Over the past few decades, genomics has undergone profound evolution. What began as a scientific discipline, striving to obtain initial glimpses, has now grown into global scientific endeavors with the potential to sequence genomes across all extant life forms. The advent of next generation sequencing, genotyping platforms, and the concurrent development of bioinformatics tools have facilitated extensive genomic research across populations and species. Genomics is rapidly making a tremendous impact on aquaculture, with thousands of publications covering relevant aquaculture research domains every year. This is further demonstrated by the significant growth of the industry in recent decades. Aquaculture has achieved a record production of 223.2 million tons in 2022 (FAO, 2023). It is projected to provide 60% of the fish supply by 2030 (FAO, 2022), positioning it as the primary source of fish proteins (Naylor et al., 2021). However, challenges such as inadequate genetically improved stocks, emerging diseases, low feed conversion rates, various environmental stressors, and slow growth rates necessitate a greater emphasis on genomic applications. Integrating genomics-informed management is essential for aquaculture and fisheries to meet sustainable food security challenges as global demand for high-quality protein sources rises.
To date, substantial advancements have been achieved in genomics for numerous aquaculture species, encompassing the development of high-quality whole-genome sequences, transcriptome databases, microarrays, and molecular markers, mainly single nucleotide polymorphisms (SNP), followed by the discovery of high-density linkages and physical maps (Houston et al., 2020). These resources have enabled comprehensive data utilization to solve many complex problems in several fields (Figure 1). For example, the incorporation of genome-based technologies into selection breeding schemes through advanced genomic selection (GS), genome-wide association studies (GWAS), and marker-assisted selection (MAS) has revolutionized aquaculture breeding for many important species (Gjedrem and Rye, 2018). More dynamically, gene editing has been a subject of intensive research and has shown a heightening role in improving genetic advances in aquaculture (Gutási et al., 2023). Similarly, high-resolution transcriptome-wide profiling and miRNA analysis have been employed in aquaculture research to identify key transcripts, potential genes, and ncRNAs involved in growth, reproduction, stress responses, immunity, and disease. Emerging genomic methodologies in fisheries management, including genome scans and eDNA, provide insights into population genetic structure, phylogenetics, and local adaptation, advancing biodiversity assessments and aquatic ecosystem conservation (Huang et al., 2022; Chouhan et al., 2023). Metagenomic analysis, a powerful tool for analyzing complex microbial genomes, is increasingly used in aquaculture to detect novel and potential pathogen antibiotic resistance genes, microbial roles in microcosms, microbial communities forming bioflocs, and probiotics (Xu et al., 2020 a). Epigenomic research increasingly demonstrates its significant impact on aquaculture, influencing many aspects of biological processes such as growth, reproduction, nutrition, and disease (Roy et al., 2022; Navarro-Martín et al., 2023; Wang and Shen, 2023). Recent single-cell genomic studies in non-model aquatic species have shown promise in advancing our understanding of cellular biology in aquaculture, offering potential solutions to sustainability challenges, particularly the critical need to control disease outbreaks (Daniels et al., 2023). Figure 2 illustrated some major events and progress in aquaculture genomics, genetics, and breeding research.
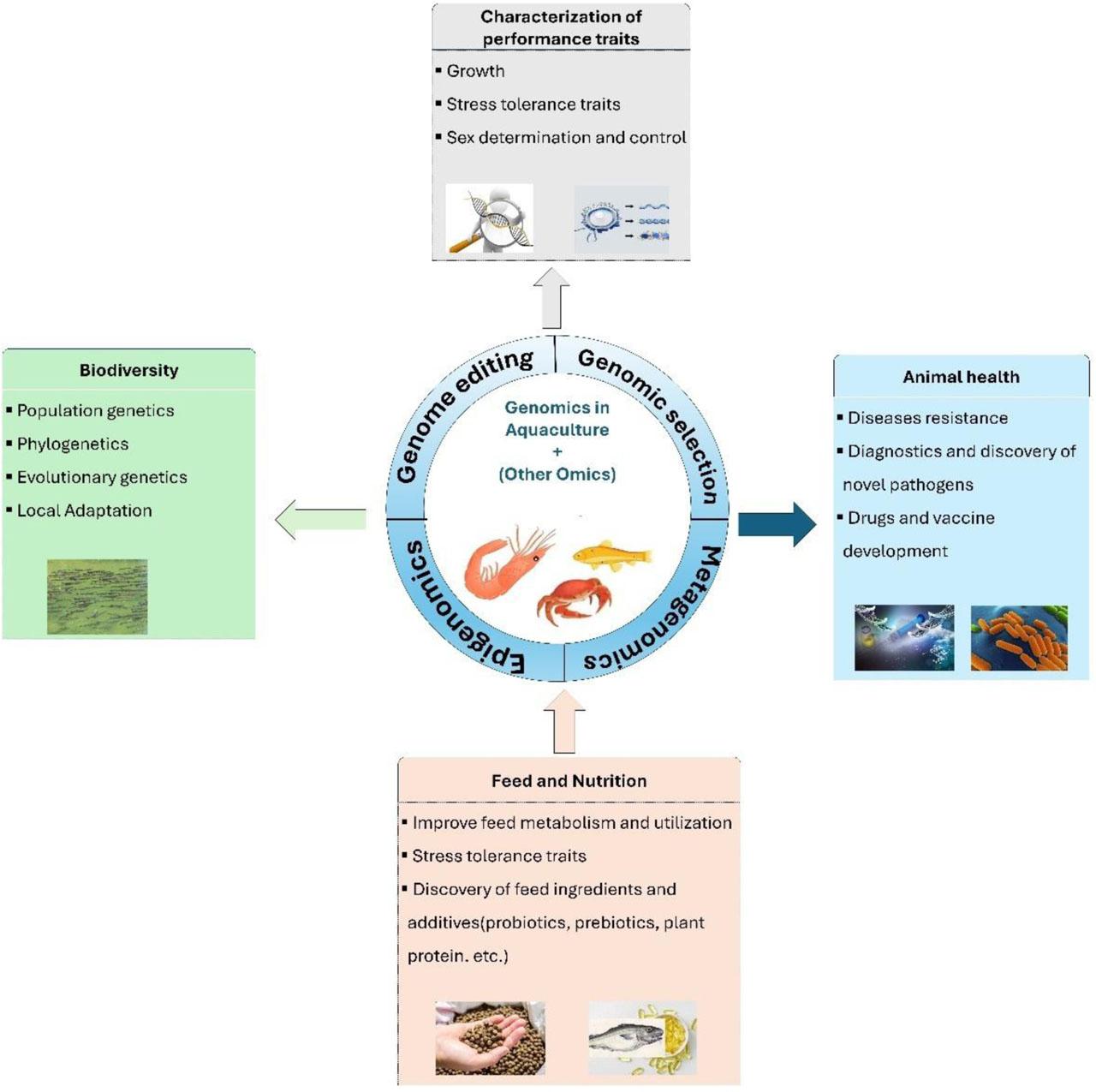
An overview of applications of genomics technology in aquaculture
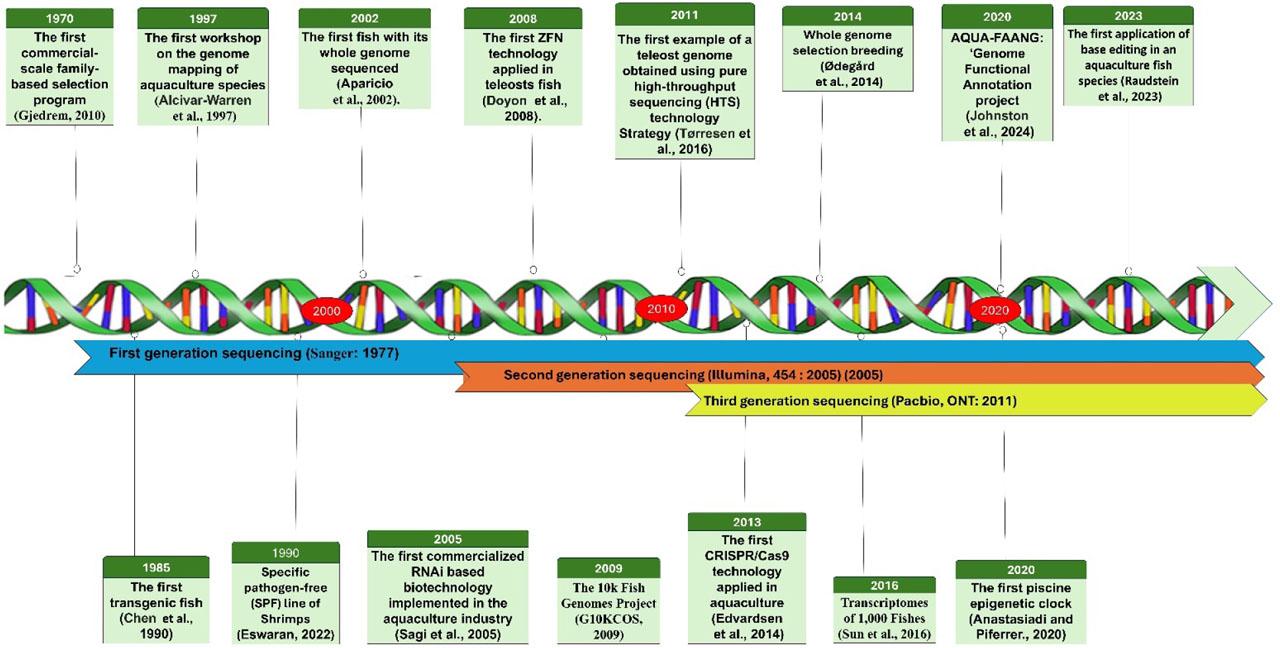
Major events in aquaculture genetics, genomics, and breeding
Although substantial progress has been made in applying genomics to aquaculture, the full potential of this field remains to be explored and realized (Canário, 2019; Friedman et al., 2022). Current awareness of genomic resource utilization is insufficient, and misinformation is prevalent, leading to an increased demand for guidance on genomic resource utilization. Significant changes in established practices, research, innovation processes, communication, and resulting policies are essential for the secure and organized use of genomic technologies and their products. This review provides an overview of the status of genomics in aquaculture practices. It highlights the available opportunities, including discussing future priorities for technologies, auspicious science, and governmental policies to realize the fullest benefit of genomic advancement in improving the aquaculture industry.
Aquaculture genomics emerged as a distinct discipline in the 1990s, marked by the inaugural workshop on genome mapping in Dartmouth, Massachusetts in 1997, which facilitated the genomic mapping of five key aquaculture species: Pacific whiteleg shrimp (Litopenaeus vannamei), Atlantic salmon (Salmo salar), channel catfish (Ictalurus punctatus), Nile tilapia (Oreochromis niloticus), and the Pacific oyster (Crassostrea gigas) (Alcivar-Warren et al., 1997). Subsequently, there has been a significant increase in the sequencing capacity and speed, accompanied by several genomic and transcriptomic projects. Notable examples include the Cetacean Genomes Project (Morin et al., 2020), Fish10K (The 10k Fish Genomes Project) (G10KCOS, 2009), and Fish-T1K (transcriptomes of 1,000 fish) (Sun et al., 2016). Recently, the Institute of Hydrology, the Chinese Academy of Science, and collaborators launched the Fish 10,000 Genomes Project (Fan et al., 2020). This project intends to sequence 10,000 fish species worldwide and to create a massive, high-quality genomic database. The advent of high-throughput sequencing (HTS) technologies and decreasing costs have led to a rapid increase in whole-genome sequencing projects on aquaculture species, frequently carried out by individual research groups with technical support from private companies. Table 1 provides a summary of recent reference genomes for several commercially significant aquaculture species.
Genome assembly and gene annotation of major aquaculture species
| Species name | Assembled size (Mb) | Genome size (Mb) | Number of genes | N50 scaffold | Reference |
|---|---|---|---|---|---|
| Chinese tapertail anchovy | 870.0 | 870 | 20,837 | 2.1 | (Xu et al., 2020 b) |
| Labeo catla | 1010 | 1.01 Gb | 25,812 | 0.7 Mb | (Sahoo et al., 2020) |
| Yellow catfish | 732.8 | – | 24,522 | 1.1 Mb | (Gong et al., 2018) |
| Large yellow croaker | 189.3 | 723.86 | 23,657 | 2.83 | (Chen et al., 2019 a) |
| N/A | 669.78 | 26,100 | 6.55 Mb | (Mu et al., 2018) | |
| Snout otter clam | 544 | 26,380 | 2.14 Mb | (Thai et al., 2019) | |
| Pacific oyster | 283 | 587 | 26,811 | 581 kb | (Wang et al., 2019 b) |
| Chinese shrimp | 147 | 1,384.88 | 25,026 | 36.87 | (Wang et al., 2022 c) |
| Australian black tiger | 31,922 | 1.89 | 35,517 | 496,398 | (Huerlimann et al., 2022) |
| Kuruma shrimp | 15,969 | 1,700 | 26,381 | 234.9 kbp | (Kawato et al., 2021) |
| Oriental river prawn | 4,500 | 2,933 | 44,086 | 86.8 Mb | (Jin et al., 2021) |
| Giant grouper | 1.128 Gb | 999.69 | 24,794 | 76,419 | (Wang et al., 2019 a) |
| Potato grouper | – | 1.13 Gb | 435 | 42.65 Mb | (Wang et al., 2022 a) |
| Red-spotted grouper | 1.135 Gb | 106.29 Gb | 23,923 | 46.03 Mb | (Ge et al., 2019) |
| Yellow perch | 877.4 | – | – | 37.4 | (Feron et al., 2020) |
| Hard-shell mussel | 1.57 Gb | – | 37,478 | 1.49 Mb | (Yang et al., 2021) |
| Rohu carp | 1480 | 1.5 Gb | 26,400 | 1.95 | (Das et al., 2020) |
| Indian catfish | 941 | 1.02 G | 23,748 | 1.3 | (Kushwaha et al., 2021) |
| Spiny red gurnard | 624.7 Mb | 637.64 Mb | 25,358 | 28.11 Mb | (Wang et al., 2023 e) |
| Channel catfish | – | 1.01 Gb | 950 | 26.7 Mb | (Bao et al., 2019) |
| Striped catfish | 788.4 Mb | 713.9 Mb | 21.8 Mb | (Hai et al., 2022) | |
| Tilapia | 1,007 Mb | 28,902 | 11.38 Mb | (Tao et al., 2021) | |
| Yellow perch | 877.4 Mb | 16,579 | 37.4 Mb | (Feron et al., 2020) | |
| Swamp eel | 799 Mb | 22,373 | 67.24 Mb | (Tian et al., 2021) | |
| Humpback grouper | 1.08 Gb | 24,442 | 43.78 Mb | (Liu et al., 2024) | |
| Acrossocheilus fasciatus | 879.52 Mb | 24,900 | 32.7 Mb | (Zheng et al., 2024 a) | |
| Topmouth culter | 1.052 Gb | 28,228 | 43.09 Mb | (Zhao et al., 2024) | |
| Black tiger shrimp | 2.39 Gb | 30,038 | (Uengwetwanit et al., 2021) | ||
| Kuruma shrimp | 665.19 Gb | 1.54 Gb | 24,317 | 38.27 Mb. | (Ren et al., 2022) |
| Ridgetail white shrimp | 5.86 Gb | 44, 288 | 138.24 Mb | (Wang et al., 2024 a) | |
| Giant river prawn | 3.18 Gb | 17,436 | 62.73 Mb | (Zheng et al., 2024 b) | |
| Pacific white shrimp | 1.87 Gb | 24,861 | 39.7 Mb | (Peng et al., 2023) | |
| Swimming crab | 125.99 Gb | 1.47 Gb | 25,026 | 36.87 Mb | (Tang et al., 2020) |
| Chum salmon | 2.6 Gb | 40,661 | 2 Mbp | (Rondeau et al., 2023) |
Aquaculture research employs various sequencing technologies, including Sanger sequencing, pyrosequencing, Ion Tor-rent, Oxford Nanopore, and single-molecule real-time sequencing. During the last decade, long-read sequencing using PacBio SMRT and ONT platforms has been applied to generate more contiguous and complete assemblies of various teleosts with complex genomes characterized by duplication events and repetitive genomic regions, including channel catfish (Ictaluru punctuatus) (Bao et al., 2019), southern catfish (Silurus meridionalis) (Zheng et al., 2022 b), striped catfish (Pangasianodon hypophthalmus) (Hai et al., 2022), Nile tilapia (Oreochromis niloticus) (Tao et al., 2021), and yellow perch (Perca flavescens) (Feron et al., 2020). Nevertheless, short reads are crucial for determining genome size and refining long reads (Wong et al., 2022), and hybrid assembly methods combining short- and long-read sequencing have become more prevalent in aquaculture research in recent times and have been proven to enhance the precision and resolution of genome assemblies (Wong et al., 2022). Combined assembly approaches, utilizing two or more sequencing technologies, have been applied to generate draft genomes with higher quality assemblies in various aquatic species, such as the swamp eel (Monopterus albus) (Tian et al., 2021), humpback grouper (Chromileptes altivelis) (Liu et al., 2024), Pacific whiteleg shrimp (Peng et al., 2023), Chinese shrimp (Fenneropenaeus chinensis) (Wang et al., 2022 c), and oriental river prawn (Macrobrachium nipponense) (Jin et al., 2021), which serve as reference genomes, invaluable for selective breeding programs for these important species. Recently, a multiplatform sequencing strategy was applied to sequence and assemble the genomes of several aquaculture species (Yang et al., 2023 a).
The availability of high-quality reference genomes has revolutionized aquaculture genetics by enabling the development of sophisticated molecular markers through diverse genomic approaches; these include targeted sequencing, expressed sequence tag (EST) data mining, bacterial artificial chromosome (BAC) end sequencing, and whole genome sequencing (WGS), each contributing uniquely to genome characterization and quantitative trait locus (QTL) mapping. SNPs are increasingly used as the marker of choice due to their genome-wide distribution, ease of detection through modern sequencing technologies, and cost-effectiveness in large-scale genotyping applications. In the last two decades, the aquaculture research community has identified numerous cost-effective methodologies for genotyping thousands of SNPs with data quality. These include genotyping by sequencing (GBS), which offers a streamlined protocol for library preparation and the ability to interrogate thousands of loci simultaneously (Kho et al., 2021); whole-genome resequencing (WGR), which provides comprehensive genomic coverage for detailed population analyses (Cádiz et al., 2020; Yáñez et al., 2020); and restriction site-associated DNA sequencing (RAD-seq), which delivers cost-effective solutions for population genetics studies (Wu et al., 2018; Vendrami et al., 2019). GBS in particular has gained prominence due to its operational simplicity, economic viability (Robledo et al., 2018 b), analytical reliability (Bian et al., 2020), and capacity for high-multiplex marker analysis (Zhang et al., 2020 a). These approaches have proven valuable in developing population-specific or species-diagnostic SNPs for several commercially important aquaculture species. Over recent years, an increasing number of SNPs and QTLs and for traits like growth performance, disease resistance, and environmental stress tolerance have been uncovered in aquaculture species (Tables 2 and 4). As sequencing costs continue to decline and genomic tools become more refined, analysis of structural variants (SVs) including deletions, duplications, inversions, and translocations is increasingly being recognized as a powerful complement to traditional SNP array genotyping in aquaculture genetic studies. The integration of SV analysis with existing SNP datasets facilitates species-wide genetic diversity analysis across populations and closely related species, offering enhanced resolution for characterizing large-effect variants at unprecedented scale and accuracy, even in complex aquaculture genomes with high repeat content. For instance, in salmonid, SV mapping has helped elucidate the genomic basis and architecture of local adaptation (Tigano and Russello, 2022; Lecomte et al., 2024), while in oysters and shrimps, SV detection has identified genomic regions associated with rapid growth and disease resilience (Modak et al., 2021; Taengchaiyaphum et al., 2024).
Aquaculture species for which commercial high-density SNP chips have been recently developed
| Species | SNP array platform | Density | Aim | Reference |
|---|---|---|---|---|
| 1 | 2 | 3 | 4 | 5 |
| Atlantic salmon | Affymetrix Axiom | Comparing genomic signatures of domestication in two populations | (López et al., 2019) | |
| Affymetrix Axiom | 17 K | GWAS and GS for amoebic gill disease (AGD) resistance | (Robledo et al., 2018 a) | |
| Affymetrix Axiom | 50 K | Reveal genetic relationship and chromosomal fusions to contribute to local adaptation | (Wellband et al., 2019) | |
| GWAS for age at maturity | (Sinclair-Waters et al., 2020) | |||
| Channel catfish | GWAS to identify intra-specific QTL associated with resistance to enteric septicemia disease | (Shi et al., 2018) | ||
| Nile tilapia | 50 K | Linkage maps to improve selection accuracy of sex determination | (Joshi et al., 2018) | |
| 58 K | Determination of the genetic basis of complex traits and genomic selection in this species | (Yáñez et al., 2020) | ||
| 17 K | Whole-genome pooled sequencing (Poolseq) approach | Population structure analysis and relationship resolution within Nile tilapia populations | (Barría et al., 2023) | |
| Rainbow trout | Affymetrix Axiom | 665 K | Genome-wide genotyping | (Bernard et al., 2022) |
| Compare the accuracy of GEBVs of P-BLUP to ssGBLUP for infectious pancreatic necrosis virus disease resistance | (Yoshida et al., 2019) | |||
| Affymetrix Axiom | 32 K / 35 K | LD analysis / GS for BCWD resistance | (Vallejo et al., 2018) | |
| 50 K | GWAS to identify QTL associated with muscle yield | (Salem et al., 2018) | ||
| Affymetrix Axiom | 23 K | GS for growth trait | (Gutierrez et al., 2018) | |
| – | 50 K | To investigate phenotype–genotype associations and determine the genomic basis of economically important traits | (Yáñez et al., 2023) | |
| – | 1.3 M | Identification of selective breeding signatures | (Cádiz et al., 2020), | |
| Pacific abalone | – | 56 K | QTL mapping and detection of candidate genes of growth-related traits | (Kho et al., 2021) |
| Affymetrix Axiom | 40 K | Development and validation of multiple SNP array for Pacific abalone | (Liu et al., 2022 a) | |
| Sea bass | ThermoFisher Axiom | 57 K | GWAS for resistance to viral nervous necrosis | (Griot et al., 2021) |
| Large yellow croaker | 55 K | Development of first liquid SNP array | (Wang et al., 2023 a) | |
| 600 K | Development of first high-throughput genotyping SNP array | (Zhou et al., 2020) | ||
| 55 K | Development and evaluation of a SNP array for genomic selection | (Zhou et al., 2022) | ||
| Eastern oyster | Affymetrix Axiom | 200 K | Development and evaluation of high density SNP array | (Xuereb et al., 2023) |
| Affymetrix Axiom | 566 K | Development and evaluation of high density SNP array | (Guo et al., 2023) | |
| Chinese tongue | Affymetrix Axiom | 38 K | Genomic selection for resistance to Vibrio harveyi resistance | (Lu et al., 2021) |
| Arctic charr | Affymetrix Axiom | 86 K | Genome-wide variation studies | (Nugent et al., 2019) |
| Sea cucumber | Affymetrix Axiom | 24 K | Genomic selection with MCP regularized deep neural networks | (Lv et al., 2022) |
| Japanese flounder | Affymetrix Axiom | 50 K | Genotyping and selective breeding programs for economically important traits | (Zhou et al., 2021) |
| Mud crab | Affymetrix Axiom | 40 K | GWAS for growth trait | (Ye et al., 2024) |
Complementing these DNA-based approaches, transcriptome sequencing (RNA-Seq) remained a powerful tool for functional SNP discovery, particularly for identifying coding variants associated with economically important traits. This approach has proven exceptionally valuable for developing diagnostic SNP panels in key aquaculture species.
The implementation of genomics tools in aquaculture breeding has marked a revolutionary shift in genetic improvement strategies, offering a powerful complement to traditional selective breeding strategies. Genome-enabled breeding begins with the identification of genetic variants associated with production traits of interest. Currently, whole-genome association analysis (GWAS) have been the most commonly used and reliable approach for screening markers linked to quantitative trait loci (QTL) into aquaculture breeding programs. The resulting GWAS datasets serve as foundational resources for breeding programs, including genomic selection (GS) and marker-assisted selection (MAS), thereby accelerating genetic improvement programs. For MAS, a landmark success story is the discovery of a major QTL controlling 80% of the genetic variance for infectious pancreatic necrosis virus (IPNV) resistance in Atlantic salmon, which significantly reduced disease-related mortality in commercial populations (Gutierrez et al., 2018). Moreover, the identification of sex-linked markers in species like blue tilapia (Oreochromis aureus) has enabled the development of mono-sex breeding lines, which have shown considerable economic benefits (Wu et al., 2021).
Unlike MAS, which often depends on the phenotypic expression of traits, GS leverages dense genome-wide marker data capturing both additive and non-additive genetic effects that enables the identification of genetic markers that can predict genomic breeding values (GEBVs), of individuals even before the traits are expressed. This predictive power greatly accelerates breeding programs, making GS more efficient and precise, particularly effective for complex, polygenic traits like disease resistance, growth, feed conversion efficiency, and environmental adaptability, which are crucial for the sustainability of the aquaculture industry. Despite the predictive accuracy of GS, widespread adoption is compromised by several limitations, including; genotype-environment interactions, limited diversity of the training population, and financial resources for genotyping and infrastructure development remain the most paramount challenges for the widespread adoption of GS. Current research indicates that integration of multi-omics data (transcriptomic, genomic, proteomic and epigenomics) significantly enhances prediction accuracy, enhancing our understanding of complex and polygenic traits by extrapolating genetic markers with phenotypic outcomes mainly through genotype-to-phenotype (G2P) associations. Notably, the convergence of AI/ML with multi-omics frameworks has demonstrated particular promise for complex trait dissection, as evidenced by recent applications in breeding programs of aquatic species including in channel catfish (Ott et al., 2024), striped bass (M. saxatilis) (Rajab et al., 2024), eastern oyster (Crassostrea virginica) (Proestou et al., 2023), rainbow trout (Ali et al., 2021), etc. Moving forward, the continued refinement of long-read sequencing platforms, coupled with computational advancement and artificial intelligence (AI), will further democratize GS in aquaculture, paving the way for next-generation precision breeding strategies.
In the last decade, advances in genome manipulation technologies including genome editing, gene transfer and RNA interference (RNAi) have opened new frontiers in aquaculture breeding through precise modifications in key genes (at both DNA and RNA levels) associated with economically important traits, offering unprecedented opportunities for genetic improvement (Wang et al., 2025). These technologies have been used to generate several improved fish and shell fish lines (Puthumana, et al., 2024), such as disease resistance (Wang and Shen, 2023, 2024 b), improved growth (Coogan et al., 2022; Wang et al., 2024 c), improved feed intake (Kishimoto et al., 2018), and sex control (Xu and Ma, 2022; Booncherd et al., 2024; Yoshikawa et al., 2024) (Table 3). These advancements hold the potential to significantly boost aquaculture breeding programs.
Successful applications of genome editing in aquaculture species
| Species | Target gene | Method | Trait of interest | Notable features | Reference |
|---|---|---|---|---|---|
| Atlantic salmon | elov-2 | CRISPR/Cas9 | Omega-3 metabolism | (Datsomor et al., 2019) | |
| slc45a2, tyr | CRISPR/Cas9 | Pigmentation | (Straume et al., 2020) | ||
| dnd (slc45a2) | CRISPR/Cas9 | Reproduction and development | (Güralp et al., 2020) | ||
| Tilapia | rln3a, rln3b | CRISPR/Cas9 | Reproduction and development | (Yang et al., 2020 b) | |
| igf3 | CRISPR/Cas9 | Reproduction and development | (Li et al., 2020) | ||
| esr1, esr2a, esr2b | CRISPR/Cas9 | Reproduction and development | (Yan et al., 2019) | ||
| Amh homozygous, amh2 homozygous, amh heterozygous, amhr2 heterozygous | CRISPR/Cas9 | Reproduction and development | (Liu et al., 2020 b) | ||
| Cyb11c1 | CRISPR/Cas9 | Reproduction and development | (Zheng et al., 2020) | ||
| Red sea bream | mstn | CRISPR/Cas9 | Feed intake | (Kishimoto et al., 2018) | |
| gonadotropin-releasing hormone gene | TALENs | Reproduction | Plasmids targeting | (Qin et al., 2022) | |
| Catfish | lh, mc4r, mstn1 and mstn2 | CRISPR/Cas9 | Growth | (Wang et al., 2024 c) | |
| Channel catfish | mc4r | CRISPR/Cas9 | Growth | Fatty acid synthesis | (Coogan, et al., 2022) |
| cathelicidin gene (As-Cath) | CRISPR/Cas9 | Disease resistance | Generation f eco-friendly disease-resistant transgenic catfish | (Wang et al., 2024 b) | |
| Striped catfish | dead end 1 (dnd1) | CRISPR/Cas9 | Sex control | Generation of sterile fish | (Booncherd et al., 2024) |
| Yellow catfish | mstna | Zinc finger nucleases | Growth | Increased mass and weight | (Zhang et al., 2020 b) |
| Common carp | MCIR | CRISPR/Cas9 | Pigmentation | (Mandal et al., 2020) | |
| ASIP | CRISPR/Cas9 | Pigmentation | (Chen et al., 2019 b) | ||
| cyp17a1 | CRISPR/Cas9 | Sex reversal | Production of a monosex of common carp | (Zhai et al., 2022) | |
| Grass carp | gcjam-a | CRISPR/Cas9 | Disease resistance | (Ma et al., 2018) | |
| Rainbow trout | igfbp-2b1/2b2 | CRISPR/Cas9 | Growth | (Cleveland et al., 2020) | |
| gata2b, tcnba | CRISPR/Cas9 | Sex control | (Carrington et al., 2022) | ||
| Pacific oyster | mstn | CRISPR/Cas9 | Growth | (Yu et al., 2019) | |
| CgMELC | CRISPR/Cas9 | Muscle growth | Larval muscle contraction and myogenesis | (Li et al., 2021) | |
| Oriental prawn | EcNinaB-X1 | CRISPR/Cas9 | Immune defense | Reduced prawns mortality | (Sun et al., 2020) |
| Medaka | kitlga | CRISPR/Cas9 | Pigmentation | Regulation of melanogenesis Melanophore proliferation and migration | (Otsuki et al., 2020) |
| Giant freshwater prawn | MroDmrt11E | RNAi | Sex control | Production of all-male monosex freshwater prawn | (Xu and Ma, 2022) |
| Tiger puffer | dead end 1 (dnd1) | CRISPR/Cas9 | Sex control | Surrogate production | (Yoshikawa et al., 2024) |
| Channel catfish | elongase gene | CRISPR/Cas9 | Growth | Enhancement of nutritional quality of catfish | (Coogan et al., 2023) |
| antimicrobial peptide gene | CRISPR/Cas9 | Disease resistance | Bacterial resistance | (Wang et al., 2023 b) | |
| ticam1/rbl | CRISPR/Cas9 | Immunity | (Elaswad et al., 2018) | ||
| Blue catfish | Alligator cathelicidin gene | CRISPR/Cas9 | Disease resistance | Bacterial resistance | (Wang et al., 2023 c) |
Aquaculture genomics studies aim to uncover the genetic factors influencing performance and production traits, which can be utilized in genetic improvement programs. Multiple published studies have shown promising results regarding genetic improvements in aquatic production, particularly for economic traits such as growth rate and disease resistance.
Growth is the most preferred economic trait of aquaculture species; therefore, it is one of the most studied traits in breeding programs (Ashton et al., 2019). Growth-related traits are quantitative and controlled by numerous genes across the genome (Omeka et al., 2022). Numerous recent investigations conducted in various aquaculture species have provided valuable insights into myostatin (mstn) mutations and their pivotal functions in regulating cell proliferation and muscle growth (Yan et al., 2022; Özcan Gökçek et al., 2023; Wu et al., 2023). In zebrafish, knockout of the Activin A receptor, type II (acvr2), and pro-opiomelanocortin (pomc) induces muscle fiber development and enhances growth performance (Che et al., 2023; Yang et al., 2023 b). In mollusks, Li et al. (2021) reported a noticeable impact on larval muscle myogenesis and sarcomeric arrangement of thin filaments in the MELC-CRISPR/Cas9 mutant group, indicating that MELC is involved in larval muscle contraction and myogenesis in oyster larvae. Recently, there has been growing interest in utilizing epigenetics to enhance the growth traits of aquatic animals, including fish and shellfish (Liu et al., 2022 c; Roy et al., 2022). To explore the mechanism behind the phenomenon of serious growth retardation problem in M. rosenbergii, Jiang et al. (2020) investigated the epigenetic variations that existed between “iron prawn” and normal prawn using the fluorescent-labeled methylation-sensitive amplified polymorphism method (F-MSAP). Recently, Konstantinidis et al. (2023) demonstrated hypomethylation in muscular tissues and tissue-specific differential methylation of genes related to somatic growth, such as growth hormones and their receptors.
Sexual dimorphism, describing systematic differences between males and females of the same species, is crucial in aquaculture research due to its impact on husbandry, management, and productivity (Liao et al., 2023). While the production of monosexual fish exhibiting enhanced growth rates holds significant economic importance, it necessitates the development of sex-specific markers or sex-related genes for efficient sex-reversal technology and effective methods of sex identification and differentiation (Li et al., 2022). Over the last two decades, various conventional techniques such as AFLPs, RAPD, and SSRs have been effectively employed to identify sex-specific sequences. Nevertheless, these approaches have become obsolete because of their low throughput and labor intensity (Liu et al., 2023). Recently, highly efficient cutting-edge genomic approaches have been developed to aid sex-specific marker screening. Examples include next-generation sequencing (NGS)-based and high-resolution melting (HRM) typing systems (Gao et al., 2020; Yang et al., 2020 a: Han et al., 2023), linkage mapping of the sex quantitative trait locus (QTL) or sex determination (SD) locus (Zhang et al., 2019 a; Liu et al., 2020 a), restriction-site associated DNA sequencing (RAD-seq) (Liang et al., 2019; Fang et al., 2020), type IIB restriction endonuclease-produced sequencing (2b-RAD-seq), double digestion restriction site-associated DNA sequencing (ddRAD-seq) (Brown et al., 2016), specific-locus amplified fragment sequencing (SLAF-seq) (Waiho et al., 2019), comparative transcriptome (RNA-seq) (Sun et al., 2018), and PCR amplification refractory mutation system (Xiao et al., 2020). A RAD-Seq computational analytic workflow called RADSex was recently developed by Feron et al. (2021) to identify sex-specific markers precisely and to study the genetic basis of sex determination. This workflow enables researchers to precisely detect sex-specific genomic sequences and XY sex determination systems in various fish species (Feron et al., 2021). Using these genomic tools, a batch of sex-determining master genes, such as Dnmt3aa, dmy, gsdf6a, amhr2, sdy, pfpdz1, and bcar1, were identified in the model and non-model fish species (Gong et al., 2018; Bao et al., 2019; Wang et al., 2021; Wei et al., 2023).
In recent years, the application of genome editing technology for sex control in aquaculture has attracted much attention worldwide. Qin et al. (2022) found that double electroporation of TALEN plasmids targeting the GnRH gene effectively sterilized channel catfish. In Chinese common carp, Neo-male was created by knockout of the cyp17a1 gene using CRISPR/Cas9 technology, and the all-female population was produced by crossing XX and XX female carp (Zhai et al., 2022). Currently, complete functional sex reversal has been achieved in M. rosenbergii. They produced neo-females (Tan et al., 2020) and all males (Xu and Ma, 2022) after silencing insulin-like androgenic gland hormone (IAG). IAG is a male endocrine organ secreted by the androgenic gland (AG) and plays a crucial role in regulating sexual differentiation and maturity in crustaceans (Levy and Sagi, 2020). Functional studies in various crustacean species have confirmed the vital involvement of the IAG in gonadal development and sex-related phenotypic characteristics. A recent RNAi experiment in crayfish Procambarus clarkii (Savaya et al., 2020) has provided insights into the possibility of manipulating IAG to produce a mono-sex population of fish species. Overall, large-scale monosex production has been achieved in various aquaculture species, significantly increasing aquaculture production.
In the past two decades, the aquaculture industry has witnessed a significant leap in integrative research aimed at bridging the mechanistic understanding of cellular and molecular stress responses in both natural ecosystems and aquaculture systems (Connon et al., 2018). Genomic tools have been pivotal enablers of these cross-scale efforts. The integration of genomic methodologies into physiological and ecological research has dramatically increased our insights into the identification of key genes, pathways, and epigenetic modifications associated with stress resilience. Consequently, the field has matured towards a more unified framework for understanding biological responses to various environmental stressors. Beyond fundamental genomics, the application of metagenomics combined with environmental DNA (eDNA) monitoring allows for real-time assessment of phenotypic plasticity and adaptive potential to environmental and anthropogenic drivers of habitat change. Blockchain technology is being explored to track stress-resistant genetic lines across supply chains and ensure transparency in breeding programs. Emerging technologies, such as artificial intelligence (AI) and portable sequencing devices, are poised to transform aquatic stress management further.
Progressive advancements from traditional genetic markers to genomics, where genome-wide sequences can be explored more effectively, have significantly increased the power of conservation genetics with unprecedented spatial resolution. Recent genomic-based studies have provided valuable insights into the molecular basis of long-standing questions, such as the domestication-related genomic signatures of Atlantic salmon (López et al., 2019), demographic history of smallmouth bass populations (Gunn et al., 2022), local adaptation of tiger shrimp (Vu et al., 2021), and detailed inbreeding depression in small yellow croaker (Larimichthys polyactis) (Zheng et al., 2022 a). Yang et al. (2020 d) used GBS technology to evaluate the population genetic structure of Sillago japonica along the coasts of Japan and China. They found significant genetic differentiation among populations, which differed from the mtDNA marker analysis. Recently, Vu et al. (2021) employed genome-wide SNP genotyping to evaluate the spatial population structure, local adaptation, and connectivity of tiger shrimp populations in the Indo-Pacific region. Their study provides novel genome-level insights directly affecting tiger shrimp aquaculture management. Genome-scan investigations have yielded novel insights into the genetic basis of temporal microevolutionary alterations in adaptive diversity resulting from fishery-induced evolution and climate change (Aramburu et al., 2023).
The ongoing development and application of the eDNA approach in conservation genomics has facilitated aquatic biodiversity surveys. It is primarily cost-effective and efficient (McColl-Gausden et al., 2021; Keck et al., 2022). Nelson-Chorney et al. (2019) developed a species-specific qPCR method by amplifying mtDNA from lake sediment eDNA to identify the colonization history of native and introduced trout (O. clarkia bouvieri) populations, thus providing an important tool for invasion assessment and management. Using eDNA metabarcoding, Zhang et al. (2022) examined the impact of urbanization and water quality variables on fish assemblages in lentic and lotic systems across Beijing. The results revealed distinct native and non-native fish diversity patterns, highlighting the ability of eDNA to resolve urban aquatic biodiversity. The utilization of genomic information in sustainable aquaculture resource management is increasingly being recognized, as evidenced by research programs such as FishPopTrace (Martinsohn et al., 2009), LabelFish (González Sotelo et al., 2014), and AquaTrace, which focus on food traceability, industry regulation, and risk assessments. Collectively, genomic tools offer a comprehensive understanding of aquaculture animal population genetics, local adaptation, and connectivity, potentially enhancing aquaculture and fishery practices.
The aquaculture sector has entered a new paradigm for disease management through strategic implementation of genomic technologies. These cutting-edge approaches have revolutionized the prevention, diagnosis, and control of aquatic diseases across production systems.
In recent years, WGS applications in aquaculture have expanded to encompass comprehensive characterization of microbial communities, including detailed analyses of resistomes, virulence, and mobilomes across diverse aquatic species (Tekedar et al., 2019; Qian et al., 2023; Salgueiro et al., 2023). As summarized in a recent review (Sciuto et al., 2022), several studies have utilized WGS and functional annotation to identify the major QTL that control disease resistance or susceptibility in many aquaculture species (see Table 4). Complementary transcriptomic analyses have further enhanced our understanding of host-pathogen dynamics by comparing gene expression profiles between susceptible and resistant phenotypes (Yang et al., 2019; Zhou et al., 2022), whereas single-cell RNA sequencing has provided unprecedented resolution in characterizing cellular responses to infection (Daniels et al., 2023). The integration of bulk segregant analysis with RNA-Seq (BSR-Seq) has proven particularly effective for identifying QTL-associated genes, as demonstrated in the studies of enteric septicemia (ESC) resistance in catfish (Jin et al., 2022). Moreover, metagenomics has emerged as a particularly powerful diagnostic tool, facilitating proactive and rapid pathogen surveillance (Taş et al., 2021) through the simultaneous sequencing of all nucleic acids present in environmental or biological samples. Metagenomics provides information regarding the diversity of microorganisms in specific areas and can reveal details regarding their functions and biological roles, even before they cause outbreaks (Tully et al., 2018). This approach has revealed complex microbial ecosystems in aquaculture environments, including the prevalence of clinically significant antimicrobial resistance genes, such as mcr, blaKPC, and blaCTX-M (Xu et al., 2020 a), as well as identifying emerging pathogenic threats through the detection of dysbiotic states in farmed fish populations (Haro-Moreno et al., 2020). Recently, Rasmussen et al. (2022) highlighted the potential of metagenomics to study the effects of bacterial synbiotics and probiotics on the microbiome of rainbow trout. The capacity of this technology to monitor microbiome alterations in response to therapeutic interventions, such as probiotic administration in rainbow trout (Rasmussen et al., 2022), has opened new avenues for microbial management. These genomic surveillance methodologies do not only provide critical early warning systems for disease outbreaks but also facilitate the tracking of antimicrobial resistance evolution challenges that directly inform the development of therapeutic interventions.
Identification of QTLs in aquaculture animals using linkage mapping
| Species | Number and traits of QTLs | Number of candidate genes | Reference |
|---|---|---|---|
| 1 | 2 | 3 | 4 |
| Atlantic salmon | 3 QTLs for sea lice resistance | 2 | (Robledo et al., 2018 a) |
| 2 QTLs for piscine myocarditis virus (PMCV) disease resistance | 6 | (Hillestad and Moghadam, 2019) | |
| 2 QTLs for cardiomyopathy syndrome resistance | 4 | (Boison et al., 2019) | |
| Common carp | 18 QTLs for head size | 10 | (Chen et al., 2018) |
| 1 QTL for koi herpesvirus (KHV) disease resistance | (Palaiokostas et al., 2018) | ||
| 17 QTL for feed conversion efficiency | (Zhang et al., 2021) | ||
| Grass carp | 4 QTLs for growth | 17 | (Huang et al., 2020) |
| Rainbow trout | 24 QTLs for Aeromonas salmonicida resistance | 33 | (Marana et al., 2021) |
| 21 QTLs for infectious hematopoietic necrosis disease resistance (IHN) | (Vallejo et al., 2019) | ||
| 5 QTLs for BCWD | (Fraslin et al., 2018) | ||
| 6 QTLs for ESC resistance | 37 | (Shi et al., 2018) | |
| 3 QTLs for ESC resistance | 55 | (Tan et al., 2018) | |
| Channel catfish | 6 QTLs for growth, 10 QTLs for sex | 25 | (Zhang et al., 2019 b) |
| Yangtze River common carp | 21 QTLs for growth, 4 QTLs for sex | 5 | (Feng et al., 2018) |
| Yellow river carp | 29 QTLs for growth | 3 | (Wang et al., 2022 b) |
| Largemouth bass | 32 QTLs for growth, 13 QTLs for sex | (Dong et al., 2019) | |
| Large yellow croaker | 7 QTLs for Cryptocaryon irritans disease resistance | 29 | (Kong et al., 2019) |
| Snapper | 4 QTLs for growth | 13 | (Ashton et al., 2019) |
| Pearl oyster | 32 QTLs for growth, 1 QTL for sex trait | 4 | (Liu et al., 2020 a) |
| Swimming crab | 20 QTLs for sex | 3 | (Lv et al., 2018) |
| 2 QTLs for salinity tolerance | 79 | (Lv et al., 2019) | |
| Mud carb | 27 QTLs for growth, 2 QTLs for sex | 13 | (Waiho et al., 2019) |
| Pacific oyster | 41 QTLs for growth | 17 | (Li et al., 2018) |
| 6 QTLs associated with orange shell color, 1 QTL for sex | (Han et al., 2021) | ||
| 2 QTLs for growth | 4 | (Kho et al., 2021) | |
| Black carp | 17 QTLs for growth | (Guo et al., 2022) | |
| 19 QTLs for growth and cold tolerance traits | 6 | (Zhang et al., 2023) | |
| South African abalone | 5 QTLs of growth | 8 | (Tshilate et al., 2024) |
| Tiger puffer | 14 QTLs for growth | (Liu et al., 2022 b) | |
| Dusky kob | 5 QTLs for of growth | 11 | (Jackson and Rhode, 2024) |
| Pacific white shrimp | 11 QTLs for growth | 4 | (Chen et al., 2024) |
The culmination of these genomic advancements has revolutionized aquaculture vaccinology, transforming the field from its early reliance on inactivated whole-organism vaccines (Ma et al., 2019) to contemporary genetic immunization strategies (Miccoli et al., 2021). While conventional vaccines remain constrained by safety concerns and variable efficacy across host species (Mohd-Aris et al., 2019), third-generation vaccines that leverage genomic insights now employ precisely targeted antigens with enhanced immunogenicity (Miccoli et al., 2021; Priya and Kappalli, 2022). DNA vaccines have demonstrated particular success against viral pathogens, including nodaviruses and rhabdoviruses, as well as against bacterial diseases (Embregts et al., 2019), with innovative delivery systems such as engineered probiotic vectors (Hua et al., 2021) and bacterial ghost platforms (Zheng et al., 2021) showing considerable promise. The regulatory approval of recombinant vaccines like Clynav against salmonid alphavirus-3 (SAV3) (Bedekar and Kole, 2022) marks a significant milestone in this technological transition, with current vaccine development programs encompassing diverse commercially important species ranging from salmonids to marine finfish (Robinson et al., 2023). These advances, grounded in fundamental genomic research and informed by comprehensive pathogen surveillance, are progressively reducing dependence on antimicrobials, while simultaneously improving vaccine stability, cost-effectiveness, and commercial scalability (Collins et al., 2019). As genomic technologies continue to converge from CRISPR-based antigen design to metagenomics-informed vaccine development, the aquaculture industry stands at the threshold of a new era in precision disease prevention.
Aquaculture research leverages genomic technologies and other omics to enhance feed utilization and understand aquatic organism's metabolic response to specific diets, feed ingredients, or additives (Hakim et al., 2018). One key success of genomics applications in aquaculture nutrition is developing and validating genetically engineered feed ingredients as adequate substitutes for fish meal and oil dependency. To date, there are numerous examples of genetically engineered organisms, such as plant-based organisms, microalgae, and other organisms that have been produced to enhance nutrient profiles in aquaculture feeds to achieve more nutritious and sustainable diets (Jones et al., 2020; Santos et al., 2020; Nagappan et al., 2021; Osmond et al., 2021). Moreover, transgenic fish demonstrate enhanced feed utilization efficiency and other advantages in production and product quality (Buwono et al., 2021). Feed efficiency is a crucial economic feature of most farmed species because animals with higher efficiency yield higher financial returns. Genetic improvements through breeding programs have shown the potential to improve feed efficiency in various aquaculture species, such as common carp (Zhang et al., 2021), rainbow trout (Kause et al., 2020), Nile tilapia (Barría et al., 2023), and sea bream (Besson et al., 2020).
Advances in epigenetics, metagenomics, and ncRNAs have provided valuable insights into the molecular changes induced by nutritional factors, enhancing our understanding of fish nutritional physiology and identifying nutritional status markers. To date, a dozen of nutrigenomic markers selection has been focused on several important aquaculture species such as spot cichlid (Amphilophus trimaculatus) (Hilerio-Ruiz et al., 2021), tropical gar (Atractosteus tropicus) (Martínez-Burguete et al., 2021), turbot (Scophthalmus maximus) (Hays et al., 2023) Atlantic salmon (Dvergedal et al., 2019; Fernández et al., 2023), rainbow trout (Abernathy and Overturf, 2019; Callet et al., 2018), gilthead sea bream (Sparus aurata) (Sáez-Arteaga et al., 2022; Naya-Català et al., 2023), shrimps (Dai et al., 2018) and important initiatives such as the ARRAINA Framework Programme (Calduch-Giner et al., 2016), AquaIMPACT (Naya-Català et al., 2022), have been developed to systematize the knowledge of expressed genes involved in species-specific metabolic networks and varying levels of metabolites in animal-fed specific nutrients. Generating nutritional markers across different developmental stages is essential for understanding the fundamental aspects of development, how different feeding regimes affect production, and identifying better feeding schemes for optimal growth in culture. As the metabarcoding approach has become more accessible, there has been a great revolution in the diet identification of aquatic species (Shum et al., 2023), which facilitates the characterization of dietary specializations, predatory feeding, feeding behavior, and dietary adaptation, thus helping to draw a more comprehensive picture of fish diets (Li et al., 2024). Li et al. (2024) compared morphological and gastrointestinal tract environmental DNA (GITeDNA) metabarcoding analyses of diets of fluviatile Masu salmon and found that GITeDNA metabarcoding offers higher efficiency in detecting diet diversity, revealing diet composition not previously recorded. Whole-genome DNA methylation patterns have been used to characterize key genes associated with feeding, such as those involved in feeding habit domestication in mandarin fish (Siniperca chuatsi) (Li et al., 2023), those involved in vegetarian adaptation in grass carp (Cai et al., 2018), and those involved in diet response in Nile tilapia (Podgorniak et al., 2022). All studies described above provide strong evidence for the impact of genomic technologies on aquaculture nutrition.
The primary problem to effective genomic applications in aquaculture is associated with the lack of high quality reference genomes, coupled with insufficiently annotated gene functions and poorly characterized genetic markers. Although recent initiatives, including Fish10K (Fan et al., 2020), FishmiRNA (Desvignes et al., 2022), Aqua-FAANG (Johnston et al., 2024), and NCBI Ref Seq updates (O'Leary et al., 2023), have made progress in improving genome annotations for aquatic species, yet many fish genomes remain incompletely assembled, with fragmented gene models and uneven taxonomic representation. Moreover, majority of sequenced fish genomes are derived from economically significant aquaculture species (e.g., salmonids, shrimp, tilapia, and catfish) or biomedical model organisms (e.g., zebrafish), leaving minor species underrepresented (Robinson et al., 2023). An additional challenge lies in the inadequate annotation of non-coding RNAs (ncRNAs) and regulatory elements, which play pivotal roles in development, environmental adaptation, and disease resistance.
Concerted effort to improve genome quality, annotation accuracy, and taxonomic inclusivity is essential to unlock the full potential of genomics in aquaculture, facilitating sustainable production and genetic resilience in the face of global environmental challenges. Strategic advancements should include: (1) expanding whole-genome assembly technologies, such as long-read sequencing (PacBio, Oxford Nanopore) and Hi-C scaffolding, to resolve complex genomic regions. (2) comparative genome re-sequencing leveraging well-annotated model species (e.g., zebrafish and medaka) to accelerate functional gene discovery, particularly for traits such as growth efficiency, disease resistance, and environmental resilience. (3) expansion of taxonomic representation in sequencing initiatives to prioritize underrepresented species, ensuring equitable coverage across diverse aquatic lineages.
The financial constraints associated with large-scale genotyping pose a substantial obstacle to the implementation of genomic selection (GS) in aquaculture breeding programs, particularly in small enterprises and developing economies. In contrast to terrestrial livestock breeding, aquaculture operations typically require genotyping of thousands of individuals per generation, especially for traits with low heritability, resulting in prohibitive costs that hinder widespread adoption (Tsairidou et al., 2020). This economic challenge is especially pronounced for lower-value species and resource-limited production systems, creating an innovation gap that threatens to exacerbate the disparities in aquaculture productivity. Nonetheless, several promising strategies have emerged for enhancing the accessibility of genomic breeding.
For example, genotype imputation has demonstrated particular promise, wherein high-density genotyping of parental stocks is combined with low-density genotyping of candidate selections followed by computational imputation to achieve high-density coverage (Yoshida et al., 2018; Tsairidou et al., 2020). This approach has been shown to maintain genomic prediction accuracy while reducing costs by 40–60% across multiple aquaculture species (Song and Hu, 2022). Collaborative efforts to develop multispecies genotyping arrays through industrial consortia represent another cost-reduction strategy, where economies of scale from high-volume species such as Atlantic salmon can subsidize array development for emerging species. This approach has been suggested to maintain prediction accuracy while reducing financial barriers in small- and lower-income economies. Genotyping pooled DNA from sib groups is another cost-reducing strategy that has been explored to minimize expenses while maintaining the effectiveness of genetic analysis. Furthermore, next-generation sequencing (NGS) techniques, such as genotyping-by-sequencing (GBS) and restriction site-associated DNA (RAD) sequencing, can create SNP panels for species lacking a reference genome, thereby lowering genotyping expenses. These methods have been successfully applied to various aquaculture species, including Atlantic salmon, large yellow croaker, rainbow trout, European sea bass, and Nile tilapia. By focusing on a limited set of restriction fragments, 3D-GBS allows fourfold more samples to be genotyped at the same minimal coverage depth. This approach allows for ultra-high-throughput and ultra-low-cost genotyping, reducing costs by approximately 86–92% for genomic selection and QTL mapping studies (De Ronne et al., 2023). The strategic integration of these cost-reduction approaches, tailored to specific breeding program requirements and economic contexts, is essential for realizing the full potential of genomic selection in the global aquaculture sector.
Bioinformatics proficiency and computational constraints remain significant obstacles hindering the extensive implementation of genomics in the aquaculture industry. Most aquaculture research institutions and production facilities lack the bioinformatics expertise needed to properly manage, analyze, and interpret sequencing data (Robinson et al., 2023). Advanced genomics, including next-generation sequencing (NGS) data analysis, complex population genetics simulations, and machine learning approaches for genotype-phenotype prediction, demands bioinformatics expertise rarely covered in standard genomics training programs. Even so, aquaculture education programs emphasize husbandry practices while underemphasizing bioinformatics training. Furthermore, the distinctive features of aquatic genomes, often characterized by higher heterozygosity and complex repeat structures, create substantial analytical hurdles that demand specialized bioinformatics approaches. Additionally, inadequate computational infrastructure further exacerbates the existing gap in bioinformatics skills in aquaculture genomics. Many institutions continue to rely on outdated computational systems that are ill equipped to handle contemporary genomic datasets (Rana et al.,, 2020). Although commercial cloud computing services (e.g., AWS, Google Cloud) offer potential solutions, their implementation remains cost-prohibitive for routine aquaculture applications (Krissaane et al., 2020), with many operations lacking funding for essential bioinformatics software licenses and HPC access (Rana et al., 2020).
To address the existing bioinformatics challenges, a multifaceted approach must be implemented, beginning with the integration of bioinformatics training into university curricula and vocational programs to develop workforce capacity, complemented by specialized workshops and online courses tailored to aquaculture applications. The development of customized analytical pipelines for non-model species will enhance variant calling and GWAS efficiency, while the creation of open-source, GUI-based platforms, such as the conceptualized AquaFAST, will improve accessibility for non-specialists. Infrastructure limitations should be addressed through the strategic adoption of cost-effective cloud computing solutions and the implementation of edge computing architectures for decentralized data processing in remote locations. Finally, establishing public-private partnerships to subsidize computational resources and forming international consortia for knowledge sharing will be critical for standardizing practices and ensuring equitable access to genomic resources across the aquaculture sector. Aquaculture may draw inspiration from successful collaboration in human genomics and crop science, as seen in The Cancer Genome Atlas (TCGA) and the International Wheat Genome Sequencing Consortium (IWGSC). This comprehensive strategy addresses both immediate technical barriers and long-term capacity-building needs, creating a sustainable framework for advancing genomic applications in aquaculture research and production systems.
Aquaculture genomics suffers from a scarcity of well-curated genomic resources, creating a significant knowledge gap that hinders genetic improvement efforts. To support the sustainable development of the aquaculture industry, there is a pressing need for a globally accessible database that integrates multi-omics data within a unified framework for both academic and industrial applications. Such a database should consolidate genomic resources with associated metadata, including evolutionary history, phenotypic traits, geographic distribution (GIS-based), and environmental parameters, all of which are accessible through an intuitive web-based interface. Currently, major repositories such as NCBI, Ensembl, and species-specific databases such as FishDB (Yang et al., 2020 c) and MiFish (Sato et al., 2018) serve as primary sources for fish genomic data. However, these platforms are not tailored to the specific demands of aquaculture and fishery research. A potential model for improvement is the AgBioData Consortium (Harper et al., 2018), which standardizes agricultural genomic data and can be adapted for aquaculture applications. Recognizing this gap, the Food and Agriculture Organization (FAO) developed AquaGRIS (https://www.fao.org/fishery/aquagris/home) (FAO, 2022), a prototype global information system for farmed aquatic genetic resources. This initiative aims to enhance data hosting, processing, annotation, and analysis, facilitating the better utilization of genomic resources in aquaculture breeding and conservation programs. Expanding such integrated databases will be crucial for advancing precision aquaculture and ensuring long-term industry sustainability.
Aquaculture genomic research faces significant challenges due to ethical, legislative, and regulatory disparities.. Ideally, new genetic technologies will encounter new and challenging regulations, and the public assessment and acceptance of this process and technology are critical. For example, public and regulatory acceptance of the application of genome-editing technology, particularly for seafood, has been rigorously debated (Wong et al., 2022). The European Union considers gene editing products as GMOs (Adamse et al., 2021), but in American countries, such as Brazil, similar products are not considered GMOs (Mutlu et al., 2024). The United States Food and Drug Administration (FDA) recently decided that GE salmon should be treated as a new “drug” (Van Eenennaam et al., 2021). These disparities can impede business and trade and hinder sustainable aquaculture development through inappropriate laws or policies. Governance structures (e.g., laws, policies, organizations, and agencies to enforce laws at national and international levels) should be involved in facilitating the use of genetic technologies in identifying, monitoring, and manipulating fisheries and aquaculture products. In addition, the international community should strive to reach a consensus on harmonized governance based on applying genetic technologies in fisheries and aquaculture management. The methodologies used should be standardized. Good governance based on genetic principles, internationally accepted principles, and inclusive and transparent communication provides consumer trust, public trust, and acceptance, eventually promoting decent fisheries and aquaculture production.
Aquaculture is a rapidly growing industry that plays a key role in meeting global food demands, with genomics playing a crucial role in this endeavor. Advancements in next-generation sequencing technologies and bioinformatics analysis pipelines have led to the publication of more aquaculture species genomes, presenting opportunities to solve many complex problems in several fields, including ecology, biochemistry, molecular phylogenetics, nutrition, physiology, and reproduction. The examples outlined in this paper illustrate the great value of genomics and genomic-derived information for the sustainable enhancement of aquaculture production. To keep moving forward, a few challenging issues must be tackled to make progress on these fronts even more rapidly. All stakeholders, including scientists and managers, must collaborate to integrate genomic-informed approaches into the management and production procedures. The way forward seems promising as more advanced genomics techniques become accessible. Over the next few years, development will focus on understanding the genetic architecture of complex traits and accelerating genetic progress in the selection for growth-, quality-, and robustness-related traits. In summary, lower-cost whole-genome sequencing approaches, the development and combined use of genomics and state-of-the-art omics technologies (metabolomics, proteomics, phenomics, and functional genomics), along with advanced computational techniques and artificial intelligence systems, are anticipated to augment aquaculture production in the coming decades significantly.