Figure 1.
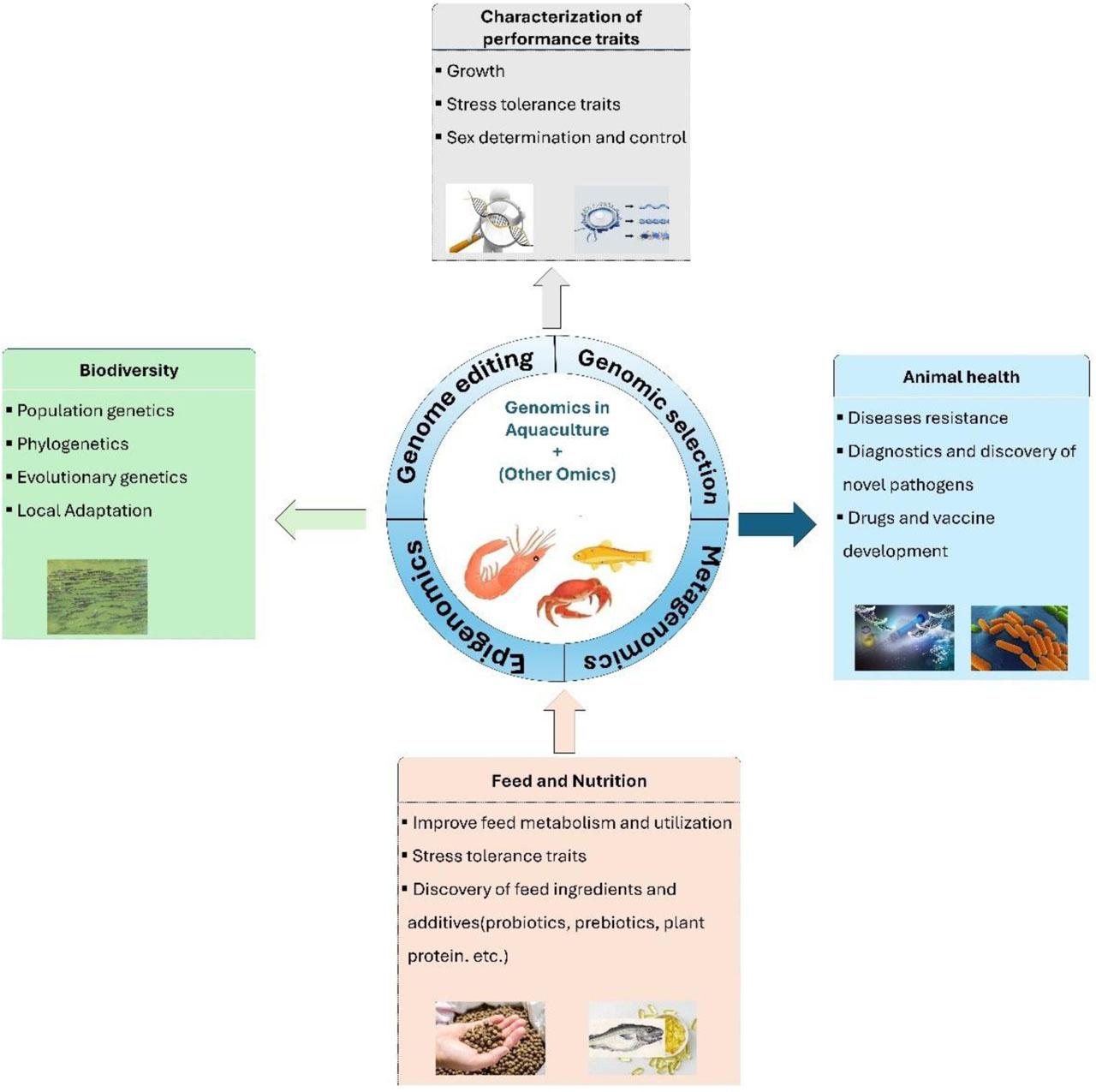
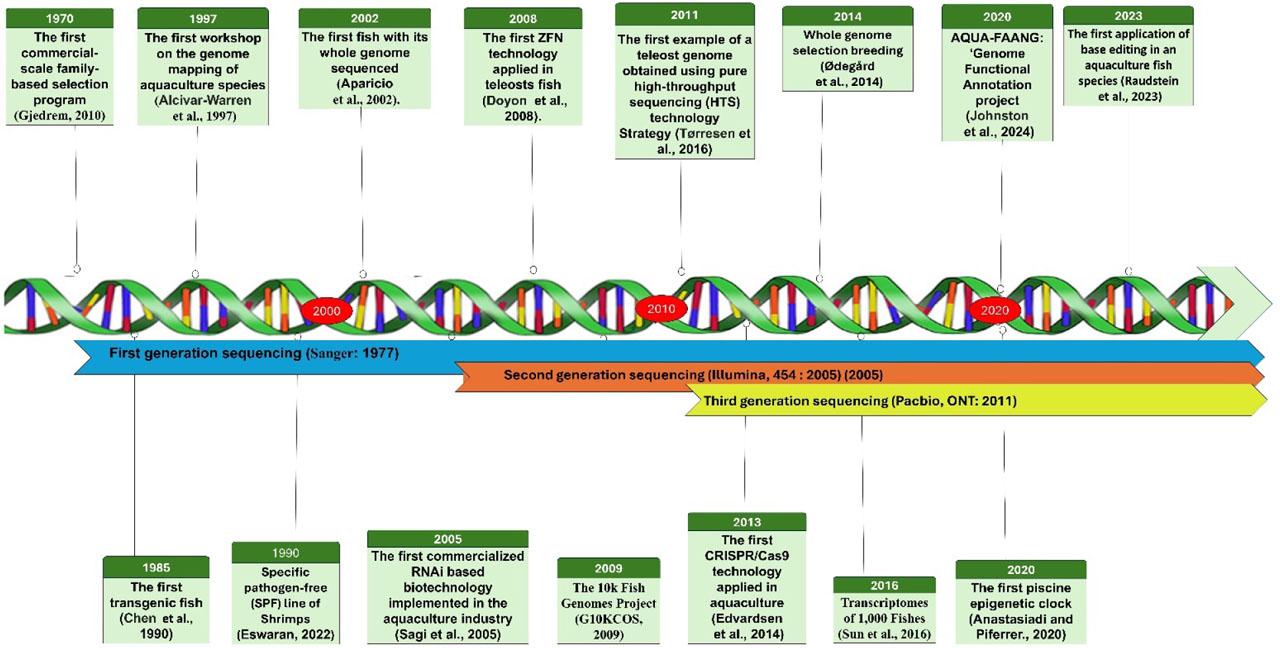
Figure 2.
Identification of QTLs in aquaculture animals using linkage mapping
| Species | Number and traits of QTLs | Number of candidate genes | Reference |
|---|---|---|---|
| 1 | 2 | 3 | 4 |
| Atlantic salmon | 3 QTLs for sea lice resistance | 2 | (Robledo et al., 2018 a) |
| 2 QTLs for piscine myocarditis virus (PMCV) disease resistance | 6 | (Hillestad and Moghadam, 2019) | |
| 2 QTLs for cardiomyopathy syndrome resistance | 4 | (Boison et al., 2019) | |
| Common carp | 18 QTLs for head size | 10 | (Chen et al., 2018) |
| 1 QTL for koi herpesvirus (KHV) disease resistance | (Palaiokostas et al., 2018) | ||
| 17 QTL for feed conversion efficiency | (Zhang et al., 2021) | ||
| Grass carp | 4 QTLs for growth | 17 | (Huang et al., 2020) |
| Rainbow trout | 24 QTLs for Aeromonas salmonicida resistance | 33 | (Marana et al., 2021) |
| 21 QTLs for infectious hematopoietic necrosis disease resistance (IHN) | (Vallejo et al., 2019) | ||
| 5 QTLs for BCWD | (Fraslin et al., 2018) | ||
| 6 QTLs for ESC resistance | 37 | (Shi et al., 2018) | |
| 3 QTLs for ESC resistance | 55 | (Tan et al., 2018) | |
| Channel catfish | 6 QTLs for growth, 10 QTLs for sex | 25 | (Zhang et al., 2019 b) |
| Yangtze River common carp | 21 QTLs for growth, 4 QTLs for sex | 5 | (Feng et al., 2018) |
| Yellow river carp | 29 QTLs for growth | 3 | (Wang et al., 2022 b) |
| Largemouth bass | 32 QTLs for growth, 13 QTLs for sex | (Dong et al., 2019) | |
| Large yellow croaker | 7 QTLs for Cryptocaryon irritans disease resistance | 29 | (Kong et al., 2019) |
| Snapper | 4 QTLs for growth | 13 | (Ashton et al., 2019) |
| Pearl oyster | 32 QTLs for growth, 1 QTL for sex trait | 4 | (Liu et al., 2020 a) |
| Swimming crab | 20 QTLs for sex | 3 | (Lv et al., 2018) |
| 2 QTLs for salinity tolerance | 79 | (Lv et al., 2019) | |
| Mud carb | 27 QTLs for growth, 2 QTLs for sex | 13 | (Waiho et al., 2019) |
| Pacific oyster | 41 QTLs for growth | 17 | (Li et al., 2018) |
| 6 QTLs associated with orange shell color, 1 QTL for sex | (Han et al., 2021) | ||
| 2 QTLs for growth | 4 | (Kho et al., 2021) | |
| Black carp | 17 QTLs for growth | (Guo et al., 2022) | |
| 19 QTLs for growth and cold tolerance traits | 6 | (Zhang et al., 2023) | |
| South African abalone | 5 QTLs of growth | 8 | (Tshilate et al., 2024) |
| Tiger puffer | 14 QTLs for growth | (Liu et al., 2022 b) | |
| Dusky kob | 5 QTLs for of growth | 11 | (Jackson and Rhode, 2024) |
| Pacific white shrimp | 11 QTLs for growth | 4 | (Chen et al., 2024) |
Aquaculture species for which commercial high-density SNP chips have been recently developed
| Species | SNP array platform | Density | Aim | Reference |
|---|---|---|---|---|
| 1 | 2 | 3 | 4 | 5 |
| Atlantic salmon | Affymetrix Axiom | Comparing genomic signatures of domestication in two populations | (López et al., 2019) | |
| Affymetrix Axiom | 17 K | GWAS and GS for amoebic gill disease (AGD) resistance | (Robledo et al., 2018 a) | |
| Affymetrix Axiom | 50 K | Reveal genetic relationship and chromosomal fusions to contribute to local adaptation | (Wellband et al., 2019) | |
| GWAS for age at maturity | (Sinclair-Waters et al., 2020) | |||
| Channel catfish | GWAS to identify intra-specific QTL associated with resistance to enteric septicemia disease | (Shi et al., 2018) | ||
| Nile tilapia | 50 K | Linkage maps to improve selection accuracy of sex determination | (Joshi et al., 2018) | |
| 58 K | Determination of the genetic basis of complex traits and genomic selection in this species | (Yáñez et al., 2020) | ||
| 17 K | Whole-genome pooled sequencing (Poolseq) approach | Population structure analysis and relationship resolution within Nile tilapia populations | (Barría et al., 2023) | |
| Rainbow trout | Affymetrix Axiom | 665 K | Genome-wide genotyping | (Bernard et al., 2022) |
| Compare the accuracy of GEBVs of P-BLUP to ssGBLUP for infectious pancreatic necrosis virus disease resistance | (Yoshida et al., 2019) | |||
| Affymetrix Axiom | 32 K / 35 K | LD analysis / GS for BCWD resistance | (Vallejo et al., 2018) | |
| 50 K | GWAS to identify QTL associated with muscle yield | (Salem et al., 2018) | ||
| Affymetrix Axiom | 23 K | GS for growth trait | (Gutierrez et al., 2018) | |
| – | 50 K | To investigate phenotype–genotype associations and determine the genomic basis of economically important traits | (Yáñez et al., 2023) | |
| – | 1.3 M | Identification of selective breeding signatures | (Cádiz et al., 2020), | |
| Pacific abalone | – | 56 K | QTL mapping and detection of candidate genes of growth-related traits | (Kho et al., 2021) |
| Affymetrix Axiom | 40 K | Development and validation of multiple SNP array for Pacific abalone | (Liu et al., 2022 a) | |
| Sea bass | ThermoFisher Axiom | 57 K | GWAS for resistance to viral nervous necrosis | (Griot et al., 2021) |
| Large yellow croaker | 55 K | Development of first liquid SNP array | (Wang et al., 2023 a) | |
| 600 K | Development of first high-throughput genotyping SNP array | (Zhou et al., 2020) | ||
| 55 K | Development and evaluation of a SNP array for genomic selection | (Zhou et al., 2022) | ||
| Eastern oyster | Affymetrix Axiom | 200 K | Development and evaluation of high density SNP array | (Xuereb et al., 2023) |
| Affymetrix Axiom | 566 K | Development and evaluation of high density SNP array | (Guo et al., 2023) | |
| Chinese tongue | Affymetrix Axiom | 38 K | Genomic selection for resistance to Vibrio harveyi resistance | (Lu et al., 2021) |
| Arctic charr | Affymetrix Axiom | 86 K | Genome-wide variation studies | (Nugent et al., 2019) |
| Sea cucumber | Affymetrix Axiom | 24 K | Genomic selection with MCP regularized deep neural networks | (Lv et al., 2022) |
| Japanese flounder | Affymetrix Axiom | 50 K | Genotyping and selective breeding programs for economically important traits | (Zhou et al., 2021) |
| Mud crab | Affymetrix Axiom | 40 K | GWAS for growth trait | (Ye et al., 2024) |
Successful applications of genome editing in aquaculture species
| Species | Target gene | Method | Trait of interest | Notable features | Reference |
|---|---|---|---|---|---|
| Atlantic salmon | elov-2 | CRISPR/Cas9 | Omega-3 metabolism | (Datsomor et al., 2019) | |
| slc45a2, tyr | CRISPR/Cas9 | Pigmentation | (Straume et al., 2020) | ||
| dnd (slc45a2) | CRISPR/Cas9 | Reproduction and development | (Güralp et al., 2020) | ||
| Tilapia | rln3a, rln3b | CRISPR/Cas9 | Reproduction and development | (Yang et al., 2020 b) | |
| igf3 | CRISPR/Cas9 | Reproduction and development | (Li et al., 2020) | ||
| esr1, esr2a, esr2b | CRISPR/Cas9 | Reproduction and development | (Yan et al., 2019) | ||
| Amh homozygous, amh2 homozygous, amh heterozygous, amhr2 heterozygous | CRISPR/Cas9 | Reproduction and development | (Liu et al., 2020 b) | ||
| Cyb11c1 | CRISPR/Cas9 | Reproduction and development | (Zheng et al., 2020) | ||
| Red sea bream | mstn | CRISPR/Cas9 | Feed intake | (Kishimoto et al., 2018) | |
| gonadotropin-releasing hormone gene | TALENs | Reproduction | Plasmids targeting | (Qin et al., 2022) | |
| Catfish | lh, mc4r, mstn1 and mstn2 | CRISPR/Cas9 | Growth | (Wang et al., 2024 c) | |
| Channel catfish | mc4r | CRISPR/Cas9 | Growth | Fatty acid synthesis | (Coogan, et al., 2022) |
| cathelicidin gene (As-Cath) | CRISPR/Cas9 | Disease resistance | Generation f eco-friendly disease-resistant transgenic catfish | (Wang et al., 2024 b) | |
| Striped catfish | dead end 1 (dnd1) | CRISPR/Cas9 | Sex control | Generation of sterile fish | (Booncherd et al., 2024) |
| Yellow catfish | mstna | Zinc finger nucleases | Growth | Increased mass and weight | (Zhang et al., 2020 b) |
| Common carp | MCIR | CRISPR/Cas9 | Pigmentation | (Mandal et al., 2020) | |
| ASIP | CRISPR/Cas9 | Pigmentation | (Chen et al., 2019 b) | ||
| cyp17a1 | CRISPR/Cas9 | Sex reversal | Production of a monosex of common carp | (Zhai et al., 2022) | |
| Grass carp | gcjam-a | CRISPR/Cas9 | Disease resistance | (Ma et al., 2018) | |
| Rainbow trout | igfbp-2b1/2b2 | CRISPR/Cas9 | Growth | (Cleveland et al., 2020) | |
| gata2b, tcnba | CRISPR/Cas9 | Sex control | (Carrington et al., 2022) | ||
| Pacific oyster | mstn | CRISPR/Cas9 | Growth | (Yu et al., 2019) | |
| CgMELC | CRISPR/Cas9 | Muscle growth | Larval muscle contraction and myogenesis | (Li et al., 2021) | |
| Oriental prawn | EcNinaB-X1 | CRISPR/Cas9 | Immune defense | Reduced prawns mortality | (Sun et al., 2020) |
| Medaka | kitlga | CRISPR/Cas9 | Pigmentation | Regulation of melanogenesis Melanophore proliferation and migration | (Otsuki et al., 2020) |
| Giant freshwater prawn | MroDmrt11E | RNAi | Sex control | Production of all-male monosex freshwater prawn | (Xu and Ma, 2022) |
| Tiger puffer | dead end 1 (dnd1) | CRISPR/Cas9 | Sex control | Surrogate production | (Yoshikawa et al., 2024) |
| Channel catfish | elongase gene | CRISPR/Cas9 | Growth | Enhancement of nutritional quality of catfish | (Coogan et al., 2023) |
| antimicrobial peptide gene | CRISPR/Cas9 | Disease resistance | Bacterial resistance | (Wang et al., 2023 b) | |
| ticam1/rbl | CRISPR/Cas9 | Immunity | (Elaswad et al., 2018) | ||
| Blue catfish | Alligator cathelicidin gene | CRISPR/Cas9 | Disease resistance | Bacterial resistance | (Wang et al., 2023 c) |
Genome assembly and gene annotation of major aquaculture species
| Species name | Assembled size (Mb) | Genome size (Mb) | Number of genes | N50 scaffold | Reference |
|---|---|---|---|---|---|
| Chinese tapertail anchovy | 870.0 | 870 | 20,837 | 2.1 | (Xu et al., 2020 b) |
| Labeo catla | 1010 | 1.01 Gb | 25,812 | 0.7 Mb | (Sahoo et al., 2020) |
| Yellow catfish | 732.8 | – | 24,522 | 1.1 Mb | (Gong et al., 2018) |
| Large yellow croaker | 189.3 | 723.86 | 23,657 | 2.83 | (Chen et al., 2019 a) |
| N/A | 669.78 | 26,100 | 6.55 Mb | (Mu et al., 2018) | |
| Snout otter clam | 544 | 26,380 | 2.14 Mb | (Thai et al., 2019) | |
| Pacific oyster | 283 | 587 | 26,811 | 581 kb | (Wang et al., 2019 b) |
| Chinese shrimp | 147 | 1,384.88 | 25,026 | 36.87 | (Wang et al., 2022 c) |
| Australian black tiger | 31,922 | 1.89 | 35,517 | 496,398 | (Huerlimann et al., 2022) |
| Kuruma shrimp | 15,969 | 1,700 | 26,381 | 234.9 kbp | (Kawato et al., 2021) |
| Oriental river prawn | 4,500 | 2,933 | 44,086 | 86.8 Mb | (Jin et al., 2021) |
| Giant grouper | 1.128 Gb | 999.69 | 24,794 | 76,419 | (Wang et al., 2019 a) |
| Potato grouper | – | 1.13 Gb | 435 | 42.65 Mb | (Wang et al., 2022 a) |
| Red-spotted grouper | 1.135 Gb | 106.29 Gb | 23,923 | 46.03 Mb | (Ge et al., 2019) |
| Yellow perch | 877.4 | – | – | 37.4 | (Feron et al., 2020) |
| Hard-shell mussel | 1.57 Gb | – | 37,478 | 1.49 Mb | (Yang et al., 2021) |
| Rohu carp | 1480 | 1.5 Gb | 26,400 | 1.95 | (Das et al., 2020) |
| Indian catfish | 941 | 1.02 G | 23,748 | 1.3 | (Kushwaha et al., 2021) |
| Spiny red gurnard | 624.7 Mb | 637.64 Mb | 25,358 | 28.11 Mb | (Wang et al., 2023 e) |
| Channel catfish | – | 1.01 Gb | 950 | 26.7 Mb | (Bao et al., 2019) |
| Striped catfish | 788.4 Mb | 713.9 Mb | 21.8 Mb | (Hai et al., 2022) | |
| Tilapia | 1,007 Mb | 28,902 | 11.38 Mb | (Tao et al., 2021) | |
| Yellow perch | 877.4 Mb | 16,579 | 37.4 Mb | (Feron et al., 2020) | |
| Swamp eel | 799 Mb | 22,373 | 67.24 Mb | (Tian et al., 2021) | |
| Humpback grouper | 1.08 Gb | 24,442 | 43.78 Mb | (Liu et al., 2024) | |
| Acrossocheilus fasciatus | 879.52 Mb | 24,900 | 32.7 Mb | (Zheng et al., 2024 a) | |
| Topmouth culter | 1.052 Gb | 28,228 | 43.09 Mb | (Zhao et al., 2024) | |
| Black tiger shrimp | 2.39 Gb | 30,038 | (Uengwetwanit et al., 2021) | ||
| Kuruma shrimp | 665.19 Gb | 1.54 Gb | 24,317 | 38.27 Mb. | (Ren et al., 2022) |
| Ridgetail white shrimp | 5.86 Gb | 44, 288 | 138.24 Mb | (Wang et al., 2024 a) | |
| Giant river prawn | 3.18 Gb | 17,436 | 62.73 Mb | (Zheng et al., 2024 b) | |
| Pacific white shrimp | 1.87 Gb | 24,861 | 39.7 Mb | (Peng et al., 2023) | |
| Swimming crab | 125.99 Gb | 1.47 Gb | 25,026 | 36.87 Mb | (Tang et al., 2020) |
| Chum salmon | 2.6 Gb | 40,661 | 2 Mbp | (Rondeau et al., 2023) |