The traits that enable the achievement of outstanding sports performance and differentiate individual predispositions for participating in specific disciplines, resulting from anatomical structure and physiological characteristics of the human body, are shaped by environmental and genetic factors as well as their interactions (Baker, 2011). Professional athletes are increasingly undergoing genetic testing to gain deeper insights into their physical and metabolic traits, while coaches utilize these analyses to design tailored training programs. In recent years, there has been a growing number of services offering genetic profiling for sports purposes, which are available not only to professional athletes but also to individuals who engage in recreational sports (Patel & Varley, 2019; Pickering et al., 2019; Roth, 2012).
Single nucleotide polymorphism (SNP) and insertion-deletion polymorphism (indels) are commonly used to identify traits influencing athletic performance. SNPs involve the substitution of a single nucleotide in the DNA sequence at a specific location in the genome, while indels include addition or removal of nucleotide sequence ranging from single nucleotide to several base pairs. Both types of genetic variants are widely spread throughout the human genome and have been investigated in relation to muscle strength and power, endurance, energy metabolism, recovery, and injury susceptibility (Lin et al., 2017, Ysabel et al., 2018).
Angiogenesis is a process essential for maintaining homeostasis in the body, enabling the reconstruction of blood vessels. It plays a crucial role in embryonic development, wound healing, and tissue regeneration (Shibuya, 2011). Vascular endothelial growth factor (VEGF) is one of the primary regulators of angiogenesis, stimulating endothelial cell proliferation and migration while increasing vascular permeability (Ferreira et al., 2024).
VEGF is a homodimeric protein that exists in several isoforms, generated through alternative splicing. Five main isoforms have been identified: VEGF-A, VEGF-B, VEGF-C, VEGF-D, and VEGF-E. Among them, VEGF-A plays the most significant role in regulating angiogenesis in humans. It acts as a ligand binding to and activating VEGF receptors (VEGFR-1 and VEGFR-2), initiating a signaling cascade that promotes angiogenesis, increases vascular permeability, facilitates cell migration, and expression of genes associated with vascular remodeling. The interaction between VEGF-A and the VEGFR-2 receptor activates the PLCγ-PKC-MAPK signaling pathway, which is critical for initiating endothelial cell proliferation (Karamysheva, 2008; Takahashi & Shibuya, 2005).
It has been shown that VEGF plays a crucial role in the post-injury regeneration of tendons and ligaments. The process of rebuilding damaged blood vessels ensures adequate oxygenation, nutrient supply, removal of waste products, and delivery of signaling molecules involved in the healing process (Liu et al., 2021).
The formation of new blood vessels and the repair of damaged ones are essential for maintaining proper oxygenation of the body, particularly in organs with high metabolic demands. In physically active individuals, the oxygen requirement of skeletal muscles increases significantly. Consequently, regulatory mechanisms that control oxygen delivery, including angiogenesis and vascular adaptation, can influence muscle performance, endurance, and overall physical efficiency (Bloor, 2005; Kissane & Egginton, 2019).
The genes responsible for the synthesis of VEGF-A and its receptor VEGFR-2 exhibit high genetic variability. SNPs within these genes can influence their expression levels, thereby affecting the amount of protein produced as well as potential changes in its structure and function (Komar, 2007; Shastry, 2009).
Due to the roles that the VEGFA and KDR genes play in tendon and ligament regeneration, as well as in ensuring proper oxygenation of organs, including skeletal muscles, variants of these genes are potential candidate biomarkers. These markers can help determine susceptibility to soft tissue injuries within the skeletal system as well as assess predisposition to physical exertion among athletes and physically active individuals (Balberova et al., 2022; Bonsignore et al., 2010).
This systematic review aims to examine the impact of SNPs within the VEGFA and KDR genes on susceptibility to tendon and ligament injuries, as well as on physical performance in athletes and highly physically active individuals. Additionally, it seeks to evaluate their potential as genetic markers in sports.
The articles for this study were selected by searching the following databases: PubMed, Scopus, World of Science, and EBSCO host. The search was completed on 9 January 2025. The following keywords according to MeSH were used: “VEGF,” “VEGFR,” “vascular endothelial factor,” “polymorphism,” “SNP,” “variant,” “gene,” “injury,” and “physical performance” combined using the Boolean operators AND and OR.
The review was performed according to the Preferred Reporting Items for Systematic Reviews and Meta-Analyses (PRISMA) guidelines. The review protocol was registered in the PROSPERO database under the ID number CRD420250653479.
The inclusion criteria involved original research articles published in English after the year 2000. Only case-control studies conducted on human participants were considered, specifically those analyzing the association of SNPs within the VEGFA and KDR genes with predisposition to the type of physical effort undertaken and the susceptibility to tendon and ligament injuries in athletes and physically active individuals.
Studies published before the year 2000, systematic reviews, and meta-analyses were excluded from this review. Additionally, animal studies, clinical trials, cancer research, and case studies were not considered. Research that did not include SNP analyses or did not involve athletes or physically active participants was also excluded (Table 1).
Inclusion and exclusion criteria for systematic review
| Inclusion criteria | Exclusion criteria |
|---|---|
| Articles published after year 2000 | Articles published before year 2000 |
| Articles written in English | Duplicates |
| Human studies | Animal studies |
| Original articles | Reviews and meta-analyses |
| Research includes SNPs analysis | Studies not concerning SNPs analyses |
| Studies include genes VEGFA and VEGFR (KDR) | Clinical trails |
| Genetic association studies | Case studies |
| Case-control studies | Cancer research |
| Studies including analyses of the correlation between genetic variants and predisposition to a specific type of physical performance | Studies not concerning athletes |
| Studies including analyses of the correlation between genetic variants and susceptibility to tendon and ligament injuries | |
| Studies concerning athletes |
The search process was conducted by a single investigator, who screened each record by analyzing the title, followed by the abstract, and finally the full text.
Initially, duplicate records and studies identified as systematic reviews or meta-analyses were removed. Titles were screened to determine whether they included SNP analyses within the VEGFA and KDR genes and whether they examined the association between genetic variants and susceptibility to injuries or physical performance. Subsequently, book chapters, dissertations, and conference materials were excluded. Studies were further excluded if only the abstract was available, if they were published in a language other than English, or if they involved animal research, clinical trials, or case studies (Figure 1).
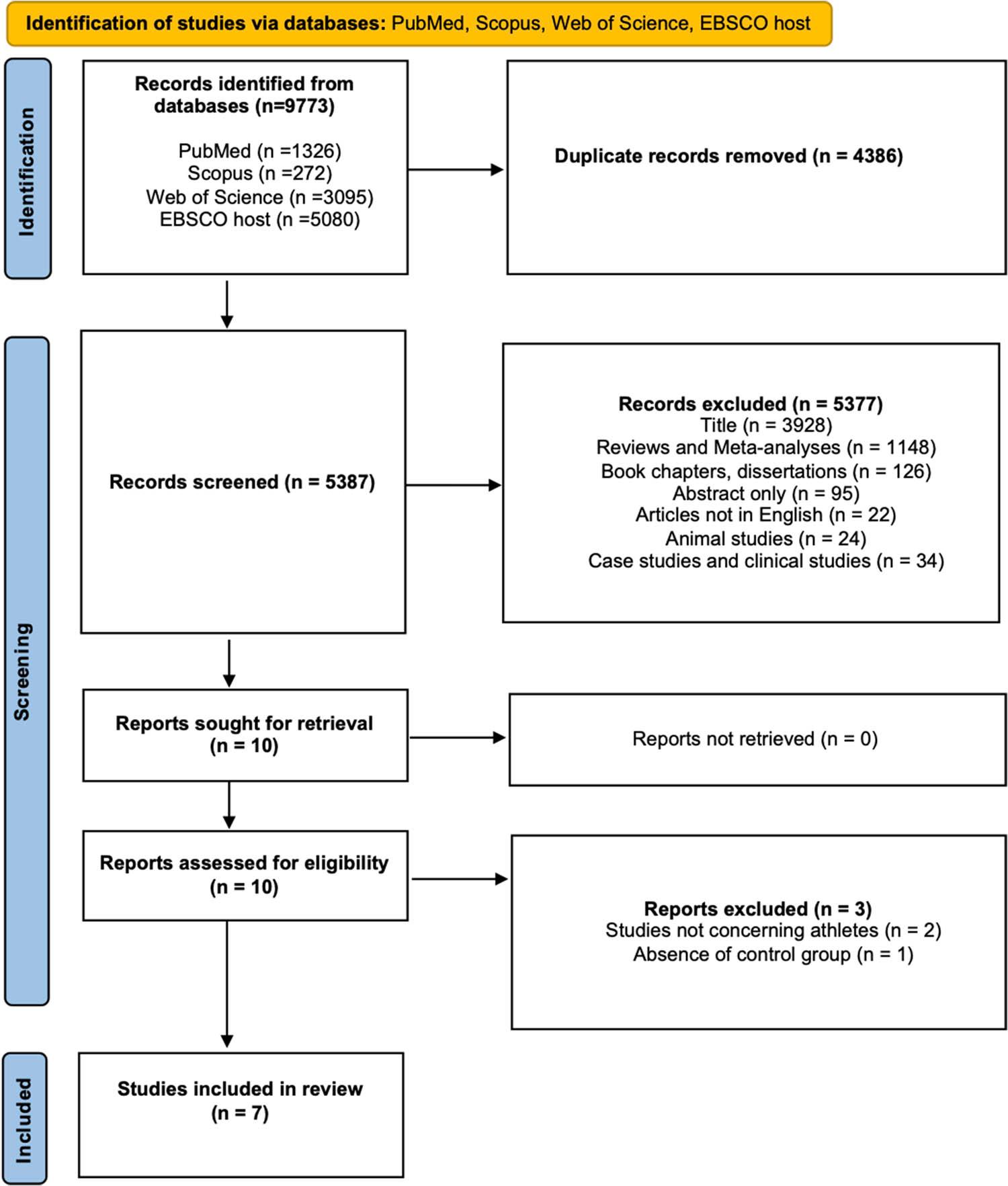
Studies selection flow diagram (PRISMA flow diagram for systematic reviews)
The Joanna Briggs Institute (JBI) checklist for case-control studies was utilized to assess the quality of the selected articles. The final articles were scored based on the JBI checklist, with a minimum acceptable score of 60% required for inclusion in this systematic review.
The following data were extracted from the selected studies, as reported: population details (sample size, ethnicity, and athletic status); genetic variants analyzed in the VEGFA and KDR genes; endurance performance and susceptibility to tendon/ligament injuries; and key findings reported by the authors, specifically allele/genotype associations, odds ratios (ORs), confidence intervals (CIs), and p-values.
The search yielded a total of 9,773 records. Of these, 4,386 duplicates, 1,148 systematic reviews and meta-analyses, 126 book chapters, 95 records with only an abstract available, and 58 studies not meeting the inclusion criteria were excluded. Additionally, 3,928 articles were removed based on their titles, as they did not focus on the VEGFA and KDR genes or did not include SNP analyses. From the 10 remaining articles, 3 studies were further excluded as they did not meet the inclusion criteria. These studies were not conducted on athletes or physically active individuals or did not include a control group.
The systematic review includes seven studies examining the association between SNP polymorphisms in the VEGFA (rs699947, rs1570360, and rs2010963, rs35569394) and KDR (rs1870377 and rs2071559) genes and their influence on endurance capacity or susceptibility to tendon and ligament injuries in athletes and physically active individuals. The summary of the analyzed articles is presented in Table 2.
Characteristics of the studies included in systematic review
| No. | First author, year | Gene | SNP variant | Participants | Country; population | Research focus | Main outcome |
|---|---|---|---|---|---|---|---|
| 1 | Ahmetov 2008 | VEGFA | rs2010963 [G/C] | Athletes | Russian; Caucasian | Physical performance | The C allele of VEGFA rs2010963 variant is associated with elite athlete status and a predisposition for enhanced aerobic performance. |
| 2 | Ahmetov 2009 | KDR (VEGFR2) | rs1870377 [T/A] | Athletes | Russian; Caucasian | Physical performance | The A allele and the AA genotype in the VEGFR2 rs1870377 variant is associated with elite athlete status in endurance sports as well as higher aerobic capacity. |
| 3 | Eider 2013 | KDR (VEGFR2) | rs1870377 [T/A] | Athletes | Poland; Caucasian | Physical performance | The A allele in the VEGFR2 rs1870377 variant is associated with predisposition to endurance sports. |
| 4 | Cięszczyk 2017 | VEGFA | rs699947 [A/C] | Athletes and physically active individuals | Poland; Caucasian | ACL injury risk (non-contact) | No significant differences between allele and genotype frequency and susceptibility to non-contact ACL ruptures in VEGFA rs699947. |
| 5 | Lulińska-Kuklik 2018 | VEGFA | rs699947 [A/C] | Physically active individuals. | Poland; Caucasian | ACL injury risk (non-contact) | The CC genotype in the VEGFA rs2010963 variant is associated with an increased susceptibility to a non-contact ACL rupture. |
| rs1570360 [G/A] | |||||||
| rs2010963 [G/C] | |||||||
| 6 | Shukla 2020 | VEGFA | rs699947 [A/C] | Athletes | India; South Asian | ACL injury risk (non-contact) | The A allele in VEGFA rs699947 and the I allele in VEGFA rs35569394 is associated with an increased risk of ACL. The C–D haplotype (rs699947, rs35569394) may present a protective effect against ACL injury. |
| rs35569394 [I/D] | |||||||
| 7 | Feldmann 2022 | VEGFA | rs699947 [A/C] | Physically active individuals. | Sweden, Poland, Australia | ACL injury risk (non-contact) | The CC genotype and C allele in VEGFA rs2010963 are associated with an increased risk of ACL injury. The A–A–G haplotype (rs699947, rs1570360, rs2010963) in VEGFA may have a protective role against ACL rupture. Additionally, gene–gene interactions between VEGFA and KDR, particularly the A–G–A–A combination, are correlated with a reduced risk of ACL injury. |
| rs1570360 [G/A] | |||||||
| rs2010963 [G/C] | |||||||
| KDR (VEGFR2) | rs1870377 [T/A] | ||||||
| rs2071559 [A/G] |
As a variant located in the 5′ UTR of the VEGFA gene, rs2010963 may influence translational efficiency and protein production, affecting VEGFA expression at both the transcriptional and translational levels and supporting its functional relevance as a regulatory SNP (Table 3). The C allele of this polymorphism is associated with higher VEGF production, which may lead to an accelerated rate of angiogenesis, particularly under hypoxic conditions. In practical terms, this may result in enhanced tissue oxygenation and improved aerobic capacity (Aziz et al., 2022; Di Stefano et al., 2015).
Genomic location and potential functional relevance of analyzed VEGFA and KDR polymorphisms
| Gene | Reference SNP number | Polymorphism type | Nucleotide change | Genomic location | Potential functional effect | Reference |
|---|---|---|---|---|---|---|
| VEGFA | rs699947 | SNP | A/C | Promoter | May alter transcription factor binding sites (TFBS) and modulate VEGFA transcription | Buroker (2014) |
| VEGFA | rs1570360 | SNP | G/A | Promoter | Regulatory SNP that may modify transcription factor binding and promoter activity | Buroker (2014) |
| VEGFA | rs2010963 | SNP | G/C | 5′ untranslated region (UTR) | May influence mRNA stability and translational efficiency, affecting VEGF-A protein levels | Buroker (2014) |
| VEGFA | rs35569394 | Indel (18 bp) | Insertion/Deletion | Promoter | May create or disrupt TFBS and influence VEGFA gene transcription | Buroker (2015) |
| KDR | rs1870377 | SNP (missense) | T/A | Coding sequence (exon 11) | Missense substitution (His472Gln); in silico analyses suggest minimal impact on protein structure, splicing regulation, or microRNA binding. | Berardi et al., (2015) |
| KDR | rs2071559 | SNP | G/A | Promoter | May alter TFBS, leading to reduced transcription | Rahim et al., (2018) |
The study by Ahmetov et al. (2008) “Polymorphism of the vascular endothelial growth factor gene (VEGF) and aerobic performance in athletes,” aimed to analyze the allele distribution and assess the relationship between genotypes and predisposition to aerobic performance in the VEGFA rs2010963 (G/C) variant.
The study included 670 cyclic sport athletes, while the control group consisted of 1,073 healthy individuals who did not engage in regular sports activities. The participants were divided into three groups based on the nature of their sport, primary energy source, and exercise duration. The first group comprised endurance athletes, where the main energy sources were fatty acids and glycogen, and exercise duration exceeded 30 min. The second group included athletes from high-power disciplines, characterized by exercise lasting between 5 and 30 min, with fatty acids and glycogen as the primary energy substrates. The third group consisted of athletes from sports where maximal exertion lasted less than 5 min, relying predominantly on glycogen and lactate metabolism.
Additionally, physiological tests assessing aerobic performance were conducted on 90 rowers (56 men and 34 women). A correlation analysis was performed between allele and genotype frequency and mean maximal oxygen uptake (VO2max) and maximal power output (Wmax). The athletes were further categorized based on sex and sports classification – Masters of Sport (MS) or Candidates for Masters of Sport (CMS).
The results indicated a higher frequency of the C allele in athletes compared to the control group (p = 0.0026). Moreover, a trend (p = 0.09) was observed, where the C allele frequency was higher in endurance athletes (Group 1) than in the control group. The mean maximal power output was significantly higher (p = 0.049) in male athletes carrying the C allele (CC + GC) compared to GG homozygotes. A trend was also observed for an association between the presence of the C allele and mean VO2max values in male MS and CMS (p = 0.09) and female CMS (p = 0.08).
The study demonstrated an association between the presence of the C allele in the VEGFA rs2010963 variant and elite athlete status, as well as a predisposition for aerobic performance.
The VEGFR2 rs1870377 (His472Gln) polymorphism, located in exon 11 of the gene encoding VEGFR-2, results in the substitution of histidine (His) with glutamine (Gln) at position 472 of the protein. This amino acid substitution may affect the structure and function of the VEGFR-2 protein, potentially modifying its tyrosine kinase activity and its ability to transmit angiogenic signals (Kim et al., 2010)
Alterations in VEGFR-2 function can influence angiogenesis, the process of new blood vessel formation, which directly impacts tissue perfusion and oxygenation. Disruptions in VEGFR-2 signaling may lead to abnormal blood pressure regulation and altered blood flow through organs, potentially resulting in insufficient tissue oxygenation (Facemire et al., 2009).
The study by Ahmetov et al. (2009), entitled “Association of the VEGFR2 gene His472Gln polymorphism with endurance-related phenotypes,” aimed to investigate the relationship between the VEGFR2 rs1870377 (T/A) polymorphism and predisposition to endurance performance and muscle fiber composition.
The study included 471 Russian athletes from various sports disciplines and 603 physically inactive individuals as a control group. Additionally, muscle fiber composition and maximal aerobic capacity (VO2max) analyses were conducted on 23 speed skaters, 76 rowers, and 45 physically active men. The athletes were categorized into four groups:
Endurance athletes (group 1), strength-endurance athletes (group 2), sprinters (group 3), and strength-oriented athletes (group 4). The control group consisted of physically inactive individuals.
An allele and genotype distribution analysis revealed a higher frequency of the A allele in endurance athletes (p = 0.0006) and sprinters (p = 0.007) (groups 1 and 3). Furthermore, the A allele was associated with elite athlete status in endurance sports compared to the control group (p = 0.0026).
Analysis of the genotype – VO2max association showed a significantly higher mean VO2max in A allele carriers (AA + AT) compared to TT homozygotes in elite male rowers (p = 0.0058) and sub-elite female rowers (p = 0.038). Muscle fiber composition analysis and genotype correlation indicated that individuals carrying the A allele exhibited a higher percentage of slow-twitch fibers compared to TT genotype individuals in both the athlete (p = 0.01) and control groups (p = 0.037).
The study demonstrated an association between the presence of the A allele and the AA genotype in the VEGFR2 rs1870377 variant with elite athlete status in endurance sports. Additionally, in high-level rowers, the A allele was linked to higher maximal aerobic capacity and an increased proportion of slow-twitch fibers. Individuals carrying the A allele may be predisposed to endurance sports, as it is associated with higher aerobic capacity.
Another study by Eider et al. (2013), entitled “The VEGFR2 gene His472Gln polymorphism in Polish endurance athletes,” investigated the association between the VEGFR2 rs1870377 polymorphism and elite endurance athlete status.
The study included 176 Polish endurance athletes, analyzed both as a whole group and further divided into specific sports disciplines: runners, cyclists, rowers, and swimmers. The control group consisted of 161 students from the University of Szczecin with low levels of physical activity.
Allele and genotype frequency analysis revealed that the A allele was significantly more frequent in the athlete group compared to the control group (p = 0.001). Additionally, the A allele was significantly overrepresented in cyclists (p = 0.006) and rowers (p = 0.019) compared to the control group.
A similar trend was observed in genotype frequency analysis. AA homozygotes were significantly more prevalent in athletes than in the control group (p = 0.014). The AA genotype was also more common in cyclists (p = 0.039) and rowers (p = 0.047) compared to physically inactive individuals.
The findings suggest that carriers of the A allele in VEGFR2 rs1870377 variant may be genetically predisposed to endurance sports.
An increased rate of angiogenesis can significantly impact the stability of connective tissue in ligaments, both by supporting regeneration and potentially contributing to their structural weakening. The formation of new blood vessels enhances vascularization and nutrient delivery, which facilitates ligament repair after injuries, such as anterior cruciate ligament (ACL) ruptures. However, chronic excessive angiogenesis may lead to structural abnormalities, including reduced collagen synthesis (Molloy et al., 2003).
The study by Cięszczyk et al. (2017), “Are genes encoding proteoglycans really associated with the risk of anterior cruciate ligament rupture?” examined the association between polymorphisms in the ACAN, BGN, DCN, and VEGFA genes and susceptibility to ACL rupture.
The analysis of the VEGFA gene focused on the rs699946 (A/C) polymorphism. The study included 372 physically active individuals of Caucasian descent, primarily soccer players. The case group comprised 229 individuals diagnosed with non-contact ACL rupture, while the control group consisted of individuals with no history of tendon or ligament injuries.
Allele and genotype frequency analysis for the VEGFA rs699947 variant revealed no significant differences between the case and control groups.
The study by Lulińska-Kuklik et al. (2018), entitled “The VEGFA gene and anterior cruciate ligament rupture risk in the Caucasian population,” analyzed the association of three VEGFA polymorphisms (rs699947, rs1570360, and rs2010963) with ACL rupture risk.
The study included 412 physically active individuals, primarily soccer players. The case group consisted of 222 individuals with a history of non-contact ACL rupture, while the control group comprised 190 individuals with no history of tendon or ligament injuries.
Significant differences in genotype distribution were observed only for the rs2010963 (G/C) variant. The CC genotype was significantly more frequent in the ACL rupture group in the codominant model (p = 0.047; OR: 2.02; 95% CI: 1.13–3.6). A similar trend was noted in the recessive model (GG + GC vs CC), where CC homozygotes were more common in the case group than in the control group.
The findings suggest that individuals with the CC genotype in the VEGFA rs2010963 polymorphism may have an increased susceptibility to non-contact ACL rupture.
The study titled “VEGFA Promoter Polymorphisms rs699947 and rs35569394 Are Associated with the Risk of Anterior Cruciate Ligament Ruptures Among Indian Athletes: A Cross-sectional Study” by Shukla et al. (2020) aimed to investigate the association between susceptibility to ACL rupture and VEGFA polymorphisms rs699947 and rs35569394.
The study included 90 Indian athletes diagnosed with ACL ruptures, with a subgroup of 62 individuals who had experienced non-contact ACL ruptures. The control group consisted of 76 healthy, physically active individuals with no history of tendon or ligament injuries.
Genotypic frequency analysis of the rs699947 (A/C) polymorphism revealed a significantly higher frequency of the A allele in the ACL group (p = 0.021; OR: 1.68; 95% CI: 1.08–2.60). In the dominant model (CC vs CA + AA), an increased prevalence of the A allele was observed in the ACL group (p = 0.004; OR: 2.69; 95% CI: 1.37–5.26). Similarly, analysis of the rs35569394 (I/D) polymorphism demonstrated that the I allele was more frequent in the ACL group (p = 0.025; OR: 1.64; 95% CI: 1.06–2.55). In the dominant model (DD vs ID + II), the frequency of the I allele was significantly higher in the ACL group (p = 0.006; OR: 2.61; 95% CI: 1.31–5.21).
Haplotype analysis of VEGFA (rs699947, rs35569394) showed that the A–I haplotype was overrepresented in the ACL group compared to the control group (p = 0.021).
These findings suggest that individuals carrying the A allele in rs699947 (A/C) and the I allele in rs35569394 (I/D) may have an increased risk of ACL rupture. Additionally, the C–D haplotype (rs699947, rs35569394) may present a protective effect against ACL injury.
Feldmann et al. (2022), in their study “Investigation of multiple populations highlight VEGFA polymorphisms to modulate anterior cruciate ligament injury,” analyzed the association between VEGFA variants (rs699947, rs1570360, rs2010963) and KDR variants (rs1870377, rs2071559) with susceptibility to ACL injury across Swedish, Polish, and Australian populations. A combined analysis of all cohorts was also conducted.
The study participants were physically active individuals. The case group consisted of individuals diagnosed with non-contact ACL rupture, while the control group included individuals with a similar level of physical activity but no history of tendon or ligament injuries.
An analysis of genotype frequency for VEGFA rs2010963 in the Swedish population revealed an increased frequency of the CC genotype (p < 0.001; OR: 6.6; 95% CI: 3.20–13.55) and the C allele (p < 0.001; OR: 2.9; 95% CI: 1.92–4.42) in the case group. Similarly, in the combined cohort analysis, the CC genotype (p = 0.0001; OR: 2.16; 95% CI: 1.47–3.19) was also more frequent in the ACL injury group.
Haplotype analysis of VEGFA (rs699947, rs1570360, rs2010963) indicated that the A–A–G haplotype was underrepresented in the case group in both the Swedish population analysis (p = 0.005) and the combined population analysis (p = 0.010).
The analysis of genotypic, allelic, and haplotype frequencies for KDR gene variants did not reveal any statistically significant differences between the case and control groups in any of the analyzed cohorts or the combined analysis.
An investigation of gene–gene interactions for VEGFA rs699947 (A/C), VEGFA rs2010963 (G/C), KDR rs2071559 (A/G), and KDR rs1870377 (T/A) revealed significant differences. The A–G–A–A (p = 0.005; OR: 0.51; 95% CI: 0.30–0.81) inferred were observed less frequently in the case group than in the control group.
The study suggests that the CC genotype and C allele in VEGFA rs2010963 (G/C) may be associated with higher ACL injury risk. VEGFA A–A–G haplotype (rs699947, rs1570360, rs2010963) was found less frequently in the ACL injury group, suggesting a potential protective role against ACL rupture. The gene–gene interactions between VEGFA and KDR, specifically for the A–G–A–A combination, were observed less frequently in the ACL-injured group, suggesting a potential protective genetic interplay that may reduce the risk of ACL rupture. No statistically significant differences for KDR gene variants (rs1870377, rs2071559) were observed between ACL-injured and control groups.
This systematic review aimed to provide further insights into the role of VEGFA and KDR polymorphisms in aerobic performance and susceptibility to ACL injury as well as to evaluate polymorphism within these genes as potential genetic markers in sports science and to contextualize VEGFA and KDR variants as potential susceptibility modifiers rather than deterministic predictors. Both genes are involved in the regulation of angiogenesis, encoding proteins that participate in a signaling pathway in which VEGF-A binds to its primary receptor, VEGFR-2, and initiates intracellular cascade that promotes endothelial cell proliferation and expansion of capillary network. This process ultimately contributes to the formation of new blood vessels and the repair of damaged capillaries. Proper regulation of this pathway ensures adequate oxygen delivery to skeletal muscles, which in athletes is associated with improved tolerance to prolonged physical exertion. Adequate tissue oxygenation is essential for the repair of microdamage within ligaments resulting from repetitive mechanical load during training or following injury. Particularly important during the phases of collagen synthesis and tissue remodeling, crucial for maintaining structural integrity of ligament tissue (Kissane & Egginton, 2019; Rahim et al., 2022).
In the analyzed studies concerning the VEGFA rs699947, Cięszczyk et al. (2017) and Feldmann et al. (2022) did not find significant differences in allele and genotype frequencies that could contribute to an increased risk of ACL injury in athletes. However, the presence of the A allele associated with a higher risk of ACL rupture was identified by Shukla et al. (2020). The VEGFA rs699947 polymorphism is located within the promoter region of the VEGFA gene and may influence its transcriptional activity, thereby modulating VEGF-A expression levels (Stevens et al., 2003). Altered expression of VEGF-A may affect the regulation of angiogenesis and extracellular matrix remodeling, processes that are crucial for both inflammatory joint diseases and ligament integrity (Rivron et al., 2012). The association of the A allele with increased susceptibility to rheumatoid arthritis (RA) and ACL rupture may be connected to mechanisms involving dysregulated vascular responses and soft tissue remodeling due to altered VEGF protein levels (Chen et al., 2012; Shukla et al., 2020).
Nevertheless, a study published by Rahim et al. (2014), which investigated the association between this polymorphism and ACL rupture risk, reported an increased frequency of the CC genotype in individuals diagnosed with ACL rupture. Similar findings were later reported by the same research team in 2022, following a study conducted on patients with surgically confirmed ACL rupture (Rahim et al., 2022).
The localization of the VEGFA rs2010963 (G/C) polymorphism within the 5′ UTR may influence mRNA stability or translational efficiency, thereby affecting VEGF-A expression and protein levels, a key factor in angiogenesis (Metzger et al., 2015). Studies suggest that the C allele of this polymorphism is associated with increased VEGF-A production, which may lead to an intensification of the angiogenic process (Ma et al., 2018). These results are consistent with previous research by Ahmetov et al. (2008), where the C allele was linked to increased maximal oxygen uptake (VO2max) and power output (Wmax), key determinants of aerobic performance. Increased VEGF-A availability may promote skeletal muscle capillarization, enhancing oxygen diffusion capacity and supporting endurance-related adaptations to training (Bloor, 2005).
When evaluating tendon and ligament injury risk, our findings indicate a significant association between the VEGFA rs2010963 CC genotype and increased ACL injury risk, consistent with the Lulińska-Kuklik et al. (2018) and Feldmann et al. (2022) studies. Additionally, the A–A–G haplotype (rs699947, rs1570360, rs2010963) in VEGFA appeared to have a protective role against ACL rupture. Notably, gene–gene interactions between VEGFA (rs699947 A/C, rs2010963 G/C) and KDR (rs2071559 G/A and rs1870377 T/A), particularly the A–G–A–A combination, were observed less frequently in ACL-injured individuals, suggesting a potential protective genetic interplay that may contribute to ligament resilience. Haplotype analyses and gene–gene interaction approaches may better capture the combined influence of multiple variants located within the same genomic region and provide a more comprehensive understanding of their functional effect than single-locus analyses, which focus on individual genetic effects. In the case of VEGFA, haplotype structure involves regulatory regions affecting mRNA stability and gene expression levels. Furthermore, gene–gene interaction analyses between VEGFA and KDR variants provide additional insight into the functional complexity of the VEGF signaling pathway. The analyzed variants are located in the promoter region, 5′ UTR, and coding sequence, and therefore may influence both gene expression and protein structure. Consequently, these variants may affect the efficiency of VEGF–VEGFR-2 signaling and the regulation of angiogenesis. Individual polymorphisms within the VEGFA and KDR genes should be interpreted as components of broader biological networks regulating angiogenesis, vascular adaptation, and tissue remodeling. Moreover, recent research in sports genomics increasingly emphasizes polygenic and multifactorial models rather than single-SNP associations. Athletic performance and injury susceptibility are complex traits influenced by the combined influence of multiple genetic variants and their interactions with environmental factors such as training load, biomechanics, and recovery strategies (Juginović et al., 2025).
The VEGFA rs35569394 polymorphism is an 18-bp indel variant located in the promoter region of the VEGFA gene and may influence changes in gene expression levels. This variant was associated with a higher ACL injury risk in I allele carriers in the study conducted by Shukla et al (2020). Additionally, the C–D haplotype (rs699947 and rs35569394) may have protective characteristics against ACL injury risk.
The KDR rs1870377 (T/A) polymorphism is a missense variant located in the coding sequence of the gene, resulting in a histidine to glutamine substitution (His472Gln). This amino acid change may influence the conformation of the VEGFR-2 receptor and potentially affect the efficiency of downstream signaling within the VEGF pathway. The A allele occurred more frequently in endurance athletes, supporting its role in vascular adaptation and slow-twitch muscle fiber composition in research conducted by Ahmetov et al. (2009). This suggests that variations in VEGFA and KDR genes may influence aerobic performance through enhanced angiogenesis, oxygen delivery, and muscle fiber composition.
Interestingly, no significant associations were found for KDR gene variants rs1870377 and rs2071559 in ACL injury susceptibility. This contrasts with their more consistently reported role in endurance-related phenotypes, suggesting that the influence of KDR polymorphisms may vary depending on the studied phenotype, indicating that while KDR polymorphisms may influence endurance performance, their role in ligament injury risk remains inconclusive. However, in 2014, Rahim et al. linked GA genotype of KDR rs2071559 polymorphism with lower risk of ACL injury in non-athlete participants (Rahim et al., 2014). Moreover, in the following study, the rs2071559 GG genotype was associated with a 2.8-fold increased risk of ACL rupture (Rahim et al., 2018). These findings suggest that, although significant associations have not yet been demonstrated in athletic populations, KDR variants may still contribute to ACL injury susceptibility in the general population and therefore should not be excluded from consideration as potential genetic markers. The current evidence regarding the role of KDR variants in ACL injury susceptibility in athletes remains limited. To date, studies conducted in athletic populations have investigated only two polymorphisms within the KDR gene (rs1870377 and rs2071559), which restricts the ability to draw broader conclusions about the involvement of this gene in ligament injury risk. Moreover, studies conducted in general populations have reported inconsistent findings regarding the functional significance of these variants.
Beyond their relevance to athletic traits, several of the analyzed variants have also been investigated in the context of human diseases. For example, the VEGFA rs699947 polymorphism has been associated with susceptibility to RA, with studies conducted by Hussain et al. and Paradowska-Gorycka et al. linking the A allele to an increased risk of RA development (Hussain et al., 2023; Paradowska-Gorycka et al., 2016), highlighting the broader biological relevance of VEGF signaling in inflammatory and connective tissue disorders. Moreover, enhanced angiogenesis can influence various physiological and pathological processes, including tissue regeneration and tumor development (Melincovici et al., 2018). In line with this, indel VEGFA variant, rs35569394, has been associated with bladder cancer development, a disease in which tumor growth, progression, and metastasis are strongly dependent on angiogenic activity (El Azzouzi et al., 2023).
The conducted systematic review has some limitations. Major among them is the small number of eligible studies. In addition, many of the included studies were conducted with small samples that included groups primarily from Caucasian populations, limiting the applicability of the results to other ethnic groups. A difference in sample characteristics, definitions of injury or performance outcomes, and genetic variants analyzed prevented a meta-analysis. Therefore, no statistical assessment of heterogeneity or publication bias was performed. Due to the limited number of eligible studies and the lack of quantitative synthesis, the possibility of bias could not be formally assessed. Additionally, most of the analyzed variants are classified in clinical databases as non-pathogenic modifiers rather than pathogenic mutations, which limits their direct clinical applicability. Moreover, the study selection was conducted by a single investigator, which may increase the risk of selection bias. Despite these limitations, this review provides a structured synthesis of current knowledge on VEGFA and KDR gene polymorphisms in the context of endurance performance and injury susceptibility, highlighting key genetic variants and directions for future research.
Research on the impact of VEGFA and KDR gene polymorphisms on physical performance and susceptibility to tendon and ligament injuries in athletes remains limited. Therefore, VEGFA and KDR variants should be interpreted as susceptibility modifiers rather than as deterministic predictors of performance or injury risk. However, the available evidence suggests that certain VEGFA variants may be associated with susceptibility to ACL injuries. Furthermore, the VEGFA and KDR variants may serve as potential biomarkers for evaluating predispositions to endurance sports. These findings may contribute to the development of genetic profiling strategies in sport, supporting individualized approaches to training optimizations, injury prevention, and recovery strategies in physically active individuals and professional athletes.
This research was funded by the Ministry of Science and Higher Education in 2025/2026 as part of the University Research Project of the University of Physical Education in Warsaw UPB Nr 12 – The influence of genetic conditions on sports performance – performance profiling based on candidate genetic variants.
Study design – K. Jabłońska-Paszko, K. Krawczak-Wójcik; Data collection – K. Jabłońska-Paszko; Data interpretation – K. Jabłońska-Paszko, K. Krawczak-Wójcik, E. Maculewicz; Manuscript preparation – K. Jabłońska-Paszko, E. Maculewicz; Literature search – K. Jabłońska-Paszko; Funding acquision – E. Maculewicz.
Authors state no conflict of interest.