Floods are among the most common and destructive extreme hydrometeorological phenomena, and their impact on the environment extends beyond direct material damage to include long-term changes in the quality of terrestrial and aquatic ecosystems. One of the less visible but significant effects of flooding is the transfer of microorganisms of human and animal origin from urbanized areas, agricultural land, landfills, and cemeteries to river systems and, through precipitation and sedimentation, to riparian soils and floodplains. Such transfer can introduce various pathogens (bacteria, viruses, parasites) and mobile elements, such as antibiotic resistance genes (ARGs), into the soil, with direct health and sanitation implications for communities that use these areas (Mavrouli et al. 2022; Yu et al. 2022).
The literature indicates that microbial loads in surface waters increase after contamination. For example, studies following Hurricane Harvey in Texas and other hazard events have shown increases in fecal indicators (e.g., E. coli and enterococci), the appearance of antibiotic resistance genes (ARGs), and the presence of human fecal markers (Mukherjee et al. 2024). Moreover, analyses of microbial DNA in river studies have revealed a broad spectrum of enteropathogens – from bacteria (Salmonella spp., Campylobacter spp., Arcobacter spp., pathogenic strains of Escherichia coli) through parasites (Cryptosporidium oocysts, Giardia cysts) to enteric viruses (adenoviruses, noroviruses, rotaviruses) – which can then be deposited in sediments and flood soils (Potgieter et al. 2020; Barnes et al. 2023).
Specific studies of post-flood soil indicate that some microorganisms can survive and be detected in soil samples collected after water recedes. Examples include the detection of Burkholderia pseudomallei in post-flood soil samples in the Kelantan region (Malaysia) (Shaharudin et al. 2016) and the increased relative abundance of taxa associated with opportunistic or pathogenic genera (including members of Enterobacteriaceae and Yersiniaceae) in periodically flooded soils in river valleys (Furtak and Marzec-Grządziel 2025). Studies of urban soils inundated with floodwaters in Wrocław confirmed the presence of numerous potentially pathogenic taxa, including Klebsiella oxytoca and Leclercia adecarboxylata (Płoneczka-Janeczko et al. 2025). These observations indicate that floods not only distribute pathogens but also modify soil microbial structure, favoring the persistence or multiplication of specific potential pathogens (Das et al. 2025; Furtak and Marzec-Grządziel 2025). However, due to the unpredictable nature of floods, pre-flood data are often lacking, which hampers the conduct of such studies.
Although the deterioration of water quality after floods has been well documented, considerably less attention has been given to soils, despite their direct relevance to agriculture, human exposure, and food production. This review is novel in its focus on soils as a secondary, yet potentially long-term, reservoir of flood-borne pathogens. It is essential to understand the persistence of microbes and the routes of exposure in post-flood soils to develop evidence-based guidelines for farmers, land managers, and public health authorities.
Potential human pathogens naturally occur in aquatic environments, including Vibrio cholerae, Staphylococcus aureus, Mycobacterium tuberculosis, and Helicobacter pylori (Xie et al. 2022). These bacteria are commonly isolated from various aquatic environments, such as lakes, ponds, rivers, and their estuaries. Unfortunately, at the same time, river waters worldwide are losing their quality and purity (Stec et al. 2022; Furtak and Marzec-Grządziel 2025). River water is not usually a natural habitat for many pathogens, such as enterococci, but they are present in rivers due to human activity. The United Nations (UN) notes that one of the leading causes of this situation is the discharge of sewage into rivers; it estimates that over 80% of sewage is discharged directly into the environment (United Nations World Water Development 2017).
Examples of pathogens detected in river water (based on (Global Water Pathogen Project; Potgieter et al. 2020))
| Pathogen | Classification* |
|---|---|
| Aeromonas spp. | Waterborne |
| Escherichia coli | Water-based |
| Shigella spp. | Water-based |
| Legionella pneumophila | Water-based |
| Mycobacterium avium | Water-based |
| Pseudomonas aeruginosa | Water-based |
| Vibrio cholerae | Waterborne |
| Salmonella spp. | Water-based |
| Helicobacter pylori | Waterborne |
Waterborne – transmitted by water; water-based - spend part of life cycle in water
Pathogens in river water can be grouped into two categories based on their mode of transmission in the aquatic environment: (1) waterborne, transmitted through ingestion of contaminated water; and (2) water-based, which can survive or multiply in aquatic environments (Potgieter et al. 2020). Detected micro-organisms include opportunistic bacteria and pathogens with high epidemic potential, highlighting the importance of surface waters as a reservoir of microbiological hazards (Payment and Locas 2011). The high diversity of pathogens detected in rivers indicates their potential deposition in sediments and soils during flooding, creating secondary contamination hotspots (Sosah et al. 2025). Examples of pathogens classified as water-based include Escherichia coli, Shigella spp., Salmonella spp., Legionella pneumophila, Mycobacterium avium, and Pseudomonas aeruginosa (Global Water Pathogen Project; Potgieter et al. 2020). These micro-organisms can survive and, in some cases, multiply in the aquatic environment, particularly in biofilms and sediments (Cabral 2010). Their presence in river waters is often associated with municipal sewage inflows, surface runoff from agricultural areas, and anthropogenic activities (Sosah et al. 2025). These pathogens pose a particular risk to immunocompromised individuals and in contexts involving the use of surface waters for recreational purposes or as drinking water sources. Pathogens classified as waterborne, such as Aeromonas spp., Vibrio cholerae, or Helicobacter pylori, are primarily transmitted via water, which acts as a transmission vector; however, it does not always favor their long-term multiplication (Global Water Pathogen Project; Potgieter et al. 2020). The presence of these microorganisms in river water is of significant epidemiological importance, as they can cause gastrointestinal disease outbreaks and transmit these diseases to distant areas during flooding. A particular threat is posed by epidemics of cholera or gastroenteritis caused by other vibrios or Aeromonas spp. (Cabral 2010). Identifying both water-based and waterborne pathogens is crucial for assessing health risks and developing effective strategies to manage water resources and protect public health (Su et al. 2024).
The problem of diseases caused by waterborne microorganisms poses a serious challenge to public health. Waterborne pathogens and opportunistic pathogens are the most important causes of morbidity and mortality worldwide. According to the World Health Organization (WHO), over 50% of deaths from water-related diseases are caused by bacterial intestinal infections (World Health Organization 2016). General WHO data indicate that every year, 3.4 million people die from water-related diseases. Illness can occur not only from consuming water contaminated with pathogens but also from using contaminated recreational bathing areas (Stec et al., 2022; Custodio et al., 2025).
As mentioned above, the UN report states that sewage discharge is one of the primary sources of pathogens in the river (United Nations World Water Development 2017). Research indicates that this discharge promotes the proliferation of pathogenic bacteria, including species from the Enterobacteriaceae family, Coxiella burnetii, Vibrio spp., Helicobacter spp., Mycobacterium tuberculosis, and Staphylococcus spp. (Xie et al. 2022). It is important to emphasize that river pollution is caused not only by the inflow of untreated sewage but also by the introduction of treated sewage, which alters the parameters of the aquatic environment and promotes the proliferation of pathogens (Xie et al. 2022) (Figure 1). Both untreated and treated hospital wastewater are hazardous to watercourses because of their potential to transfer pathogens (Męcik et al. 2024). Hospital sanitary waste (feces, urine, biological waste) may contain pathogens and antibiotic resistance genes (Yuan and Pian 2023). Research indicates that antibiotic resistance genes continue to transfer from the river environment to drinking water and the human gut (Yu et al. 2022).
A second important source of pathogens in water reservoirs is agriculture, particularly livestock farming (Stec et al. 2022) (Figure 1). Animal husbandry (cattle, pigs, poultry, and other livestock) generates animal feces that may contain zoonotic bacteria (capable of infecting humans), as well as protozoa and parasites. During rainfall or flooding, feces can be washed into water systems (including rivers) (Díaz-Gavidia et al. 2022). Such microorganisms include, among others: Brucella spp., Chlamydia spp., Leptospira spp., Rickettsia spp., Salmonella spp., Brachyspira hyodysenteriae, Bacillus anthracis, Erysipelothrix rhusiopathiae, Mycobacterium spp., and Campylobacter spp.
A less obvious source of pathogens and antibiotic resistance genes is burial sites. A review of the literature indicates that biological material from bodies can enter groundwater or, through surface runoff, enter watercourses (Tarnawska et al. 2024).
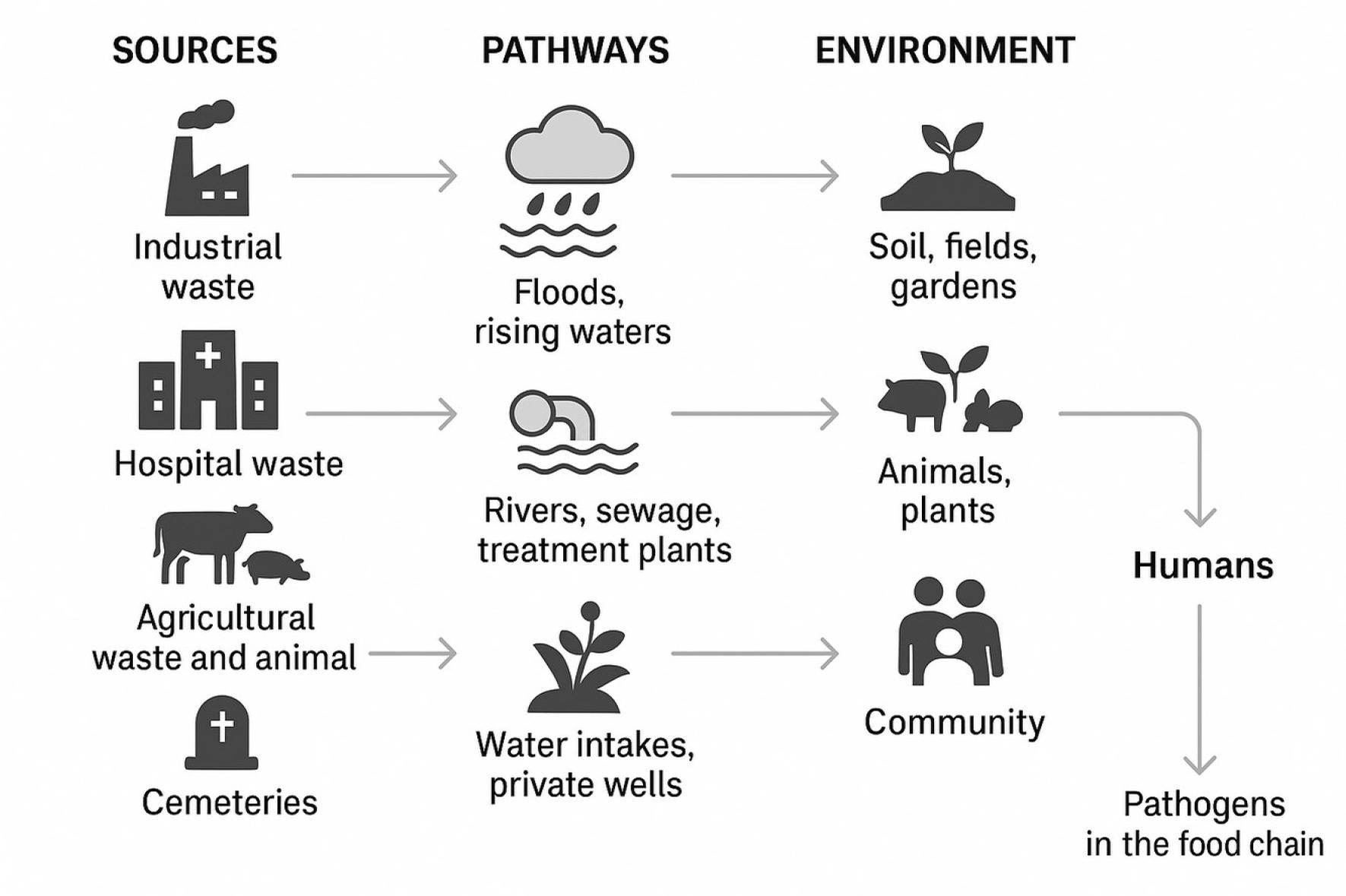
Sources of microbial pathogens during flood (author’s elaboration using bioRender software).
River water, as shown above, can itself be contaminated with pathogenic bacteria from the sources described. It is estimated that about one-third of all rivers in Latin America, Africa, and Asia are contaminated with pathogens. At the same time, severe organic contamination occurs in about one-seventh of all rivers worldwide (Annual Report 2022 | UNEP - UN Environment Programme). Floodwater can therefore carry pathogens, including opportunistic pathogens already present in it, into flooded areas (fields, drinking water intakes, plants) (Figure 1). Rising water levels promote the mobilization of microorganisms from sediments, soils, and sanitary infrastructure.
River water that overflows its banks during flooding may come into contact with numerous potential sources of pathogens (Figure 1): sewage treatment plants, landfills, cemeteries, farm buildings (cowsheds, piggeries, poultry houses), and manure piles, as well as plant and animal waste (Furtak and Marzec-Grządziel 2025). Like pathogens flowing from agricultural areas, cemeteries, and landfills into watercourses during rainfall, floods transfer microorganisms on a larger scale because of the size of the flooded area and the extent and duration of flooding at these sources.
Rainfall, floods, or rising floodwaters can transport pollutants from land areas (agricultural, cemetery, residential) to rivers and waterways (Cheung et al. 2025). Stagnant floodwater can pick up microorganisms from soil, plants, dead plants, feces, or animal waste (Figure 1). Floodwaters can transport pathogens over long distances, linking sources to agricultural areas, home gardens, and water intake points. This means that many sources that are usually distant can end up in river waters during a flood (Potgieter et al. 2020). Although this load is diluted, pathogens are carried further and can be detected in the river even after the flood has subsided. This is indicated by reports from studies conducted after hurricanes/floods, where noroviruses, rotaviruses, Cryptosporidium sp., and Giardia sp., as well as enteropathogenic bacteria such as Salmonella sp. and Campylobacter sp., were often detected in flood surface waters (Acosta-España et al. 2024).
The transfer of pathogens via floodwater is associated with the accumulation of microorganisms in water, soil, and plants (Dzodzomenyo et al. 2022). These microorganisms can persist in soil for extended periods and be transferred to subsequent crops during the next growing season. This introduces pathogens into the human and animal food chain (Nichols et al. 2018). Animals can become infected by grazing on contaminated land, drinking untested water, and consuming feed from flooded areas. Not only can farm animals become ill themselves, but they can also cause secondary transmission of pathogens to humans (zoonoses).
The final link in the chain is humans (Figure 1), who can become infected through direct contact with contaminated environments, by consuming contaminated water or food, or by contact with infected animals (Potgieter et al. 2020; World Health Organization 2006). Pathogen accumulation in the food chain increases the risk of waterborne and foodborne disease outbreaks, particularly after flooding, when standard sanitary barriers are disrupted (Boelee et al. 2019). These pathways are particularly pertinent to agricultural floodplains, justifying targeted soil monitoring after flooding (Howard et al. 2016). Farmers who clean their farms and clear fields and who are in direct contact with carrier animals are among those most at risk of post-flood diseases. Plants from flooded areas can also introduce pathogens into the human food chain (Food and Agriculture Organization of the United Nations 2023).
The presence of pathogens in floodwater and post-flood water has been confirmed, as has their partial transfer to soil. However, their transfer to plants or animals has been poorly studied. There is a risk of microorganisms spreading in environments flooded with such water, particularly when weather conditions are favorable or when flooding lasts for a long time. This knowledge gap is significant because pathogens can be transmitted to humans during fieldwork and cleanup, to animals fed feed from flooded areas, and to crops destined for food processing plants.
For safety reasons, areas affected by flooding, especially drinking water intakes and, sometimes, soil, are tested for contaminants such as pesticides, fuels, and heavy metals. Global research in flood-affected areas primarily focuses on controlling microbiological contamination of soil and water by Salmonella spp., Clostridium spp., and Escherichia spp. (Divakaran et al., 2019; Tyszkiewicz and Banaszuk, 2016).
Numerous studies in the literature confirm that groundwater and drinking water intakes become contaminated after flooding subsides. A comprehensive literature review by Andrade et al. (2018) found that floods cause groundwater contamination and numerous cases of gastrointestinal illness in humans. The microorganisms identified in these cases include coliform bacteria, fecal streptococci, Vibrio cholerae, Legionella spp. (including L. pneumophila), and Clostridium spp. (including C. tetani) (Dai et al. 2019; Moghadam et al. 2022).
While the proliferation of pathogenic microorganisms in floodwater is rapid and short-lived, traces of pathogens can persist in the sediments left behind after the floodwaters recede for much longer (Mukherjee et al. 2024). At the same time, there is still little research on the persistence of pathogens in soil after flooding. This is surprising, given ongoing climate change. Studies confirm that floods increase the risk of microbial contamination of fields and crops, but direct contamination of such post-flood areas is rarely studied.
Some studies indicate the long-term persistence of pathogens in soil. Table 2 provides examples of pathogens detected in post-flood soils across different regions. Notably, several studies report prolonged survival of bacteria such as E. coli and Salmonella spp., with detection up to 44 days after flooding (Bergholz et al. 2016) (Table 2). Multiple factors, including soil texture, organic matter content, temperature, moisture, oxygen availability, and specific microbial traits such as spore formation or biofilm production, influence persistence. Regional differences are evident: warmer climates and fine-textured, organic-rich soils generally favor prolonged pathogen survival. The transport of bacteria through soil in floodwater depends on soil type, moisture content, and terrain slope (Ning et al. 2023). Microorganisms move with varying efficiency through soils with different structures; in some soils, they can travel farther horizontally, whereas in others they disappear more quickly the farther they are from the flood source (Callahan et al. 2017). For instance, soils with a higher concentration of fine particles (e.g., clay) have been shown to facilitate prolonged accumulation of specific pathogens, such as Salmonella enterica, compared to sand (Phan-Thien et al. 2020). However, many findings are based on limited case studies or laboratory experiments, underscoring the need to distinguish well-supported conclusions from preliminary observations.
Examples of pathogens detected in post-flood soils
| Microorganisms | Range of MICs (µg/mL) | Methodological remarks (medium type, colony counts, incubation time, and temperature) | References |
|---|---|---|---|
| Gram-positive bacteria | |||
| Clostridium perfringens | 1 | MHB, 105 cfu/mL, 24–48 h, 36°C | (Koburger et al. 2010) |
| Enterococcus faecalis | 4 | MHB, 105 cfu/mL, 24–48 h, 36°C | (Koburger et al. 2010) |
| 3.125–6.25 | TSB, 105 cfu/mL, 24–48 h, 37°C | (Karpiński, et al. 2025a) | |
| E. faecium | 0.49–1.95 | TSB, 105 cfu/mL, 24 h, 37°C | (Dydak et al. 2021) |
| 3.125–6.25 | TSB, 105 cfu/mL, 24–48 h, 37°C | (Karpiński, et al. 2025a) | |
| E. hirae | 0.6–10 | TSB, 108–109 cfu/mL, 24–72 h, no data | (Schug et al. 2022) |
| Staphylococcus aureus | 2 | MHB, 105 cfu/mL, 24–48 h, 36°C | (Koburger et al. 2010) |
| 0.49–0.98 | TSB, 105 cfu/mL, 24 h, 37°C | (Dydak et al. 2021) | |
| 2–4 | SCS, 1.5–5×105 cfu/mL, 48 h, 37°C | (Denkel et al. 2022) | |
| 0.9 | MHB, 105 cfu/mL, 24 h, 37°C | (Krasowski et al. 2021) | |
| 0.3–5 | TSB, 108–109 cfu/mL, 24–72 h, no data | (Schug et al. 2022) | |
| 3.125–6.25 | TSB, 105 cfu/mL, 24–48 h, 37°C | (Karpiński, et al. 2025a) | |
| Methicillin-resistant S. aureus (MRSA) | 1 | MHB, 105 cfu/mL, 24–48 h, 36°C | (Koburger et al. 2010) |
| 1–4 | MHB, 5×105 cfu/mL, 24–48 h, 37°C | (Dittmann et al. 2019) | |
| S. epidermidis | 0.49–7.8 | TSB, 105 cfu/mL, 24 h, 37°C | (Dydak et al. 2021) |
| Coagulase-negative staphylococci | 2–4 | SCS, 1.5–5×105 cfu/mL, 48 h, 37°C | (Denkel et al. 2022) |
| Streptococcus pneumoniae | 8–32 | MHB, 105 cfu/mL, 24–48 h, 36°C | (Koburger et al. 2010) |
| S. pyogenes | 3.125–6.25 | TSB, 105 cfu/mL, 24–48 h, 37°C | (Karpiński, et al. 2025a) |
| Gram-negative bacteria | |||
| Acinetobacter baumannii | 0.25–3.9 | TSB, 105 cfu/mL, 24 h, 37°C | (Dydak et al. 2021) |
| Enterobacter cloacae | 3.9 | TSB, 105 cfu/mL, 24 h, 37°C | (Dydak et al. 2021) |
| 6.25 | TSB, 105 cfu/mL, 24–48 h, 37°C | (Karpiński, et al. 2025a) | |
| Escherichia coli | 2 | MHB, 105 cfu/mL, 24–48 h, 36°C | (Koburger et al. 2010) |
| 1.95–3.9 | TSB, 105 cfu/mL, 24 h, 37°C | (Dydak et al. 2021) | |
| 2–4 | SCS, 1.5–5×105 cfu/mL, 48 h, 37°C | (Denkel et al. 2022) | |
| 1.95–3.9 | MHB or artificial urine, 105–106 cfu/mL, 20 ± 2 h, 37°C | (Loose et al. 2021) | |
| 1–4 | MHB, 106 cfu/mL, 20 ± 2 h, 37°C | (da Silva et al. 2023) | |
| 0.6–20 | TSB, 108–109 cfu/mL, 24–72 h, no data | (Schug et al. 2022) | |
| 3.125–6.25 | TSB, 105 cfu/mL, 24–48 h, 37°C | (Karpiński, et al. 2025a) | |
| Haemophilus influenzae | 1 | MHB 105 cfu/mL, 24–48 h, 36°C | (Koburger et al. 2010) |
| Klebsiella spp. | 2–4 | SCS, 1.5–5×105 cfu/mL, 48 h, 37°C | (Denkel et al. 2022) |
| K. pneumoniae | 1.95–7.8 | TSB, 105 cfu/mL, 24 h, 37°C | (Dydak et al. 2021) |
| 3.125–6.25 | TSB, 105 cfu/mL, 24–48 h, 37°C | (Karpiński, et al. 2025a) | |
| Proteus mirabilis | 1.95–3.9 | MHB or artificial urine, 105–106 cfu/mL, 20 ± 2 h, 37°C | (Loose et al. 2021) |
| 3.125–6.25 | TSB, 105 cfu/mL, 24–48 h, 37°C | (Karpiński, et al. 2025a) | |
| Pseudomonas aeruginosa | 2–8 | MHB, 105 cfu/mL, 24–48 h, 36°C | (Koburger et al. 2010) |
| 3.9–15.7 | TSB, 105 cfu/mL, 24 h, 37°C | (Dydak et al. 2021) | |
| 8–32 | SCS, 1.5–5×105 cfu/mL, 48 h, 37°C | (Denkel et al. 2022) | |
| 2.25±0.95 | MHB, 105 cfu/mL, 24 h, 37°C | (Krasowski et al. 2021) | |
| 3.9–7.8 | MHB or artificial urine, 105–106 cfu/mL, 20 ± 2 h, 37°C | (Loose et al. 2021) | |
| 3.91–15.63 | TSB, 105 cfu/mL, 24 h, 36°C | (Karpiński, et al. 2025b) | |
| 1.25–80 | TSB, 108–109 cfu/mL, 24–72 h, no data | (Schug et al. 2022) | |
| 3.125–12.5 | TSB, 105 cfu/mL, 24–48 h, 37°C | (Karpiński, et al. 2025a) | |
| Salmonella enterica | 6.25 | TSB, 105 cfu/mL, 24–48 h, 37°C | (Karpiński, et al. 2025a) |
| Shigella flexneri | 6.25–12.5 | TSB, 105 cfu/mL, 24–48 h, 37°C | (Karpiński, et al. 2025a) |
| Yersinia enterocolitica | 6.25 | TSB, 105 cfu/mL, 24–48 h, 37°C | (Karpiński, et al. 2025a) |
| Fungi | |||
| Ascophera apis | 0.78–3.125 | TSB, 105 cfu/mL, 24–48 h, 37°C | (Karpiński, et al. 2025a) |
| Candida albicans | 1 | MHB, 105 cfu/mL, 24–48 h, 36°C | (Koburger et al. 2010) |
| 0.49–0.98 | TSB, 105 cfu/mL, 24 h, 37°C | (Dydak et al. 2021) | |
| 0.45 | RPMI with 2% glucose, 105 cfu/mL, 24 h, 37°C | (Krasowski et al. 2021) | |
| 0.5 ± 0.25 and 0.9 ± 0.4 | TSB, 106 cfu/mL, 24 h, 36°C | (Korbecka-Paczkowska and Karpiński 2024) | |
| 1.95–3.91 | Sabouraud broth, 106 cfu/mL, 24 h, 36°C | (Karpiński et al. 2024) | |
| 0.78–1.56 | TSB, 105 cfu/mL, 24–48 h, 37°C | (Karpiński, et al. 2025a) | |
| C. auris | 3.125 | TSB, 105 cfu/mL, 24–48 h, 37°C | (Karpiński, et al. 2025a) |
| C. glabrata | 0.78–3.125 | TSB, 105 cfu/mL, 24–48 h, 37°C | (Karpiński, et al. 2025a) |
| C. tropicalis | 0.78–1.56 | TSB, 105 cfu/mL, 24–48 h, 37°C | (Karpiński, et al. 2025a) |
| Cryptococcus neoformans | 3.125 | TSB, 105 cfu/mL, 24–48 h, 37°C | (Karpiński, et al. 2025a) |
| Rhodotorula mucilaginosa | 3.125 | TSB, 105 cfu/mL, 24–48 h, 37°C | (Karpiński, et al. 2025a) |
Currently, there are no clear guidelines for handling soil and agricultural produce after a flood. First, the soil should be dried and its structure loosened to aerate it and prevent anaerobic conditions (Food and Drug Administration 2011; Taylor et al. 2013). Next, tests should be conducted to determine the soil’s basic physical and chemical parameters and its microbiological contamination (Food and Drug Administration 2011). Silt and flood sediment on soils at risk of sanitary contamination may contain potential pathogens (Furtak 2025). Protective masks and gloves should be worn when removing them from fields. One basic sanitation method is to lime the soil, as it has disinfectant properties. Oxide lime acts as both a disinfectant and a heavy metal neutralizer (Li et al. 2022; Wenyika et al. 2025). Disinfectants are available on the market that can be used independently in farm buildings (Makovska et al. 2025). In areas with significant contamination, the following methods can be used: fumigation (Porter et al. 1999), physical methods (Gullino et al. 2022), and bioremediation (Hossain et al. 2025). Before disinfection, it is essential to clear the soil surface of animal and plant debris, waste, and flood sediment.
In the face of advancing climate change, increasingly frequent flooding, and deteriorating river water quality, research on the microbiological safety of flood-affected areas is essential. The lack of such data, especially for farmland, leaves farmers without explicit, simultaneous guidance on managing crops, yields, or soil. There are no universal guidelines for food safety after flooding, nor are there indications of whether to remove agricultural products affected by flooding. Research in this area is essential and should be a focus of environmental microbiologists. Key challenges include the lack of pre-flood baseline data, limited temporal monitoring, and methodological constraints.
There are apparent gaps in the literature on the impact of floods on soil environments, stemming from a lack of baseline soil property data before flooding and insufficient monitoring afterward (Furtak and Marzec-Grządziel 2025). Without measurements taken beforehand, it is impossible to accurately estimate the extent of changes caused by extreme hydrological phenomena, let alone assess cause-and-effect relationships (Das et al. 2024; Furtak et al. 2020). Sporadic, uneven monitoring after the flood recedes does not provide a complete picture of soil regeneration processes. Studies from post-flood periods often focus on specific aspects (e.g., pollution, chemical parameters) and lack a systematic, long-term approach. Consequently, knowledge of the trajectory of soil recovery to its original state, or of permanent changes in soil functions and pathogen accumulation, remains fragmentary (Das et al. 2024; Sao et al. 2025). Despite growing awareness of sanitary risks, there is a clear gap in the scientific literature due to a lack of baseline studies and monitoring of soil pathogens before and after flooding (Mapili et al. 2022).
Methods to date have mainly relied on culturing bacteria on selective growth media. This approach excludes entire groups of non-cultivable bacteria, as well as those that require specific cultivation conditions (Al-Awadhi et al. 2013; Feleni et al. 2025). Results based solely on culture may systematically underestimate actual microbiological exposure. Therefore, assessing pathogen risk in the aquatic environment requires modern detection tools and microbiological indicators that can identify the presence of and the potential pathogenicity of microorganisms in water used by humans. While culture-based methods may underestimate pathogen diversity, molecular techniques detect DNA from both viable and non-viable cells (Sheikh et al. 2012). Furthermore, the scope of sanitary testing should be expanded due to increasing antibiotic resistance in bacteria and the emergence of new, previously unanalyzed pathogens (McLellan et al. 2024).
Flood-affected areas should be monitored for microbiomes, with particular attention to pathogenic microorganisms, rather than focusing solely on basic indicator bacteria, such as fecal bacteria. The importance of systematic research on microbiological soil contamination after flooding is multidimensional. First, accurate data on which organisms and at what concentrations enter the soil are essential for assessing the health risk to the population (contact with soil, agricultural use, recreational contact). Second, monitoring correlated with hydrological analyses enables distinguishing between sources of contamination (e.g., municipal sewage, hospital sewage, animal waste, surface runoff from fields, releases from cemeteries), which is crucial for designing preventive measures and risk management. Third, identifying the presence of ARGs and opportunistic pathogens in soils after flooding has implications for controlling the spread of antibiotic resistance in the environment. Finally, the results of such studies provide a basis for developing sanitary and epidemiological guidelines for the safe use of flood-affected areas and for rapid response strategies after flooding (e.g., decontamination, use bans, water and soil testing before restoration of use) (Mavrouli et al. 2022).
Available data clearly show that flooding leads to the long-term persistence of pathogenic microorganisms in soil. This is particularly significant in agricultural soils, as these pathogens can be transferred to farmers, livestock, and plants — and therefore to crops and feed. Post-flood soil analysis should be standard practice, encompassing the full spectrum of potential soil, water, and human-origin pathogens, as floodwaters transmit microorganisms from multiple sources. Guidelines on food safety after flooding, along with instructions for handling affected agricultural products, are needed. To develop such policies, it is necessary to understand the problem and its scope. Strengthening soil-focused microbiological research, clarifying exposure routes, and developing practical mitigation guidelines are essential steps toward effective flood risk management in a changing climate.