Figure 1.
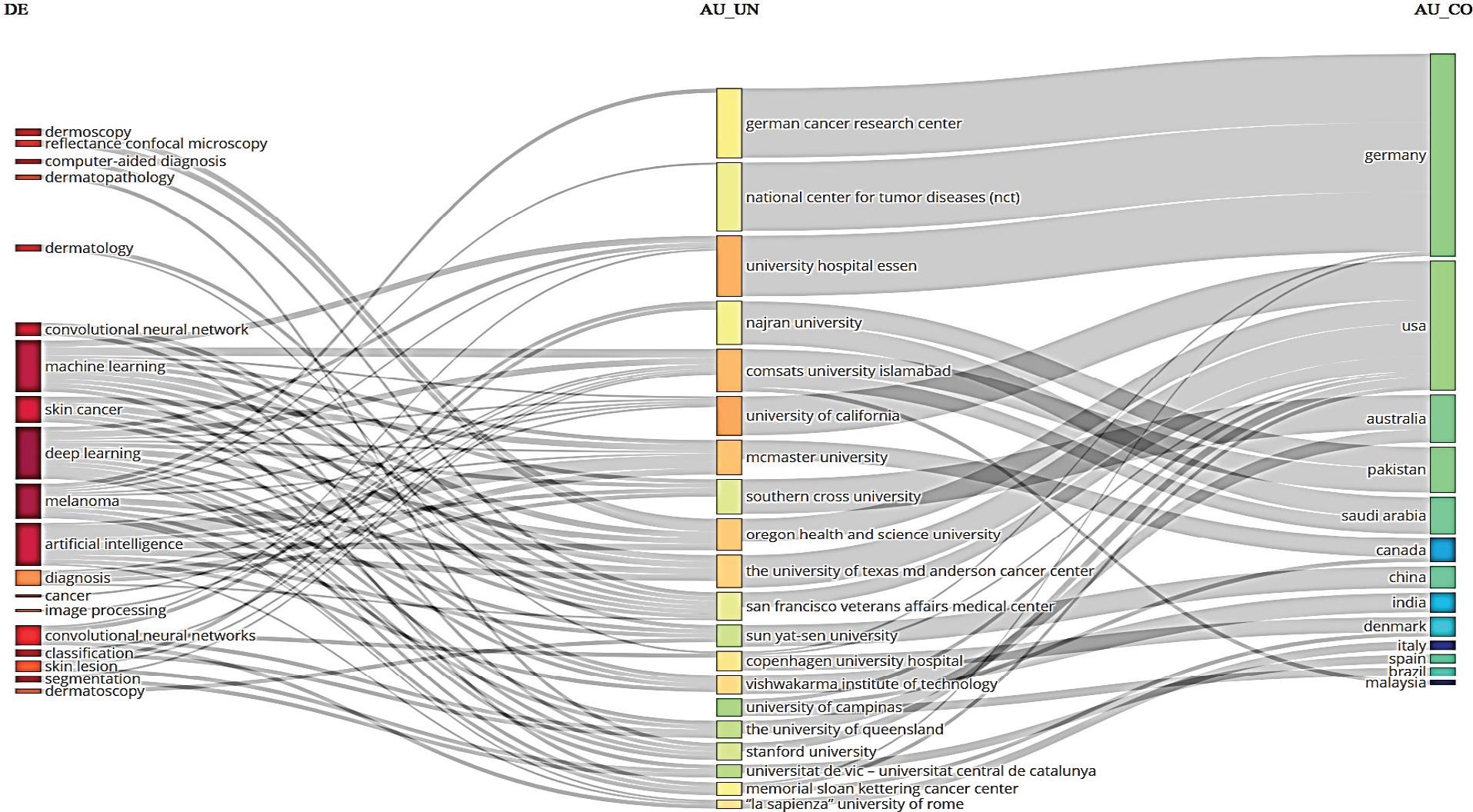
Figure 2.
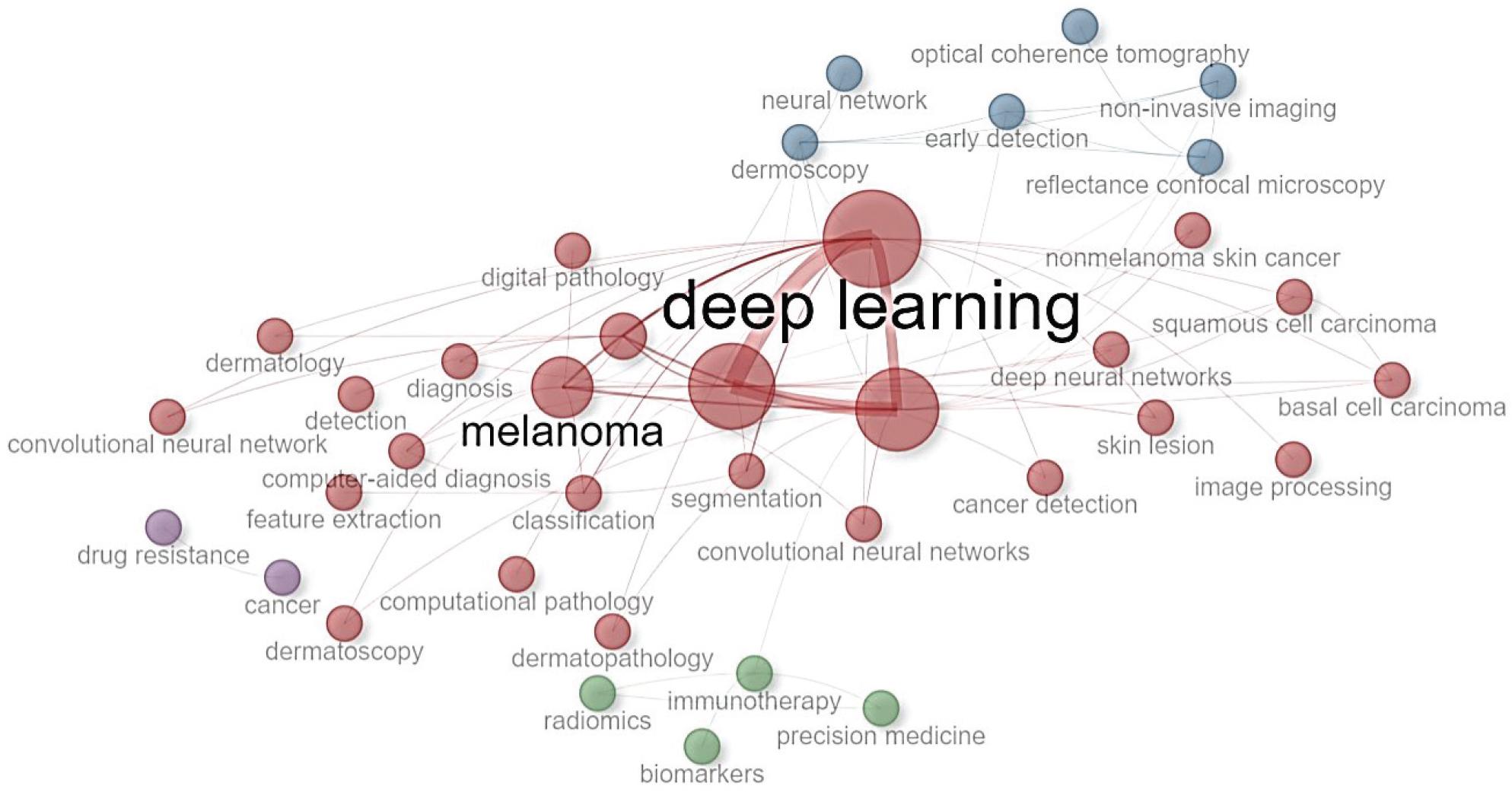
Figure 3.
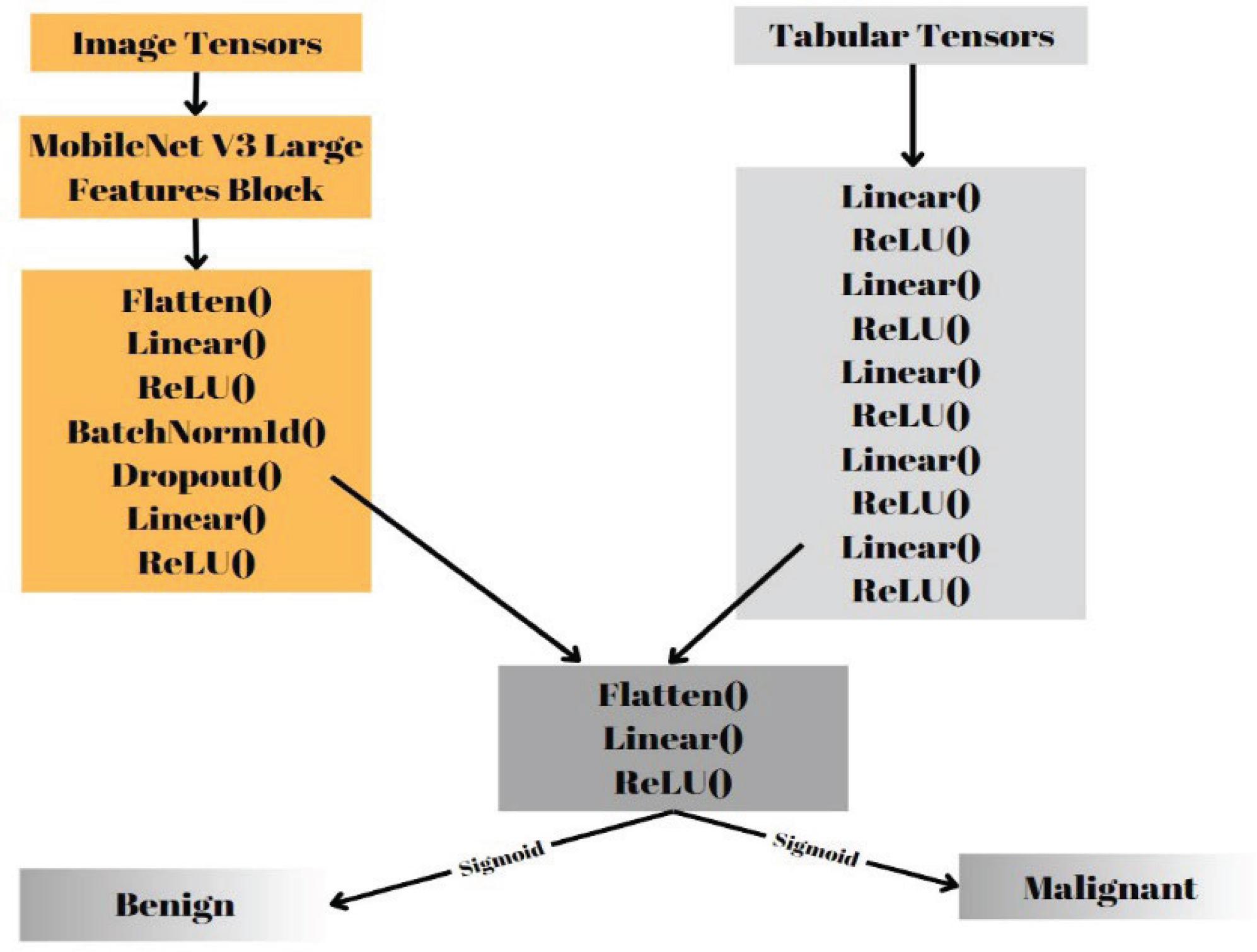
Figure 4.
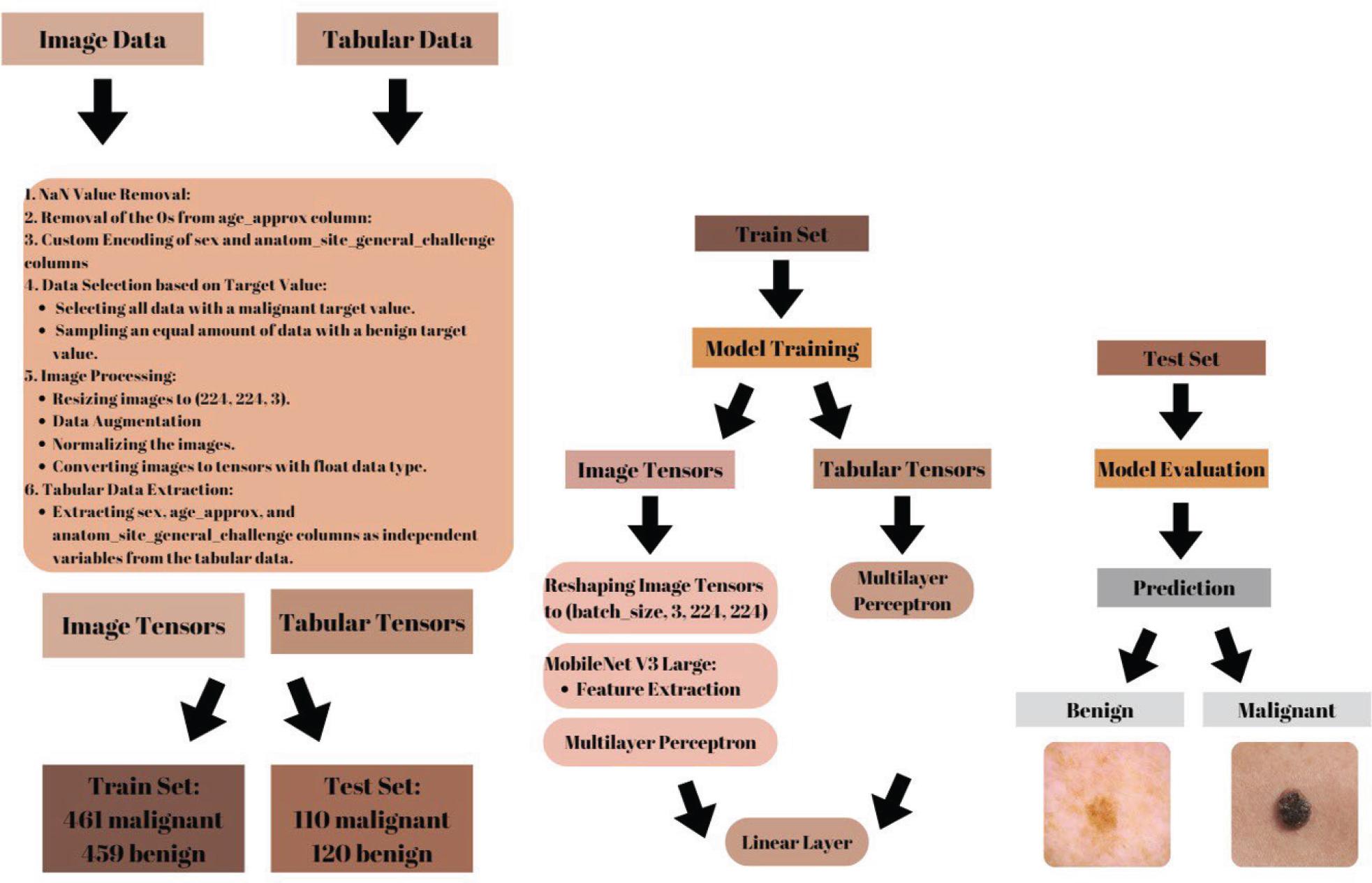
Figure 5.
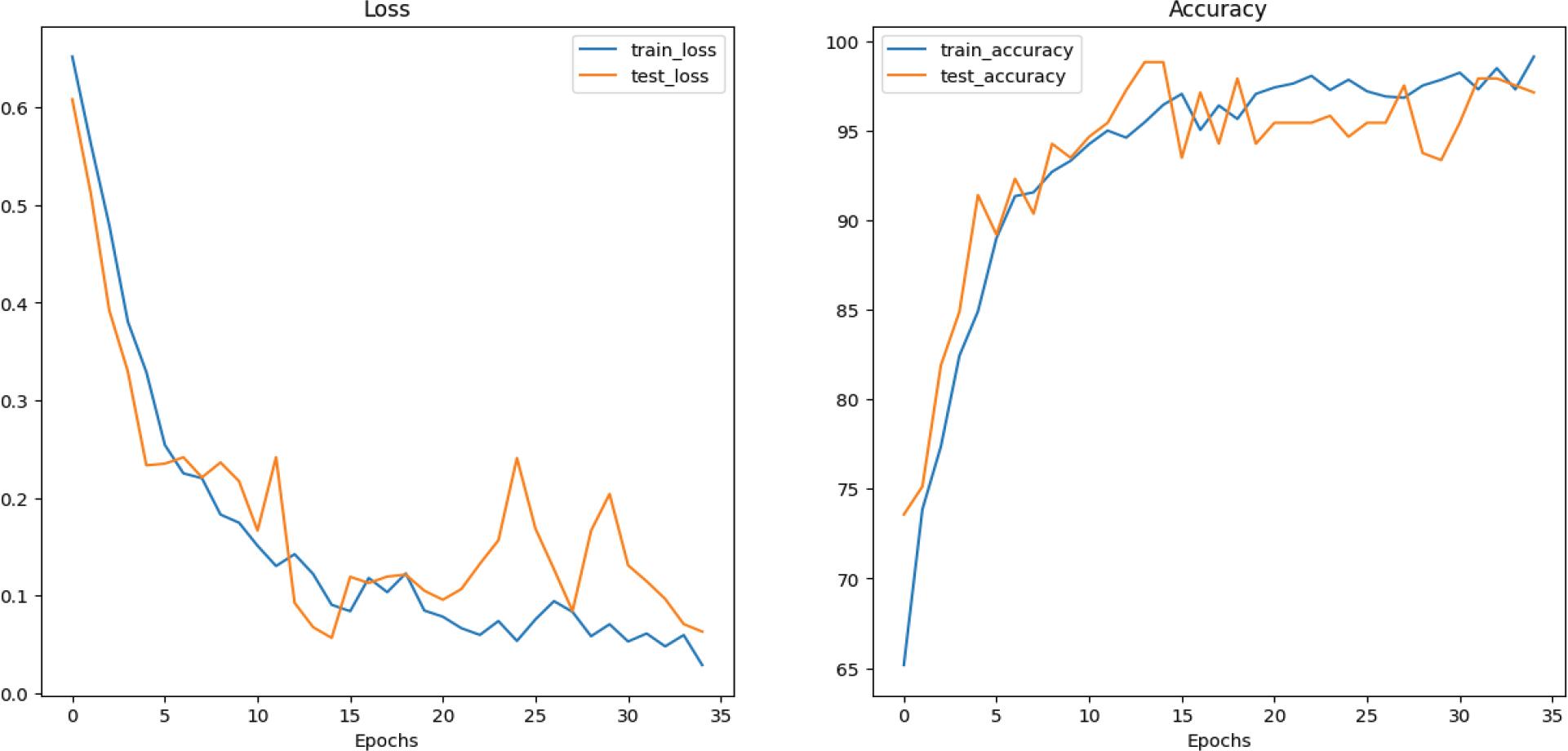
Figure 6.
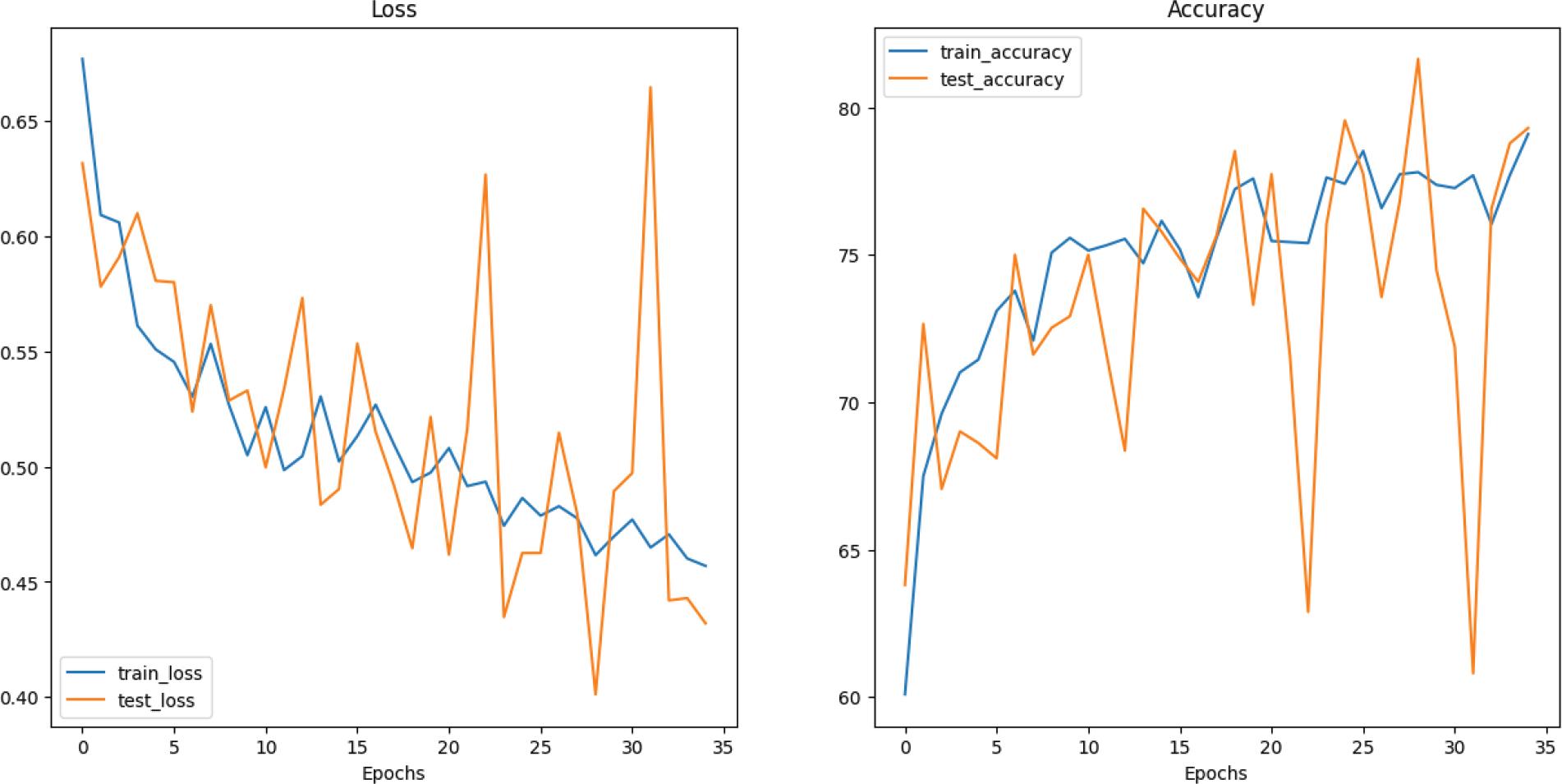
Figure 7.
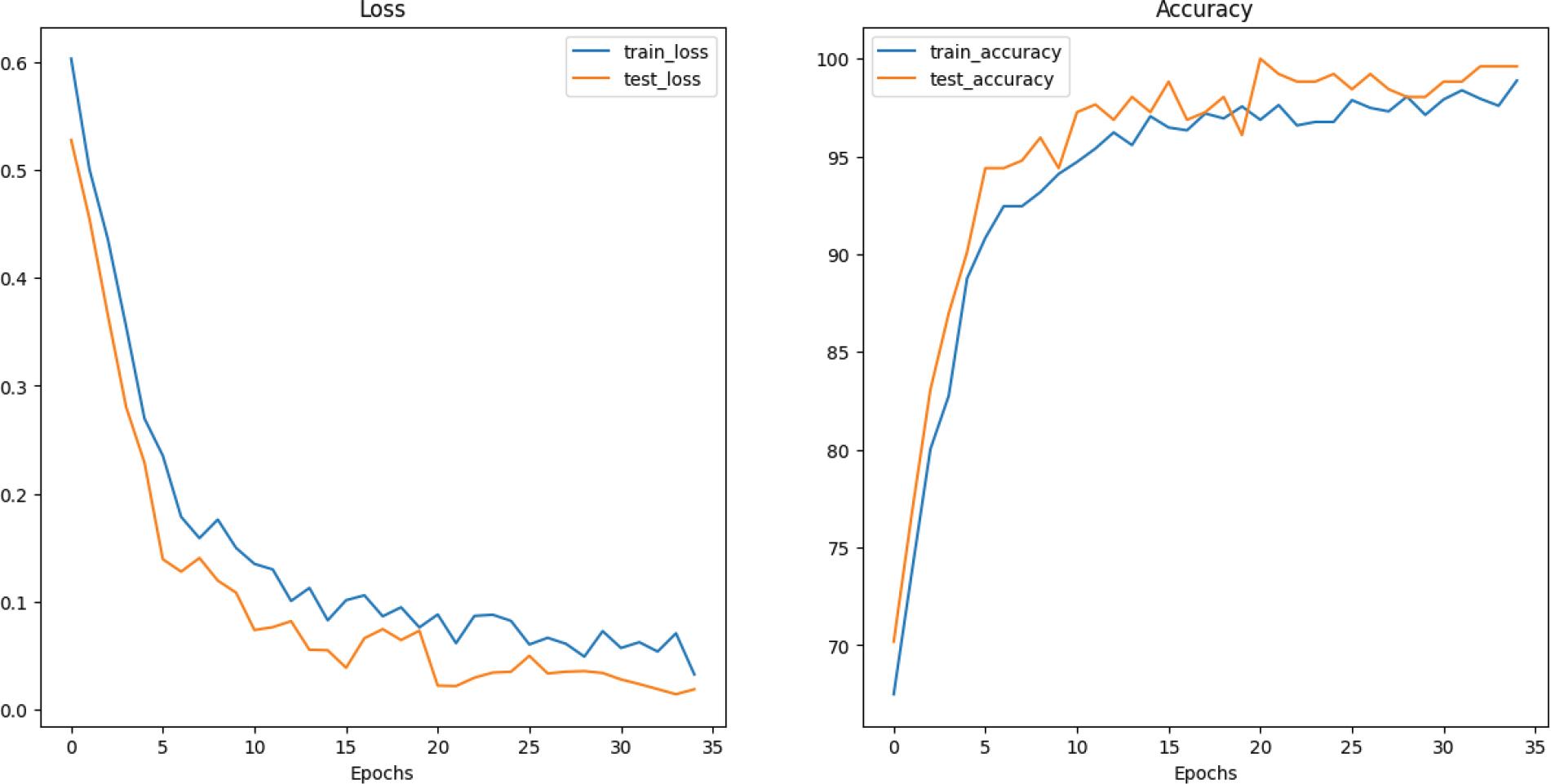
Figure 8.
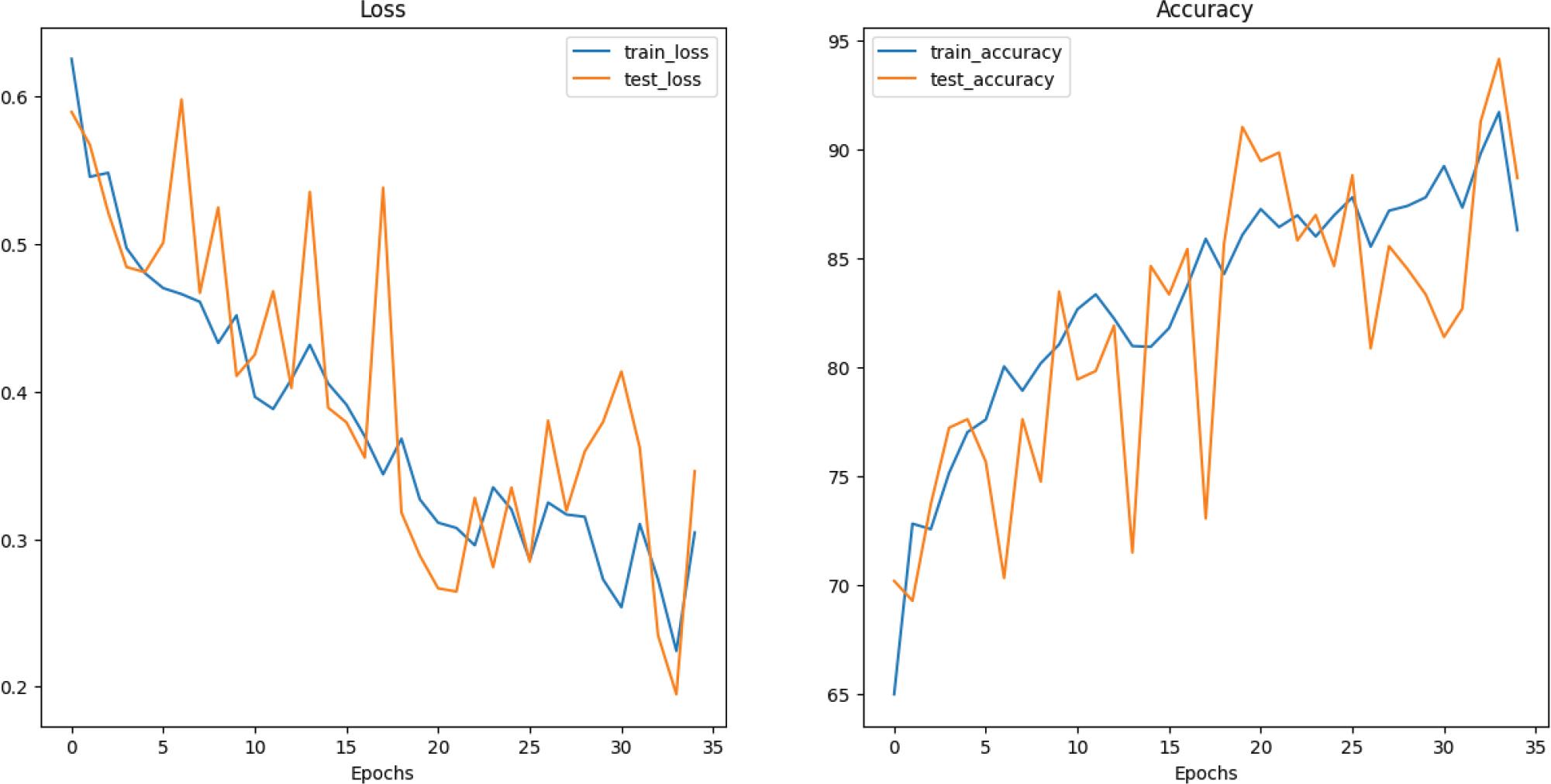
Figure 9.
Number of tabular and image data taken from SIIM-ISIC Dataset
| Class | Total | Training | Testing |
|---|---|---|---|
| Malignant | 571 | 461 | 110 |
| Benign | 579 | 459 | 120 |
| Total | 1150 | 920 | 320 |
Performance on the test set
| Accuracy(%) | Precision(%) | Recall(%) | F1-Score(%) | |
|---|---|---|---|---|
| Effcientnet_v2_s | 91.74 | 96.33(B)/87.60(M) | 87.50(B)/96.36(M) | 91.70(B)/91.77(M) |
| Convnext_base | 77.39 | 82.07(B)/73.38(M) | 72.50(B)/82.72(M) | 76.99(B)/77.77(M) |
| Densenet161 | 98.69 | 97.56(B)/1.00(M) | 1.00(B)/97.27(M) | 98.76 (B)/98.61 (M) |
| Mobilenet_v3_large | 99.56 | 1.00(B)/99.09(M) | 99.16(B)/1.00(M) | 99.58(B)/99.54(M) |
| VGG16 | 87.39 | 85.83(B)/90.83(M) | 90.83(B)/85.83(M) | 88.26(B)/88.26(M) |
Confusion matrix on the test set
| Model | TP (B) | TN (M) | FN | FP |
|---|---|---|---|---|
| Effcientnet_v2_s | 105 | 106 | 15 | 4 |
| Convnext_base | 87 | 91 | 33 | 19 |
| Densenet161 | 120 | 107 | 0 | 3 |
| Mobilenet_v3_large | 119 | 110 | 1 | 0 |
| VGG16 | 109 | 92 | 11 | 18 |
Skin cancer detection methodologies and their respective datasets
| Reference | Methodology | Dataset(s) | Evaluation Metrics |
|---|---|---|---|
| [22] | The preprocessing images and fnetuning convolutional neural networks with transfer learning, with EffcientNet B4 identifed as the top-performing model. | HAM10000 dataset | F1 Score: 87%, Accuracy: 87.91% |
| [23] | Automated Skin-Melanoma Detection (ASMD) system using image processing and SVM-based classifcation, proposing a Melanoma-Index (MI) for clinical use. | DD image dataset | Accuracy: 97.50% |
| [24] | Automatic skin cancer diagnosis system including Histogram of Gradients (HG) and Histogram of Lines (HL), combined with other features. | HPH dermoscopy database and the Dermoft standard database | Accuracy: 98.79% (HPH) and 92.96% (the standardDermoft) |
| [25] | Skin cancer detection system utilizing Genetic Programming (GP) for evolving a classifer and feature selection. | PH2dataset | Accuracy: 97.92% |
| [26] | Image processing and deep learning techniques, including Convolutional Neural Networks (CNNs), for skin cancer detection and classifcation. | MNISTHAM10000 dataset | Weighted Average Accuracy: 0.88, WeightedAverage Recall: 0.74, Weighted F1-score: 0.77 |
| [27] | Classifcation of skin lesions, utilizing dynamic-sized kernels and both ReLU and leakyReLU activation functions. | HAM10000 dataset | Overall accuracy: 97.85% |
| [28] | Soft-Attention mechanism in deep neural architectures for skin lesion classifcation. | HAM10000 dataset and ISIC-2017 dataset | Precision: 93.7% (HAM10000), sensitivity: 91.6% (ISIC-2017) |
| [29] | MobileNetV3 introducing the Improved Artifcial Rabbits Optimizer (IARO) algorithm to enhance feature selection | PH2, ISIC-2016, and HAM10000 datasets | Accuracy: 87.17% (ISIC-2016), 96.79% (PH2 dataset), and 88.71% (HAM10000) |
| [30] | SkinTrans, an improved transformer network, for skin cancer classifcation, utilizing vision transformers (VIT) with self-attention mechanism. | HAM10000 and clinical datasets | Accuracy: 94.3% (HAM10000) and 94.1% (Clinical) |
