Fig. 1.
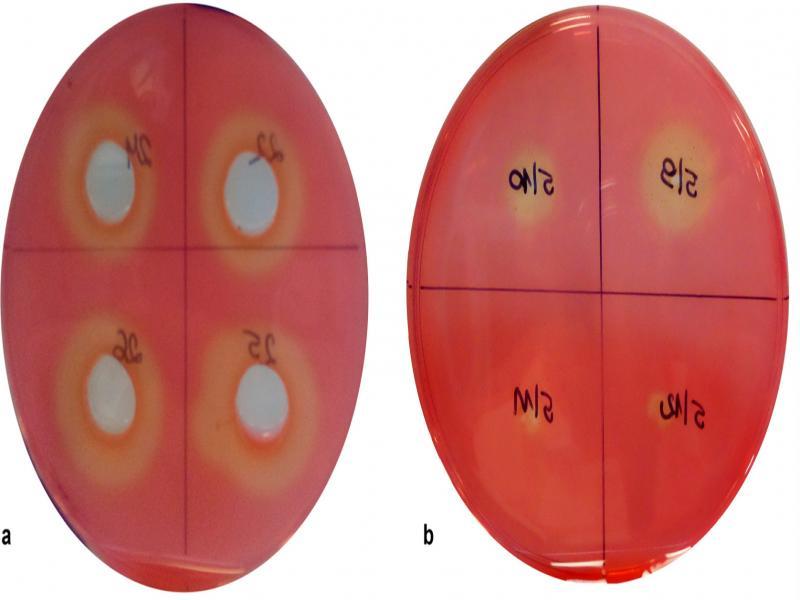
Fig. 2.
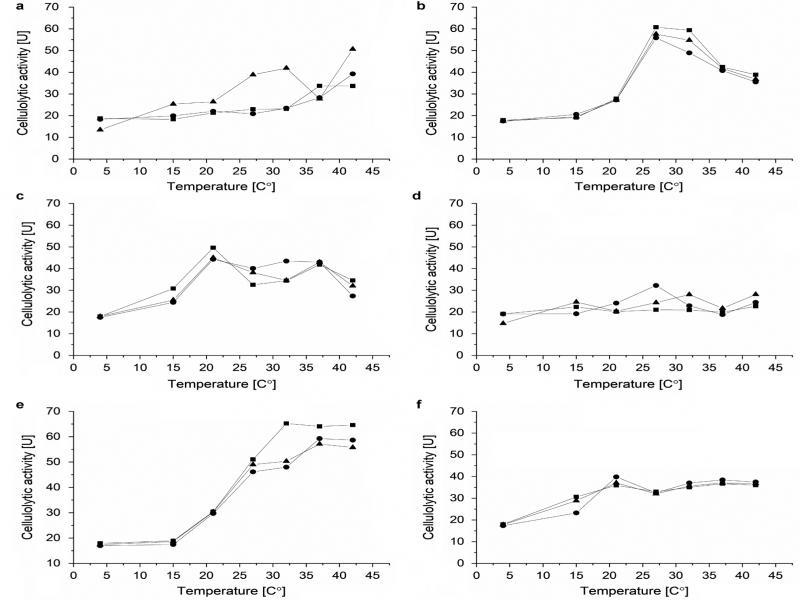
Fig. 3.
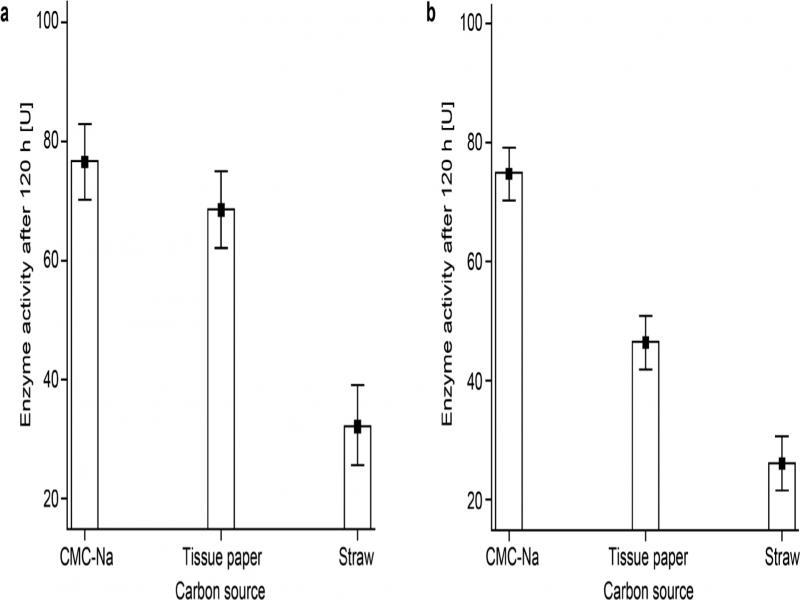
Fig. 4.
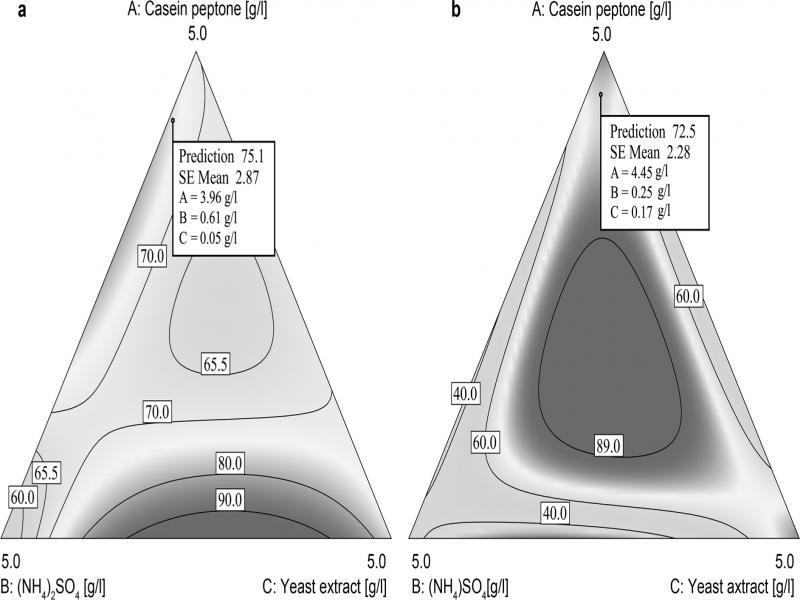
Regression coefficients and the selected fit statistics for two strains investigated_
| Component in model | 4/7 – Bacillus subtilis | 4/18 – Bacillus licheniformis | ||||
|---|---|---|---|---|---|---|
| CMC-Na | Tissue paper | Straw | CMC-Na | Tissue paper | Straw | |
| A | 14.010 | 13.758 | 15.113 | 16.453 | 9.146 | 11.111 |
| B | 9.885 | 8.696 | 11.233 | 9.128 | 9.797 | 3.789 |
| C | 15.096 | 15.718 | 13.633 | 16.565 | 8.661 | 6.956 |
| D | 8.724 | 7.120 | 7.468 | 9.490 | 3.248 | 4.099 |
| A×B | 3.277 | 3.000 | 0.597 | −3.483 | 0.494 | −2.712 |
| A×C | −0.808 | −5.549 | −3.786 | −6.638 | 3.085 | −3.472 |
| A×D | 1.294 | −0.602 | −5.714 | −2.414 | 1.225 | −2.800 |
| B×C | 4.399 | −2.491 | −1.287 | −6.412 | −1.193 | −0.983 |
| B×D | 1.619 | 4.134 | 3.687 | −4.134 | −0.166 | 3.289 |
| C×D | 1.161 | 2.324 | −1.994 | −4.717 | 1.665 | −0.530 |
| A×B×C | −5.876 | −4.340 | −31.119 | 18.744 | −8.735 | −6.175 |
| B×C×D | 5.785 | 9.842 | 24.012 | 8.648 | 8.648 | 8.648 |
| Adj-R2 | 0.886 | 0.941 | ||||
| Model p-value | < 0.0001 | < 0.0001 | ||||
Composition of the nitrogen sources blends according to the mixture design_
| Run | A: Casein peptone (g/l) | B: (NH4)2SO4(g/l) | C: Yeast extract (g/l) | D: NH4Cl (g/l) |
|---|---|---|---|---|
| 1 | 5.00 | 0.000 | 0.000 | 0.000 |
| 2 | 0.00 | 5.000 | 0.000 | 0.000 |
| 3 | 0.00 | 0.000 | 5.000 | 0.000 |
| 4 | 0.00 | 0.000 | 0.000 | 5.000 |
| 5 | 1.25 | 1.250 | 1.250 | 1.250 |
| 6 | 3.13 | 0.625 | 0.625 | 0.625 |
| 7 | 0.63 | 3.125 | 0.625 | 0.625 |
| 8 | 0.63 | 0.625 | 3.125 | 0.625 |
| 9 | 0.63 | 0.625 | 0.625 | 3.125 |
| 10 | 2.50 | 0.000 | 0.000 | 2.500 |
| 11 | 2.50 | 0.000 | 2.500 | 0.000 |
| 12 | 2.50 | 2.500 | 0.000 | 0.000 |
| 13 | 0.00 | 2.500 | 0.000 | 2.500 |
| 14 | 0.00 | 2.500 | 2.500 | 0.000 |
| 15 | 0.00 | 0.000 | 2.500 | 2.500 |
| 16 | 2.50 | 0.000 | 2.500 | 0.000 |
| 17 | 5.00 | 0.000 | 0.000 | 0.000 |
| 18 | 0.00 | 5.000 | 0.000 | 0.000 |
| 19 | 0.00 | 0.000 | 5.000 | 0.000 |
| 20 | 0.00 | 0.000 | 0.000 | 5.000 |
The response data for the cellulose-degrading enzymes activity [U] of each experimental point after the cultivation of the selected strains on the media containing various substrates for 120 h_
| Run | 4/7 – Bacillus subtilis | 4/18 – Bacillus licheniformis | ||||
|---|---|---|---|---|---|---|
| CMC-Na | Tissue paper | Straw | CMC-Na | Tissue paper | Straw | |
| 1 | 73.341 | 69.931 | 76.155 | 84.545 | 45.651 | 54.975 |
| 2 | 51.870 | 46.036 | 57.331 | 47.838 | 48.956 | 18.901 |
| 3 | 78.858 | 81.562 | 70.540 | 86.859 | 43.448 | 33.492 |
| 4 | 44.842 | 37.608 | 38.008 | 49.592 | 16.792 | 20.351 |
| 5 | 86.250 | 69.578 | 90.502 | 67.032 | 47.400 | 25.125 |
| 6 | 67.130 | 59.652 | 22.006 | 70.089 | 42.723 | 51.931 |
| 7 | 63.610 | 61.600 | 55.361 | 59.664 | 45.89 | 21.751 |
| 8 | 75.801 | 61.600 | 31.052 | 72.537 | 46.864 | 31.312 |
| 9 | 65.790 | 70.978 | 64.362 | 52.856 | 35.44 | 30.849 |
| 10 | 64.633 | 45.841 | 18.583 | 47.315 | 40.568 | 21.191 |
| 11 | 67.991 | 46.718 | 48.692 | 40.523 | 69.297 | 25.819 |
| 12 | 81.805 | 74.096 | 67.183 | 41.298 | 50.323 | 21.398 |
| 13 | 58.227 | 64.584 | 67.386 | 17.805 | 33.017 | 41.822 |
| 14 | 91.841 | 46.815 | 55.102 | 22.799 | 38.85 | 20.497 |
| 15 | 66.825 | 71.173 | 41.523 | 32.724 | 42.395 | 23.676 |
| 16 | 67.451 | 30.849 | 48.943 | 40.674 | 58.946 | 20.022 |
| 17 | 66.886 | 66.850 | 74.287 | 80.405 | 45.374 | 56.023 |
| 18 | 48.983 | 41.919 | 54.057 | 43.417 | 48.097 | 19.754 |
| 19 | 72.537 | 76.958 | 68.496 | 78.736 | 43.021 | 34.637 |
| 20 | 42.529 | 32.785 | 35.983 | 43.71 | 16.822 | 20.972 |
The results of identification of cellulose-degrading bacteria, based on the 16 rRNA gene sequencing_
| Number of strain | Identity (%) | Strain of the closest match |
|---|---|---|
| 4/3 | 86.09 | Bacillus pseudomycoides |
| 85.84 | Bacillus cereus | |
| 4/7 | 99.86 | Bacillus subtilis |
| 4/11 | 99.93 | Bacillus cereus |
| 99.86 | Bacillus thuringiensis | |
| 4/18 | 99.86 | Bacillus licheniformis |
| 6/5 | 100.00 | Achromobacter piechaudi |
| 100.00 | Alcaligenes faecalis | |
| 99.93 | Achromobacter xylosoxidans | |
| 6/6 | 99.97 | Lactococcus lactis |