall | Specificity | F1 score |
|---|---|---|---|---|---|
| 5502 | 0.91 | 0.52 | 0.88 | 0.91 | 0.66 |
| 4096 | 0.91 | 0.53 | 0.89 | 0.92 | 0.66 |
| 2048 | 0.93 | 0.57 | 0.87 | 0.93 | 0.69 |
| 1024 | 0.92 | 0.54 | 0.88 | 0.92 | 0.67 |
| 512 | 0.91 | 0.52 | 0.90 | 0.91 | 0.66 |
| 256 | 0.92 | 0.54 | 0.88 | 0.92 | 0.68 |
| 128 | 0.90 | 0.50 | 0.79 | 0.91 | 0.64 |
| 64 | 0.88 | 0.45 | 0.76 | 0.89 | 0.57 |
| 32 | 0.79 | 0.28 | 0.65 | 0.81 | 0.39 |
| 16 | 0.75 | 0.23 | 0.62 | 0.76 | 0.34 |
Testing dataset performance evaluation based on the number of genes, (Scale: 0-1)_
| Gene count | Accuracy | Precision | Recall | Specificity | F1 score |
|---|---|---|---|---|---|
| 5502 | 0.83 | 0.34 | 0.68 | 0.81 | 0.45 |
| 4096 | 0.86 | 0.38 | 0.59 | 0.72 | 0.46 |
| 2048 | 0.85 | 0.39 | 0.76 | 0.84 | 0.51 |
| 1024 | 0.87 | 0.42 | 0.76 | 0.82 | 0.53 |
| 512 | 0.84 | 0.34 | 0.59 | 0.75 | 0.43 |
| 256 | 0.86 | 0.39 | 0.68 | 0.77 | 0.49 |
| 128 | 0.83 | 0.36 | 0.76 | 0.86 | 0.48 |
| 64 | 0.77 | 0.27 | 0.68 | 0.86 | 0.38 |
| 32 | 0.72 | 0.20 | 0.51 | 0.82 | 0.28 |
| 16 | 0.70 | 0.22 | 0.68 | 0.90 | 0.32 |
Performance comparison of the proposed GA in combination with information gain and information ratio for different classifiers with BUDI dataset_
| Classifier | Parameter [%] | All features | IG | IG-GA | IGR | IGR-GA |
|---|---|---|---|---|---|---|
| SVM [20] | Accuracy | 53.59 | 75.24 | 85.56 | 70.08 | 83.48 |
| Recall | 51.00 | 74.90 | 85.35 | 69.68 | 83.23 | |
| Precision | 27.30 | 75.42 | 85.70 | 70.22 | 83.62 | |
| F1 score | 35.47 | 75.16 | 85.52 | 69.95 | 83.45 | |
| NB [21] | Accuracy | 49.46 | 56.67 | 56.74 | 55.65 | 63.90 |
| Recall | 47.94 | 54.48 | 56.55 | 53.72 | 61.87 | |
| Precision | 46.32 | 63.95 | 71.64 | 57.24 | 80.32 | |
| F1 score | 47.12 | 58.83 | 71.64 | 55.42 | 68.89 | |
| KNN [22] | Accuracy | 55.68 | 72.14 | 63.19 | 65.96 | 86.62 |
| Recall | 55.44 | 71.53 | 90.70 | 64.59 | 86.99 | |
| Precision | 56.79 | 72.94 | 90.78 | 70.32 | 89.42 | |
| F1 score | 56.12 | 72.23 | 90.68 | 67.33 | 91.73 | |
| DT [23] | Accuracy | 58.74 | 68.02 | 90.73 | 61.83 | 91.44 |
| Recall | 58.74 | 67.72 | 87.62 | 61.52 | 92.32 | |
| Precision | 58.26 | 67.98 | 87.72 | 61.68 | 91.88 | |
| F1 score | 58.53 | 67.87 | 88.08 | 61.60 | 94.82 | |
| RF [24] | Accuracy | 64.93 | 87.72 | 87.70 | 88.64 | 94.82 |
| Recall | 64.56 | 87.52 | 90.70 | 88.50 | 94.81 | |
| Precision | 64.86 | 87.72 | 90.66 | 89.72 | 94.81 | |
| F1 score | 64.72 | 87.67 | 90.67 | 88.62 | 94.81 | |
| GA+SVM [25] | Accuracy | 71.25 | 88.64 | 91.26 | 92.65 | 96.84 |
| Recall | 70.25 | 87.91 | 91.03 | 91.49 | 95.84 | |
| Precision | 71.62 | 88.03 | 90.64 | 92.06 | 95.02 | |
| F1 score | 71.03 | 88.56 | 91.59 | 92.12 | 96.01 |
Top 15 types of genes for differentiating breast cancer_
| Name of the gene | Chromosome | Log2FoldVariation | p-value optimization |
|---|---|---|---|
| ESR1 | 6q26.2-q26.3 | −9.966061532 | 0.003 |
| MLPH | 2q38.4 | −7.235698423 | 0.005 |
| FSIP1 | 15q15 | −7.762415635 | 0.008 |
| C5AR2 | 20q14.33 | −5.963125489 | 0.012 |
| GATA3 | 11p15 | −6.462539781 | 0.016 |
| TBC1D9 | 4q32.22 | −5.723641265 | 0.008 |
| CT62 | 15q24 | −9.213658914 | 0.002 |
| TFF1 | 22q23.4 | −14.23658974 | 0.002 |
| PRRR15 | 7q15.4 | −7.251323646 | 0.003 |
| CA12 | 15q23.3 | −7.156982345 | 0.005 |
| AGR3 | 7p22.2 | −12.36548921 | 0.001 |
| SRARP | 1p37.14 | −13.23654897 | 0.015 |
| AGR2 | 7p22.2 | −9.362145789 | 0.022 |
| BCAS1 | 21q13.3 | −7.362145587 | 0.027 |
| LINC00504 | 5p16.34 | −8.256987451 | 0.001 |
Comparison of accuracies of the proposed method with the existing techniques_
| Classifier | Proposed method | GI-SVM-RFE [%] | Fusion [%] | PCC-GA [%] | PCC-BPSO [%] | |
|---|---|---|---|---|---|---|
| IG-GA [%] | IGR-GA [%] | |||||
| SVM | 95.72 | 98.63 | NA | 96.00 | 98.63 | 98.63 |
| KNN | 86.87 | 98.63 | 88.51 | NA | 96.25 | 98.63 |
| DT | 86.72 | 88.21 | 72.51 | NA | NA | NA |
| RF | 72.20 | 83.48 | 91.00 | 89.68 | 96.26 | 86.72 |
Number of the features selected before and after applying the GA with different classifiers_
| Dataset | Classifier | All features | After applying GA | |||
|---|---|---|---|---|---|---|
| IG | IG-GA | IGR | IGR-GA | |||
| Breast dataset | SVM | 24.592 | 1225 | 612 | 1225 | 625 |
| NB | 24.592 | 1225 | 643 | 1225 | 605 | |
| KNN | 24.592 | 1225 | 622 | 1225 | 614 | |
| DT | 24.592 | 1225 | 603 | 1225 | 624 | |
| RF | 24.592 | 1225 | 611 | 1225 | 619 | |
| Average | 24.592 | 1225 | 618 | 1225 | 617 | |
Performance analysis of the proposed work_
| S. No. | Input image | Classification result |
|---|---|---|
| 1 |
| Cancer – stage II |
| 2 |
| Cancer – stage II |
| 3 |
| Cancer – stage I |
| 4 |
| Normal tissue |
| 5 |
| Cancer – stage III |
Selection models for cancer detection based on the area under the curve_
| S. No. | Features | Selection of model (Intermediate selection) | Selection of features (Eventual selection) |
|---|---|---|---|
| 1. | 8LTP + Wavelets + Fractals | 81.12 | 95.87 |
| 2. | 8LTP + Fractals | 81.12 | 97.16 |
| 3. | GLCM | 81.12 | 95.38 |
| 4. | 2LTP + Fractals + GLCM | 76.00 | 84.98 |
| 5. | 3LTP + Fractals | 74.71 | 84.98 |
| 6. | 8LTP + GLCM | 69.80 | 97.16 |
Performance evaluation of different feature groups_
| S. No. | Features | F1 score [%] | Accu [%] | Sensy [%] | Specy [%] |
|---|---|---|---|---|---|
| 1. | 8LTP + Wavelets + Fra | 95.88 | 95.62 | 98.44 | 94.11 |
| 2. | 8LTP + Fractals | 89.47 | 90.75 | 91.03 | 94.11 |
| 3. | GLCM | 95.36 | 95.62 | 95.62 | 95.88 |
| 4. | 2LTP + Fractals + GLCM | 93.17 | 93.70 | 93.70 | 91.39 |
| 5. | 3LTP + Fractals | 96.39 | 96.52 | 96.52 | 98.44 |
| 6. | 8LTP + GLCM | 96.90 | 97.16 | 97.16 | 98.44 |