Multidrug-resistant (MDR) Klebsiella pneumoniae presents a significant public health threat, exacerbated by the transmission of carbapenemase genes. K. pneumoniae carbapenemase (KPC) is one of the most widespread carbapenemases in China, first identified in 1996 in North Carolina (Yigit et al. 2001). The blaKPC gene, which encodes KPC, is typically located on transferable plasmids. In ST258 K. pneumoniae, blaKPC is commonly found within the transposon Tn4401, which varies in length and has a high potential for horizontal gene transfer (Bush and Bradford 2020; Queenan and Bush 2007). Continuous reports of outbreaks involving KPC-harboring K. pneumoniae highlight the ongoing challenge posed by these pathogens (Bush and Bradford 2020).
The most prevalent sequence types (STs) of KPC-harboring K. pneumoniae are ST258 and ST11 (Chen et al. 2014). ST258 poses a significant threat primarily in North America, Latin America, and Europe, while ST11 is predominant in Asia and Latin America (Chen et al. 2014). ST111 is a rarely reported type of K. pneumoniae, and its clinical relevance remains uncertain. As analyzed in our previous study (Jia et al. 2021), among the clinically prevalent K. pneumoniae types in China, ST111 K63 K. pneumoniae was most closely related to ST692 K64 and ST65 K2 in the phylogenetic tree determined by MLST. Holt et al. (2015) identified that ST111 is homologous to ST228 in the phylogenetic tree of core gene SNPs. To date, at least 8 O-antigens, based on the LPS, and 77 K-antigens, based on the CPS, have been identified (Follador et al. 2016). The K63 serotype has been reported to be susceptible to bacteriophage infection and has been used in bacteriophage research (Gao et al. 2022; Gorodnichev et al. 2021). To date, only sporadic cases of severe infection associated with K63 K. pneumoniae strain have been reported, with no evidence of sustained outbreaks.
Multidrug resistance (MDR) and virulence are typically considered incompatible in K. pneumoniae. However, with the emergence of mobile resistance or virulence elements, MDR and HvKp have recently emerged as clinically significant pathogens (Lee et al. 2017; Russo and Marr 2019; Wyres et al. 2020). Currently, the generation of MDR HvKp can be classified into three patterns: typical HvKp clones acquiring additional MDR factors (plasmids or elements), typical MDR clones acquiring additional hypervirulent factors, and typical clones acquiring hybrid vectors of both MDR and hypervirulence (Lan et al. 2021; Wyres et al. 2020).
However, genomic prediction alone cannot fully determine a strain’s biological virulence, especially when a hypervirulent lineage acquires a large, multidrug-resistant plasmid. Acquisition of such plasmids can impose a fitness cost and potentially attenuate virulence, resulting in a “genotypic hypervirulent” strain that is not highly pathogenic in vivo. Therefore, to validate whether AZJ065 remains hypervirulent after acquiring the KPC-harboring plasmid, we conducted in vivo infection experiments using Galleria mellonella and a mouse intraperitoneal model. These experiments were designed to test the hypothesis that AZJ065 retains high virulence comparable to that of classic hypervirulent strains, despite being string test-negative and multidrug-resistant. The in vivo data thus provide essential functional evidence to complement the genomic and phylogenetic analyses, confirming the clinical threat posed by the evolutionary convergence of resistance and virulence.
The aims of this study were to characterize the antimicrobial resistance and virulence phenotypes of a carbapenem-resistant hypervirulent Klebsiella pneumoniae strain, AZJ065, belonging to sequence type ST111 and capsular type K63; to elucidate the genetic features of its virulence and KPC-harboring resistance plasmids; and to investigate the evolutionary relationships of clinical ST111 K63 strains using comparative genomic and phylogenetic analyses. Additionally, we aimed to determine whether the acquisition of the MDR plasmid compromised the virulence of AZJ065 in in vivo infection models.
Among the 239 Gram-negative bacteria causing bloodstream infections in a surveillance program across 24 representative hospitals in different regions of China in 2018 (Jia et al. 2021), only four K. pneumoniae strains (1.7%), belonging to ST111 K63, were collected. The strains, AZJ065, SYF150, SCQ159, and SZS111, were all isolated from blood cultures of infected patients (Jia et al. 2021) and were identified using the Vitek MS MALDI-TOF system at authors’ Hospital.
The K. pneumoniae strains, NTUH-K2044 (K2044) and QD110, were chosen as representatives of hypervirulent and hypovirulent K. pneumoniae, respectively, for use in the Galleria mellonella and mouse intraperitoneal challenge infection models (Jia et al. 2021).
The broth microdilution method was performed according to the Clinical and Laboratory Standards Institute (CLSI) M100 guidelines (CLSI 2023). The minimum inhibitory concentrations (MICs) of colistin, levofloxacin, ceftazidime-avibactam, amikacin, ciprofloxacin, aztreonam, ceftazidime, cefoxitin, ceftriaxone, ertapenem, meropenem, imipenem, cefepime, imipenem-relebactam, ceftolozane-tazobactam, and piperacillin-tazobactam were determined. MICs were measured only when the quality control strains (Escherichia coli ATCC 25922 and Pseudomonas aeruginosa ATCC 27853) yielded acceptable results.
The hypervirulence of the four clinical strains was assessed using the string test, the G. mellonella infection model, and the mouse intraperitoneal challenge infection model (Li et al. 2020; Russo et al. 2021; Russo and Marr 2019).
The string test was performed by streaking an overnight colony onto agar plates with a loop. If the length of the viscous strings exceeded 5 mm, the phenotype of the K. pneumoniae strain was defined as hypermuco-viscous or a positive result (Fang et al. 2004).
In the G. mellonella infection model, ten healthy larvae of G. mellonella were selected for each infection challenge. A microbial suspension of 0.5 McFarland standard was prepared using single overnight colonies on agar plates and was diluted 1:10 in normal saline (approximately 1×107 CFU/ml). Ten larvae were injected with 10 microliters of the suspension (1×105 CFU) per larva from the third left foot using a 20μl syringe. After injection, larvae were incubated in a sterile petri dish at 37°C, and survival percentages were observed at 12h, 15h, 18h, 24h, 36h, and 48h. K2044, QD110, and normal saline were used as hypervirulent, hypovirulent, and negative controls, respectively (Liu et al. 2022).
In the mouse intraperitoneal challenge infection model, six- to eight-week-old female CD1 mice were obtained from Vital River, Beijing, China, and infected by intraperitoneal inoculation with 1×107 CFU in 100 μl of logarithmic culture, washed with phosphate-buffered saline (PBS). Five biological replicates were performed. The animals’ survival was monitored every 12 hours for 7 days. K2044 and QD110 were used as hypervirulent and hypovirulent controls, respectively (Liu et al. 2022).
The animal protocols were reviewed and approved by the Institutional Animal Care and Use Committee (IACUC) of the Laboratory Animal Resources Center of Tsinghua University (2022-841).
The genomic analysis of strains in this study was based on a previous study (Zhu et al. 2021). Genomic DNA was extracted using the UltraClean® Microbial DNA Isolation Kit (MO BIO Laboratories, Inc.) and the whole-genome sequence was obtained on the PacBio Sequel platform (Zhu et al. 2016).
The next-generation sequencing of AZJ065 was used to correct polymerase errors during PacBio sequencing. Paired-end libraries were prepared from 5 μg of genomic DNA using the TruSeq DNA Sample Prep Kit (Illumina Inc., San Diego, California, USA) and then sequenced on the Illumina Genome Analyzer IIx with a read length of 2×150 nucleotides. The raw Illumina sequencing reads were trimmed at a threshold of 0.01 and then assembled de novo using Unicycler (Wick et al. 2017) from both short and long reads.
Annotations of genome sequences were conducted using the rapid prokaryotic genome annotation software Prokka (Seemann 2014). The assembled plasmids were initially analyzed using NCBI BLASTn to identify similar reference plasmids. First, the replication origin (repR or repA) was identified. Subsequently, gene prediction was performed using RAST, Glimmer, GeneMarkS, and Prodigal. Finally, the plasmids were compared with reference sequences. The preliminary annotation from Prokka (Seemann 2014) was mapped to the plasmids, and detailed comments on their coding sequences (CDS) and mobile genetic elements (MGE) were added according to the NCBI database. Genome comparisons among plasmids were performed to analyze evolutionary relationships and sequence homology. Plasmid circular structure maps were generated with BRIG software. FASTA sequences of virulence genes downloaded from the Virulence Factor Database (VFDB) were used to align with genes in AZJ065 using BLAST (coverage ≥50%, identity ≥90%). Antimicrobial resistance genes were identified using ResFinder 4.0.
Sequence Types (STs) were determined using SRST2 (Inouye et al. 2014), and serotype was analyzed using SerotypeFinder 2.0 (Joensen et al. 2015) based on the assembled contigs. To elucidate the phylogenetic characteristics of plasmids in AZJ065, the genome sequences of similar plasmids were downloaded from the NCBI database. The prediction of conjugative regions of self-transmissible mobile genetic elements (Llosa and Alkorta 2017), including the oriT region, relaxase, Type IV secretion system genes (T4SS), and Type IV coupling protein (T4CP) in plasmids, was performed on the oriTfinder website (https://tool-mml.sjtu.edu.cn/oriTfinder/oriTfinder.html). The phylogenetic analysis of AZJ065 and three ST111 K63 clinical strains was based on core-genome SNPs detected by MUMmer 3.23 using the assembled genome of AZJ065 as the reference (Delcher et al. 2002). We further included an ST111 strain, two prevalent HvKp strains, and two prevalent CRKp strains in the phylogenetic tree. MAFFT (Nakamura et al. 2018) was used to align the concatenated SNP sequences, and the phylogenetic tree was generated with FastTree (Price et al. 2009). Gene homology was conducted using Roary.
The difference was estimated using GraphPad Prism 9 (GraphPad Software, Inc., USA). The survival rate in the G. mellonella mouse intraperitoneal challenge infection model was tested using the Log-rank (Mantel-Cox) test.
The detailed clinical information of the four strains, AZJ065, SCQ159, SYF150, and SZS111, is listed in Table I. All four strains were collected from positive bloodstream cultures and originated from different hospitals. The infections occurred in elderly patients aged 66, 69, 79, and 82 years. They all suffered from invasive bloodstream infections accompanied by a duration of fever. Carbapenems, such as meropenem and imipenem/cilastatin, were administered to combat the infections caused by SCQ159, SYF150, and SZS111. The condition of the patients infected with SCQ159 or SZS111 improved, while the patient infected with AZJ065 was discharged from the hospital without a cure, and the patient infected with SYF150 died. The antimicrobial susceptibility results for these four strains are listed in Table I. Briefly, AZJ065 is a multidrug- and carbapenem-resistant strain, while the other three were susceptible to all tested antibiotics.
The clinical information and antimicrobial susceptibility of the clinical strains, AZJ065 and three ST111 K63 strains
| Strain | AZJ065 | SCQ159 | SYF150 | SZS111 | |
|---|---|---|---|---|---|
| Province | Zhejiang | Chongqing | Zhejiang | Shanghai | |
| Hospital | The First Affiliated Hospital of Zhejiang University | The First Affiliated Hospital of Zhejiang University | Sir Run Run Shaw Hospital | Zhongshan Hospital of Fu Dan University | |
| Gender | Male | Male | Male | Female | |
| Age (year) | 79 | 66 | 82 | 69 | |
| Department | ICU | Medical ward | ICU | Surgery | |
| Primary Infection Site | Urinary tract infection | Liver abscess | Intra-abdominal infection | Intra-abdominal infection | |
| Duration of fever (Day) | 6-14 | >14 | 6-14 | 3-15 | |
| Peak of fever (°C) | >39 | >39 | 38-39 | >39 | |
| Laboratory data* | WBC (1×109/I) | 10.1 | 35.25 | 17.5 | 6.78 |
| N% | 92.8 | 94.4 | 94.8 | 58.9 | |
| CRP (mg/l) | 101 | >90 | 256.2 | 11.5 | |
| PCT (ng/ml) | 0.77 | >200 | 5.43 | 2.53 | |
| ICU Admission | Yes | No | Yes | No | |
| Antibiotic use 2g q12h | Fluconazole, Cefoperazone sulbactam | Imipenem/cilastatin 1g q8h | Meropenem 0.5g q6h | Meropenem 1g q12h | |
| Cefoxitin 2g q8h | |||||
| Discharge status | Treatment failure | Improved | Died | Improved | |
| MIC (μg/ml) | Colistin (S≤2 mg/L) | ≤1 | ≤1 | ≤1 | ≤1 |
| Levofloxacin (S≤0.5 mg/L) | 1 | ≤0.5 | ≤0.5 | ≤0.5 | |
| Amikacin (S≤16 mg/L) | ≤4 | ≤4 | ≤4 | ≤4 | |
| Ceftazidime-avibactam (S≤8 mg/L) | 0.5 | ≤0.12 | 0.25 | ≤0.12 | |
| Ciprofloxacin (S≤0.25 mg/L) | 1 | ≤0.25 | ≤0.25 | ≤0.25 | |
| Aztreonam (S≤ 4 mg/L) | >128 | ≤1 | ≤1 | ≤1 | |
| Ceftazidime (S≤ 4 mg/L) | 32 | ≤1 | ≤1 | ≤1 | |
| Cefoxitin (S≤ 8 mg/L) | >16 | ≤2 | 8 | ≤2 | |
| Ceftriaxone (S≤ 1 mg/L) | >8 | ≤1 | ≤1 | ≤1 | |
| Ertapenem (S≤0.5 mg/L) | >4 | ≤0.06 | ≤0.06 | ≤0.06 | |
| Meropenem (S≤ 1 mg/L) | >16 | ≤0.12 | ≤0.12 | ≤0.12 | |
| Imipenem (S≤ 1 mg/L) | >16 | ≤0.12 | ≤0.12 | ≤0.12 | |
| Cefepime (S≤ 2 mg/L) | >32 | ≤1 | ≤1 | ≤1 | |
| Imipenem-relebactam (S≤ 1 mg/L) | 1 | 0.25 | 0.25 | ≤0.12 | |
| Ceftolozone-tazobactam (S≤ 2/4 mg/L) | >16 | ≤0.12 | 0.5 | 0.25 | |
| Piperacillin-tazobactam (S≤ 8 mg/L) | >64 | ≤2 | 8 | ≤2 | |
Laboratory data was collected on 1–3 days in the initial period of infection
WBC – white blood cell, N % – the proportion of neutrophils, CRP – C-reaction protein, PCT – procalcitonin; MIC – minimum inhibitory concentration, S≤ – the susceptible breakpoints of the antibiotic for K. pneumoniae
The virulence of the strains was assessed using several methods, with animal infection models considered the more definitive standard (Russo et al. 2021). The length of the viscous string for AZJ065 was <5 mm, indicating it is a non-hypermucoviscous strain. Although the hypermucoviscous phenotype plays a vital role in differentiating HvKp from classical Kp, not all hypervirulent strains (identified by the G. mellonella infection model) exhibited a hypermucoviscous phenotype in our study. We propose that the hypermuco-viscous phenotype is not a prerequisite for HvKp to exert hypervirulence. Therefore, the mortality rates of G. mellonella for the four ST111 K63 strains in this study were further evaluated and are shown in Fig. S1. The test (Log-rank (Mantel-Cox) test) showed no significant difference between the clinical strains and K2044 and a significant difference from QD110, indicating that all ST111 K63 clinical strains in this study exhibited equivalent virulence to K2044 in the G. mellonella infection model. All mice infected by AZJ065 or K2044 succumbed to infection within 2.5 days post-infection, although those infected by K2044 survived slightly longer with a statistical difference (p-value = 0.0442) (Fig. 1). This result reveals that AZJ065 is hypervirulent, as it demonstrated a more rapid death rate than the hypervirulent control strain, K2044.
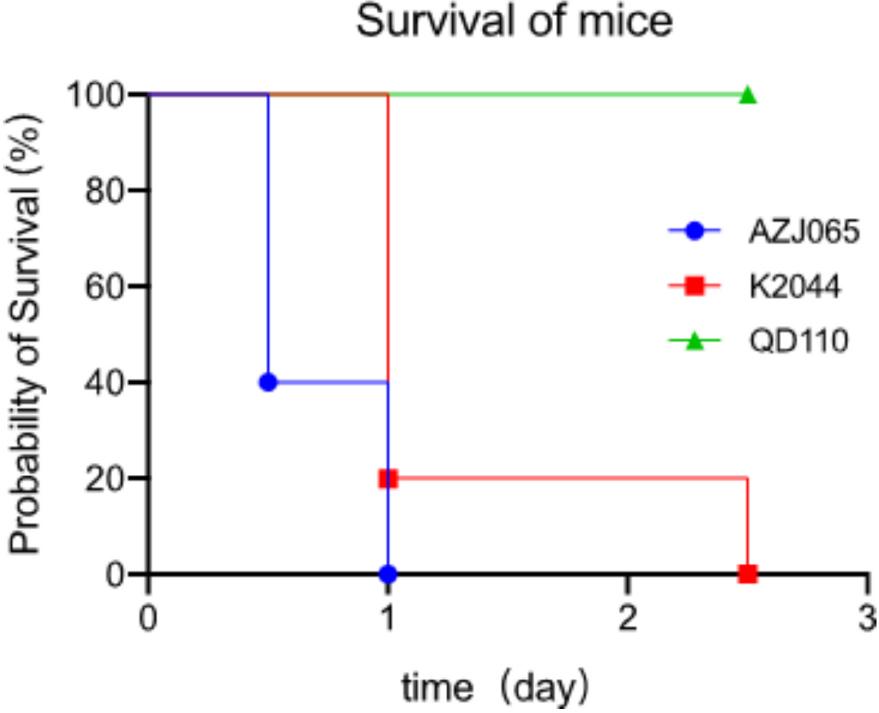
The survival curve of six-week CD1 mice after the intraperitoneal infection with the clinical strain, AZJ065 and the controls, NTUH-K2044 (K2044) and QD110. Each strain was tested by 1×107 CFU in 100μl PBS with five mice. P value from the log-rank (Mantel-Cox) test was 0.0442 (<0.05) between AZJ065 and K2044, indicating the virulence of AZJ065 was significantly different from that of K2044 and was even higher than that of K2044.
The genome sequences of AZJ065 were assembled into three circular replicons: a 5,273,117 bp chromosome, a 192,108 bp virulence plasmid (pAZJ065-Hv), and an 83,665 bp resistance plasmid (pAZJ065-KPC) (Fig. 2). The chromosome contains 5,088 coding genes, including three virulence genes or clusters (fyuA, mrkABCDFHIJ, ybtAEPQSTUX) and four resistance genes (blaSHV, fosA3, oqxA, oqxB), with an average gene size of 919 bp and a G+C content of 57.43%. The strain belonged to ST111 and serotype K63 O1. The two plasmids possessed a lower GC content, a lower proportion of coding regions, and shorter average gene lengths. There were three genes or clusters (iroBCDN, peg344, rmpA) on the virulence plasmid and eight resistance genes (dfrA14, sul2, blaKPC-2, arr-3, aph(6)- Id, aac(6’)-Ib, aac(6’)-Ib3, qnrS1) on the resistance plasmid. Neither plasmid was classified as hypervirulent or resistant. The annotated genome sequences of AZJ065, including the chromosome and two plasmids, have been deposited in the NCBI GenBank database under accession numbers CP099519, CP099520, and CP099521.
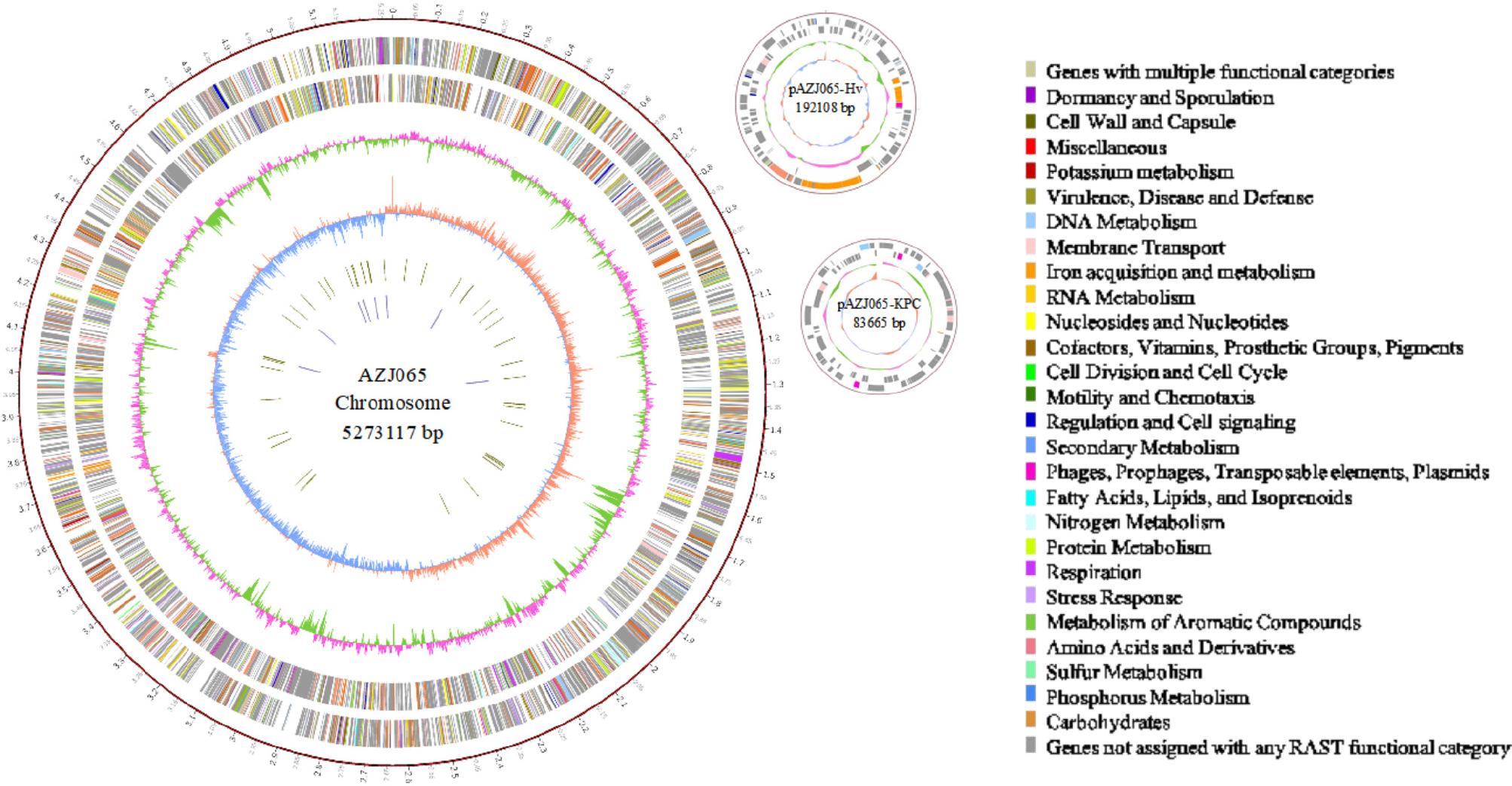
Circular representation of AZJ065 genome (chromosome and two plasmids). Circles are shown as follows (outside to inside): (1) A physical map scaled in megabases (Mb) from base 1 (the start of the putative replication origin); (2) coding sequences transcribed in a clockwise direction; (3) coding sequences transcribed in a counter-clockwise direction; (4) G + C content; (5) GC skew; (6) tRNA genes; (7) rRNA genes. Genes displayed in (2) and (3) are color-coded according to different functional categories shown on the right.
Given the presence of important virulence biomarker genes (iroBCDN, rmpA, and peg344) (Russo et al. 2018) on the plasmid pAZJ065-Hv, it was classified as a virulence plasmid. This plasmid, with a G+C content of 50.17%, harbored 202 coding genes, with an average gene size of 762 bp. It belonged to the IncHI1B incompatibility group and exhibited high similarity (query coverage of 100%, identity of 99.96%) with the 18.40-kb plasmid pM186-2 in K. pneumoniae. Notably, a forward-direction IS102 element was inserted into the siderophore gene clusters iroBCDN (between iroD and iroN) in pAZJ065-Hv (Fig. 3A and Fig. 3B). The iroBCDN in pAZJ065-Hv was found to be identical to that on the plasmid of K2044 (query coverage: 100%, identity: >99.9%), but the whole genomic sequence of pAZJ065-Hv shared very low coverage (35%). No self-transmissible mobile genetic elements were predicted in this plasmid.
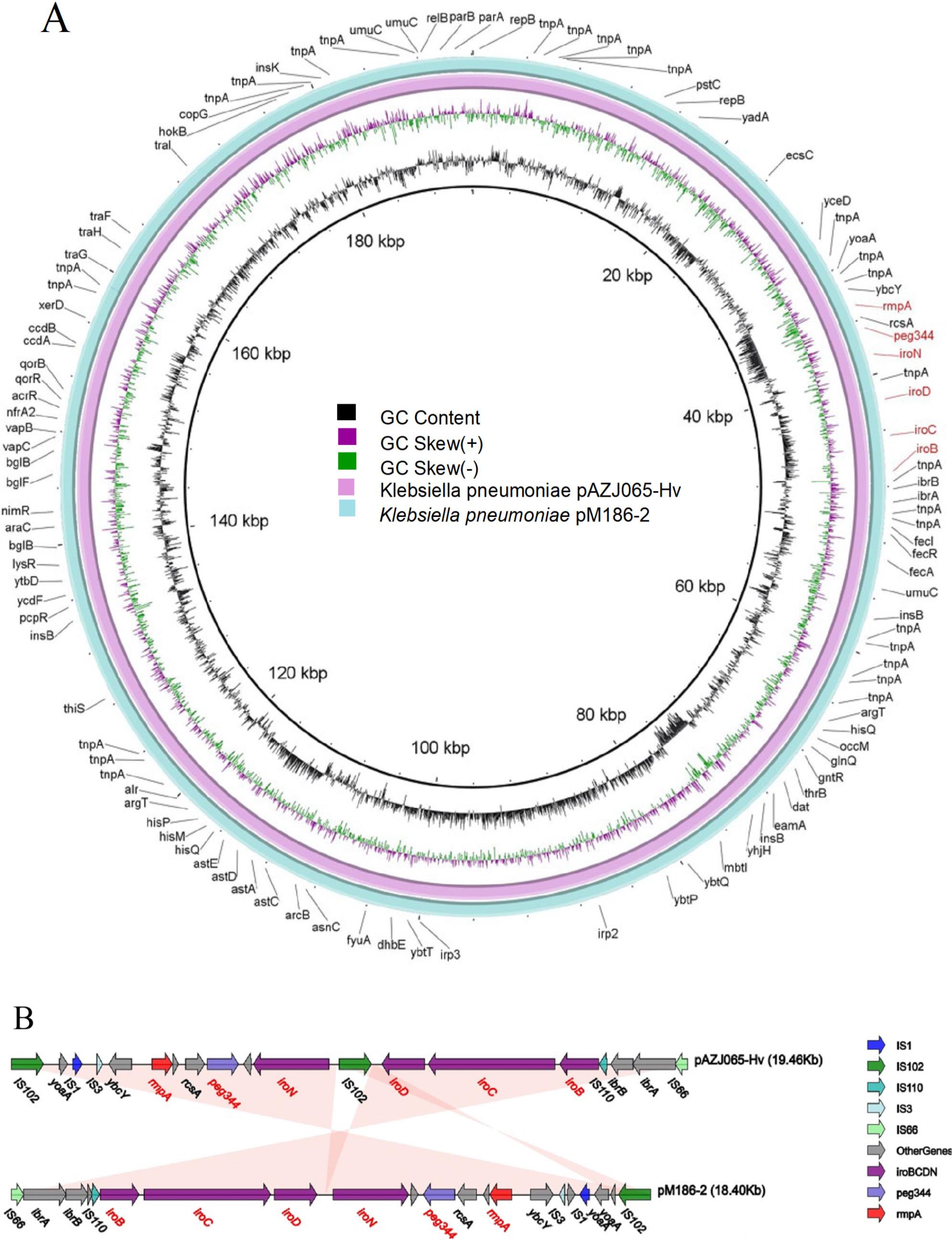
Genomic structure comparison of pAZJ065-Hv and pM186-2. (A) Alignments of resemble plasmids are shown as concentric rings. The outermost shows the main coding genes of pAZJ065-Hv. Virulence genes are highlighted in red. (B) Linear structure comparison of region harboring virulence genes on plasmids pAZJ065-Hv and pM186-2. Genes, mobile elements, and other features are colored based on functional classification. Shading denotes the regions with high homology (>95% nucleotide identity).
Ninety-one coding genes were identified on the resistance-related plasmid pAZJ065-KPC, with an average gene size of 748 bp and a G+C content of 53.53%. Nine resistance genes were detected, including one trimethoprim resistance gene (dfrA14), one sulphonamide resistance gene (sul2), four aminoglycoside resistance genes (aph(3’’)-Ib, aph(6)-Id, aac(6’)-Ib3, aac(6’)-Ib-cr), one carbapenemase gene (blaKPC-2), one rifampicin resistance gene (arr-3), and one quinolone resistance gene (qnrS1). These antimicrobial resistance genes contribute to the multidrug-resistant (MDR) phenotype of AZJ065, which is consistent with the antimicrobial resistance testing results (Table I) except for amikacin (broth microdilution MIC ≤4 μg/ml). The resistance plasmid pAZJ065-KPC belonged to the IncU type and presented a structure with the highest query coverage in BLAST of 87% with a KPC-3-harboring plasmid in Raoultella planticola (pFDAARGOS_430), followed by a KPC-3-harboring plasmid in Escherichia coli strains (query coverage of 80%) and a KPC-2-harboring plasmid in a K. pneumoniae strain (query coverage of 79%). The plasmid structure comparison of pAZJ065-KPC, K. pneumoniae p24169- KPC, Escherichia coli pBK32602, and Raoultella planticola pFDAARGOS_430 is shown in Fig. 4A. The main backbone regions of the four strains were similar, with pAZJ065-KPC having the most similar backbone structure to pBK32602. Notably, the accessory modules were different from the other three drug resistance regions. The drug resistance region on pAZJ065-KPC is novel and can be divided into three different parts. The region harboring three aminoglycoside-resistance genes, including sul2, aph(3’’)-Ib, and aph(6)- Id, was highly consistent with pFDAARGOS_430 and pBK32602 (Fig. 4B). The region harboring blaKPC-2 was highly consistent with p24169-KP. Another resistance region harboring the rifampicin-resistance gene arr-3 and the aminoglycoside-resistance gene aac(6’)-Ib3 showed no similar structure.
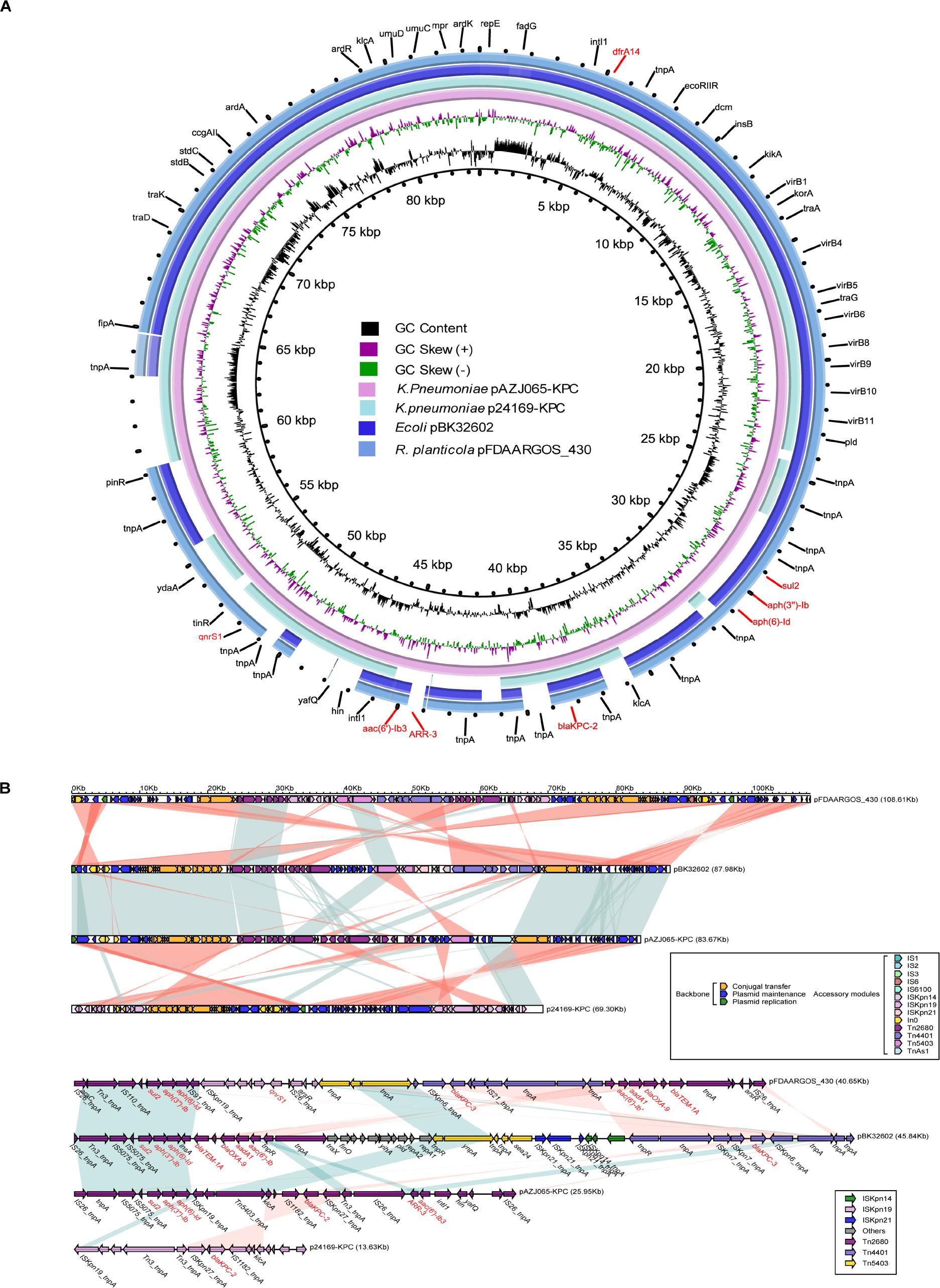
Genomic structure comparison of pAZJ065-KPC and three most similar plasmids (Klebsiella pneumoniae p24169-KPC, Escherichia coli pBK32602, Raoultella planticola pFDAARGOS_430). (A) Alignments of similar plasmids are shown as concentric rings. The outermost shows the main coding genes of pAZJ065-KPC. Resistance genes are highlighted in red. (B) The upper figure presents the related transposons and integrons comparison of plasmids pAZJ065-KPC and the three most similar plasmids in linear structure pattern. The lower one presents regions harboring resistance genes on plasmids. Genes, mobile elements, and other features are colored based on functional classification. Shading denotes the regions with high homology (>95% nucleotide identity).
The pAZJ065-KPC harbored three IS26 elements, forming a complete transposon, Tn2680, with two IS26 transposases located on both flanks of Tn2680 (Mollet et al. 1985), a transposon carrying the KPC gene, characteristic of a prokaryotic mobile genetic element. Moreover, the plasmid contained traM, traA, traB, traO, traF, traK, which belong to the T4SS; traD, which belongs to the T4CP; the oriT region; and relaxase. These genes constituted a complete conjugative system.
The results of the gene homology analysis of four ST111 K63 clinical strains (Fig. 5A) showed that although more than 90% of genes were core genes, AZJ065 was distinct from the other three strains. The analysis of virulence genes (Table S1) revealed that the virulence genes of AZJ065 and SCQ159 were identical, while those of SYF150 and SZS111 were identical, consistent with the whole-genome-wide SNP phylogenetic tree (Fig. 5B).
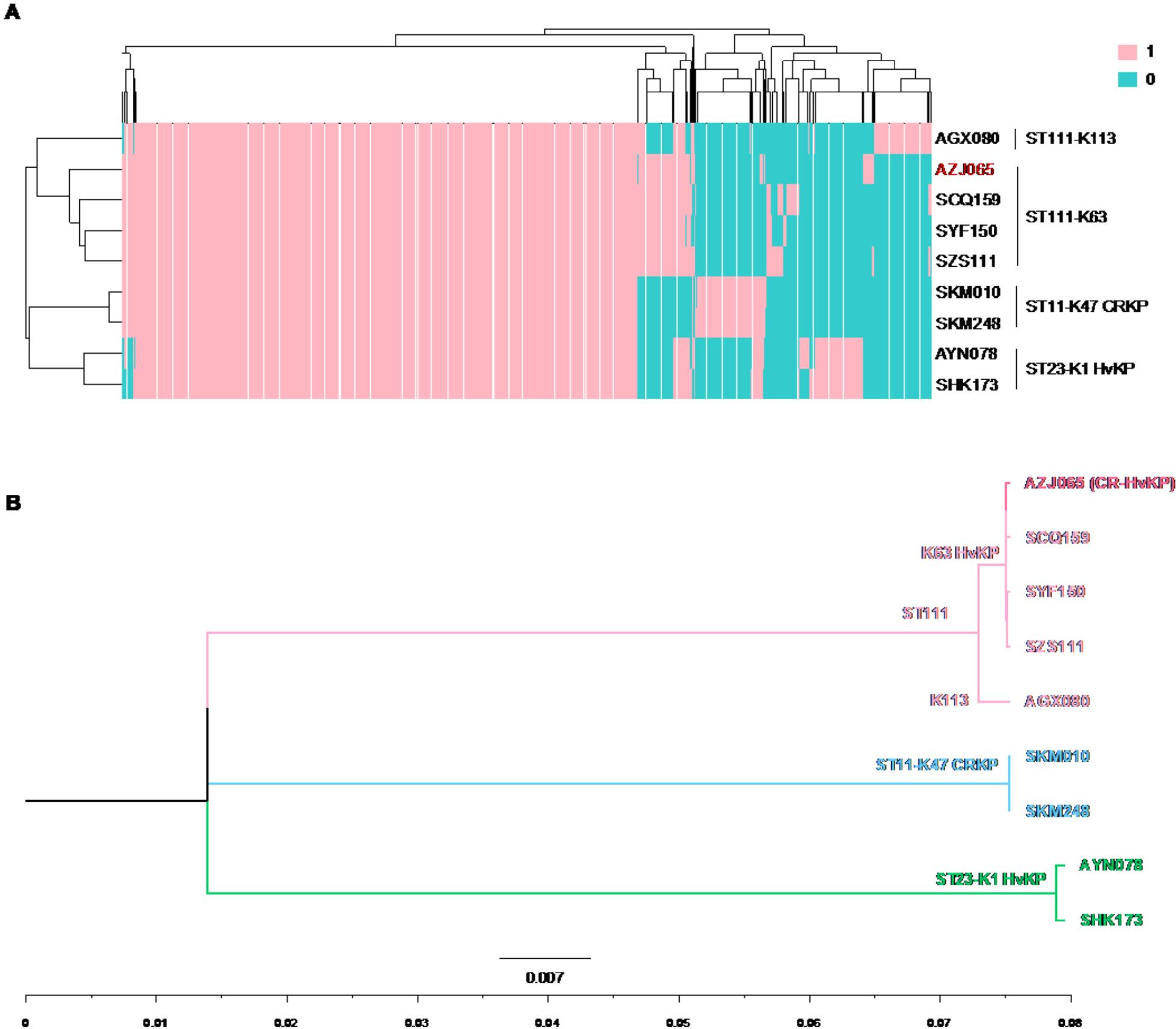
(A) Gene homology cluster of four ST111 K63 clinical strains. ‘1’ represents the presence of the genes in the strain and “0” represents the absence of the genes in the strain. (B) The phylogenetic tree of four ST111 K63 clinical strains based on core gene SNPs from whole-genome sequence.
Furthermore, analysis of six ST111 K63 K. pneumoniae genome sequences from the public database (NCBI GenBank) revealed that all ST111 K63 strains harbored three hypervirulence genes (iroBCDN, rmpA, and peg344) but lacked carbapenem resistance genes. It is strongly suggested that the basic genome feature of ST111 K63 contributes to hypervirulence, and that AZJ065 evolved into a carbapenem-resistant, hypervirulent strain by acquiring a resistance plasmid.
In this study, we describe a novel carbapenem-resistant, hypervirulent K. pneumoniae strain, ST111 K63. We identified a prevalence of 1.7% (4/239) of ST111 K63 strains in K. pneumoniae causing bloodstream infections in our study, and 0.6% (6/1014) of ST111 K63 strains among all 1014 K. pneumoniae strains with complete sequences in NCBI GenBank (as of January 21, 2021). Reports on the ST111 type are scarce and lack detailed genetic or phenotypic characterization (Jia et al. 2021; Shankar et al. 2018; Uz Zaman et al. 2014; Wang et al. 2020). The reported ST111 K. pneumoniae among neonates in a tertiary care centre in South India (Shankar et al. 2018) did not exhibit hypervirulence genes. In our study, all ST111 K63 O1 strains exhibited hypervirulence, and a hypervirulent and multidrug-resistant strain, AZJ065, was identified. Unlike strain AZJ065, which was non-hypermucoviscous (<5 mm), the other three ST111 K63 strains in this study and the control strain K2044 all presented a hypermucoviscous phenotype. The whole-genome sequences also displayed high similarity between the chromosomal genes of the four strains. These findings suggest that ST111 K63 MDR HvKp likely originated from hypervirulent clones acquiring MDR plasmids, providing evidence that AZJ065 evolved by acquiring an MDR KPC-harboring plasmid into a hypervirulent background.
Our in vivo experiments provided critical insights that genomic data alone could not reveal. Although phylogenetic analysis suggested AZJ065 evolved from a hypervirulent ancestor, genomic data cannot predict the biological impact of plasmid acquisition. The G. mellonella and mouse models demonstrated that AZJ065 induced high mortality, comparable to that of the classic hypervirulent strain K2044.
This finding is significant because it rules out the possibility of virulence attenuation due to the fitness cost of the MDR plasmid. It confirms that the evolutionary convergence of virulence and resistance in AZJ065 has resulted in a “superbug” that is both highly fatal and drug-resistant, underscoring the severe clinical threat posed by this ST111 K63 lineage.
The presence of several IS regions within the antimicrobial gene clusters suggests that pAZJ065-KPC was derived from an assembly of horizontally acquired DNA fragments. It also reveals the potential for transfer of drug resistance genes in the plasmid pAZJ065- KPC of the strain, including the carbapenem resistance gene blaKPC-2. The plasmid pAZJ065-KPC exhibited a lower transfer burden and more favorable transfer conditions, which might negatively affect future control of antimicrobial resistance. Furthermore, pAZJ065-KPC possessed a complete conjugative system, as predicted by the presence of conjugation elements. The essence of plasmid conjugation is DNA transport, which involves a DNA rolling-circle replication system (Llosa and Alkorta 2017; Llosa et al. 2002). The presence of a complete conjugative system and abundant IS elements indicates that pAZJ065-KPC tends to perform transferable novel hybrid hypervirulence-MDR plasmids or extensively drug-resistant plasmids.
In a study of 2503 K. pneumoniae genomes, the structures of iro loci in K. pneumoniae were categorized into five variants, one on the chromosome and the other four on plasmids (Lam et al. 2018). The research discovered that iro loci were not present in every ST111 K63 strain, and some strains had only one copy on a plasmid (Lam et al. 2018). The iro locus in AZJ065 is in accordance with this research and can be identified as the iro1 type, which is the one on the virulence plasmid in K2044 and prevalent in clonal groups (CG) 23, CG25, CG34, CG65, and CG85 (Lam et al. 2018). The deletion of iro loci was observed, and most of them were collected before 1960 (Lam et al. 2018).
This study identifies a carbapenem-resistant hypervirulent Klebsiella pneumoniae ST111 K63 O1 strain, AZJ065, carrying both a virulence plasmid and a KPC-harboring resistance plasmid with a unique genetic architecture. Comparative genomic analysis suggests that this strain likely evolved through the acquisition of a multidrug resistance plasmid by a pre-existing hypervirulent ST111 K63 lineage, rather than by stepwise accumulation of virulence determinants in a classical MDR background. Notably, the resistance plasmid pAZJ065-KPC harbors a complete conjugative transfer system and multiple insertion sequences flanking antimicrobial resistance regions, indicating a high potential for horizontal dissemination and structural rearrangement. This genetic configuration facilitates the mobilization of blaKPC into hypervirulent backgrounds and raises concern about the emergence of transferable hybrid plasmids that combine multidrug resistance and virulence-associated traits.
Such plasmid-mediated convergence represents a critical evolutionary mechanism driving the expansion of carbapenem-resistant hypervirulent K. pneumoniae. Although ST111 K63 remains a relatively uncommon lineage in clinical settings, the identification of a fully equipped conjugative KPC plasmid within a hypervirulent clone highlights a concrete risk for further dissemination and clonal expansion.
These findings emphasize the importance of surveillance efforts that focus not only on sequence types, but also on the acquisition of conjugative carbapenemase-encoding plasmids within hypervirulent K. pneumoniae lineages.
In conclusion, this study reports a carbapenem-resistant hypervirulent Klebsiella pneumoniae ST111 K63 strain carrying a newly structured KPC-harboring plasmid with a complete conjugative system. Our findings provide direct evidence that hypervirulent lineages can acquire highly transmissible carbapenem resistance plasmids, leading to the emergence of multidrug-resistant hypervirulent clones. The principal threat posed by this lineage lies in the plasmid-mediated convergence of hypervirulence and carbapenem resistance, rather than its current prevalence. Surveillance strategies should therefore prioritize the detection of conjugative carbapenemase-encoding plasmids, particularly those harboring blaKPC genes and abundant insertion sequences, within hypervirulent K. pneumoniae backgrounds. Early identification of such genetic configurations may be critical for preventing the establishment and dissemination of high-risk CR-HvKp clones in clinical settings.