Fig. 1.
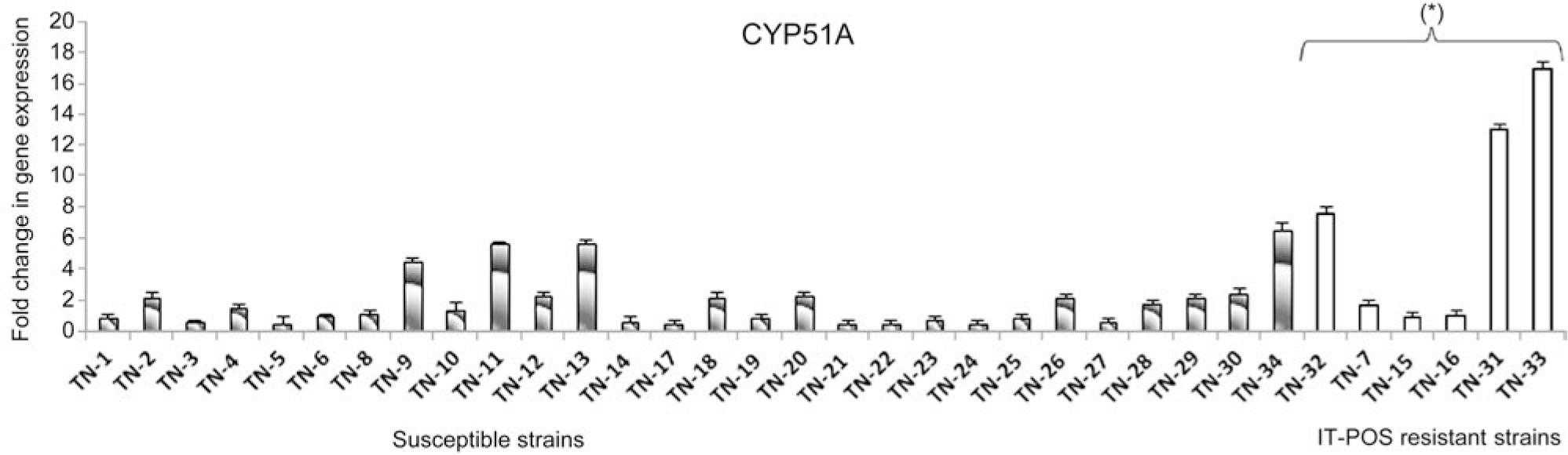
Fig. 2.
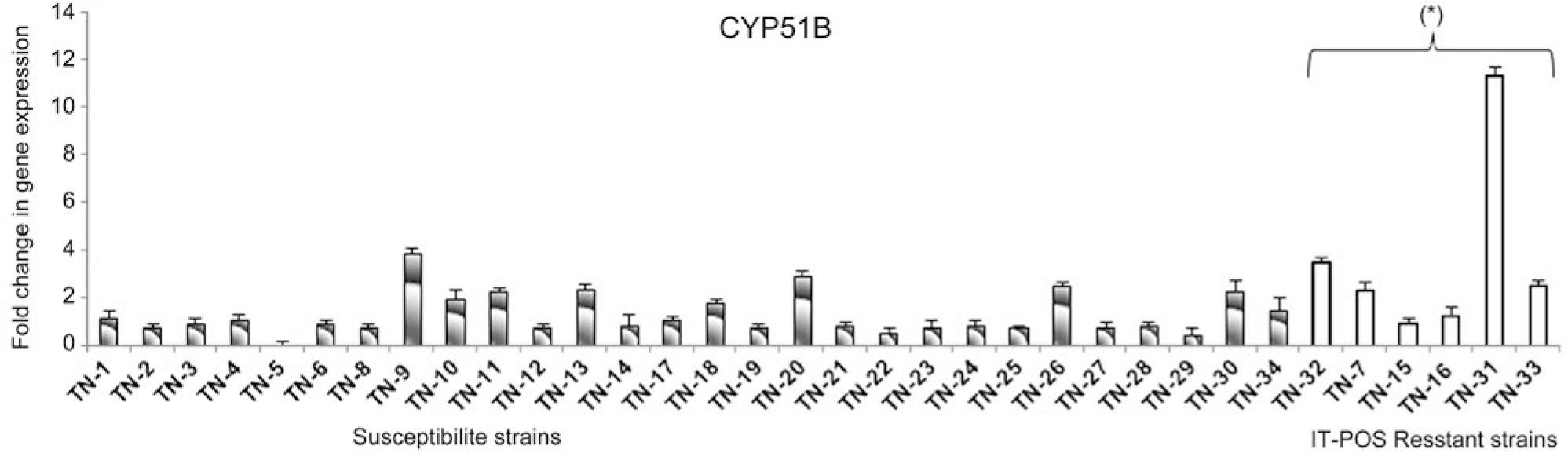
Fig. 3.
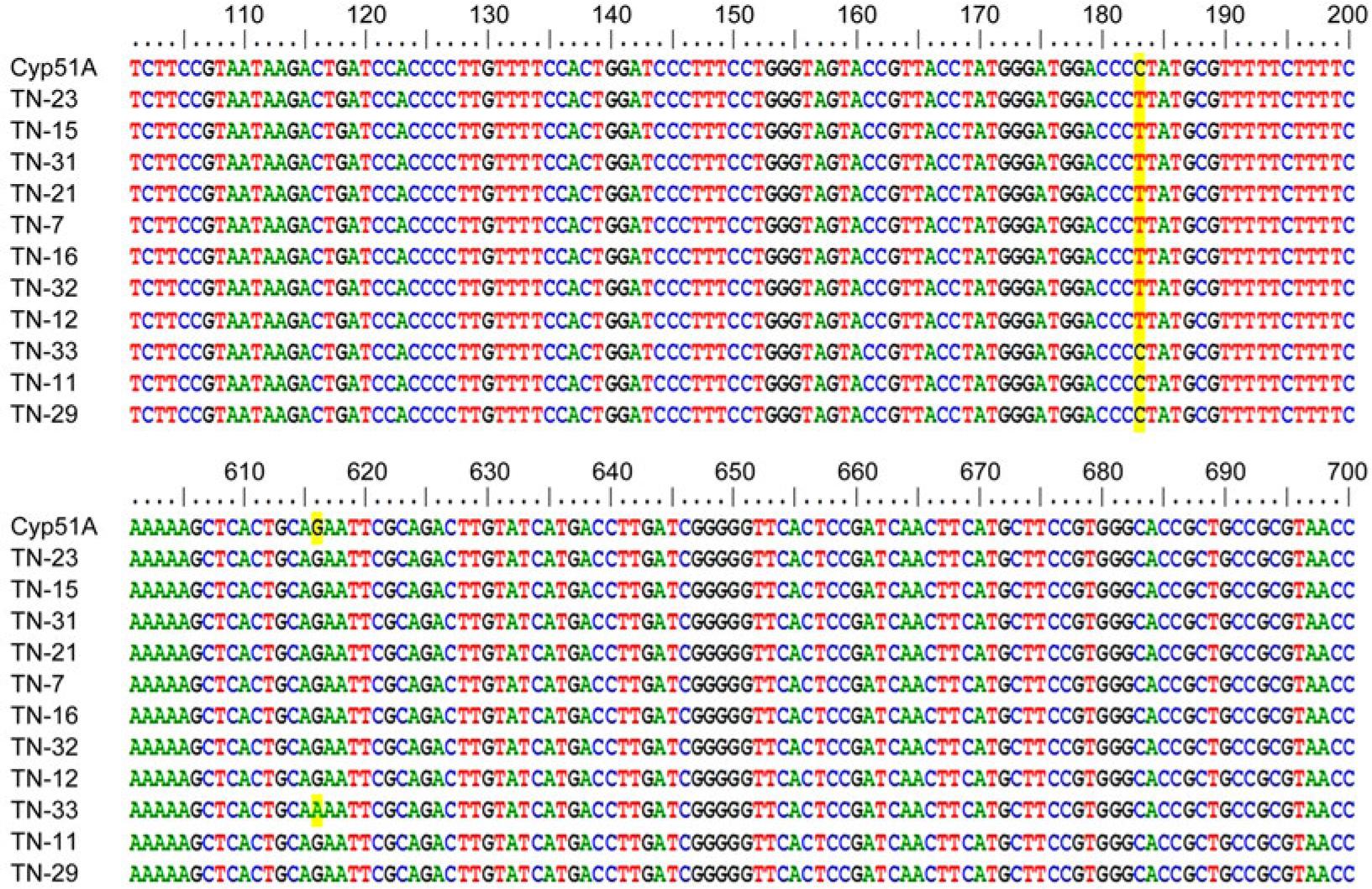
Fig. 4.
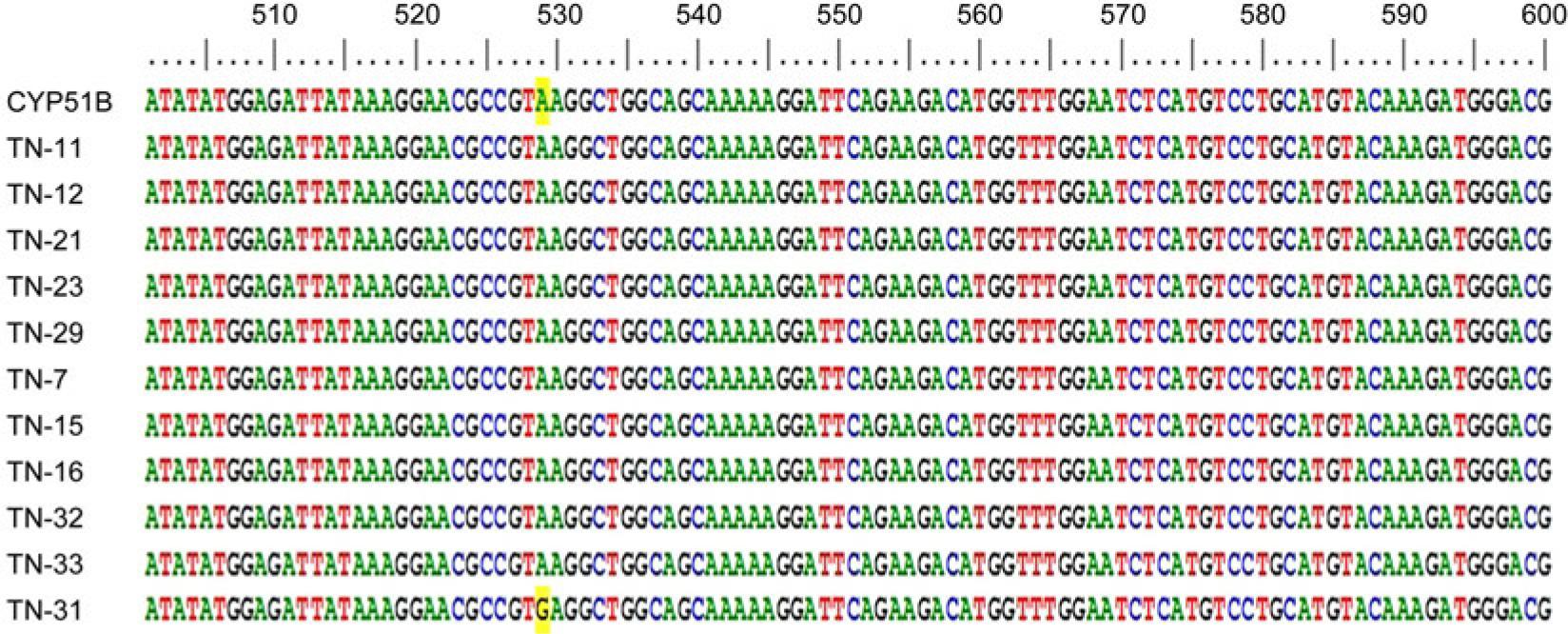
Fig. 5.
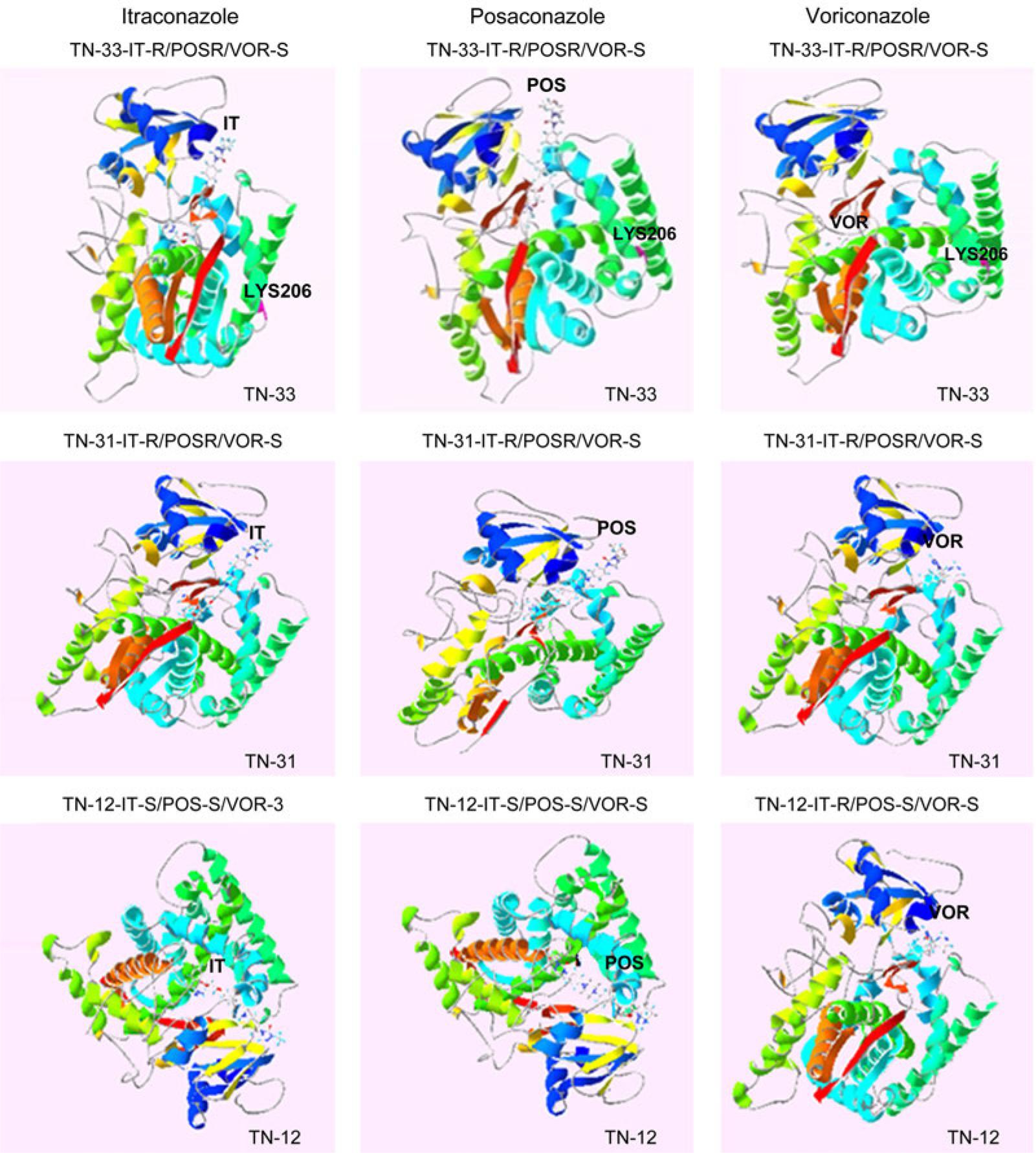
Fig. 6.
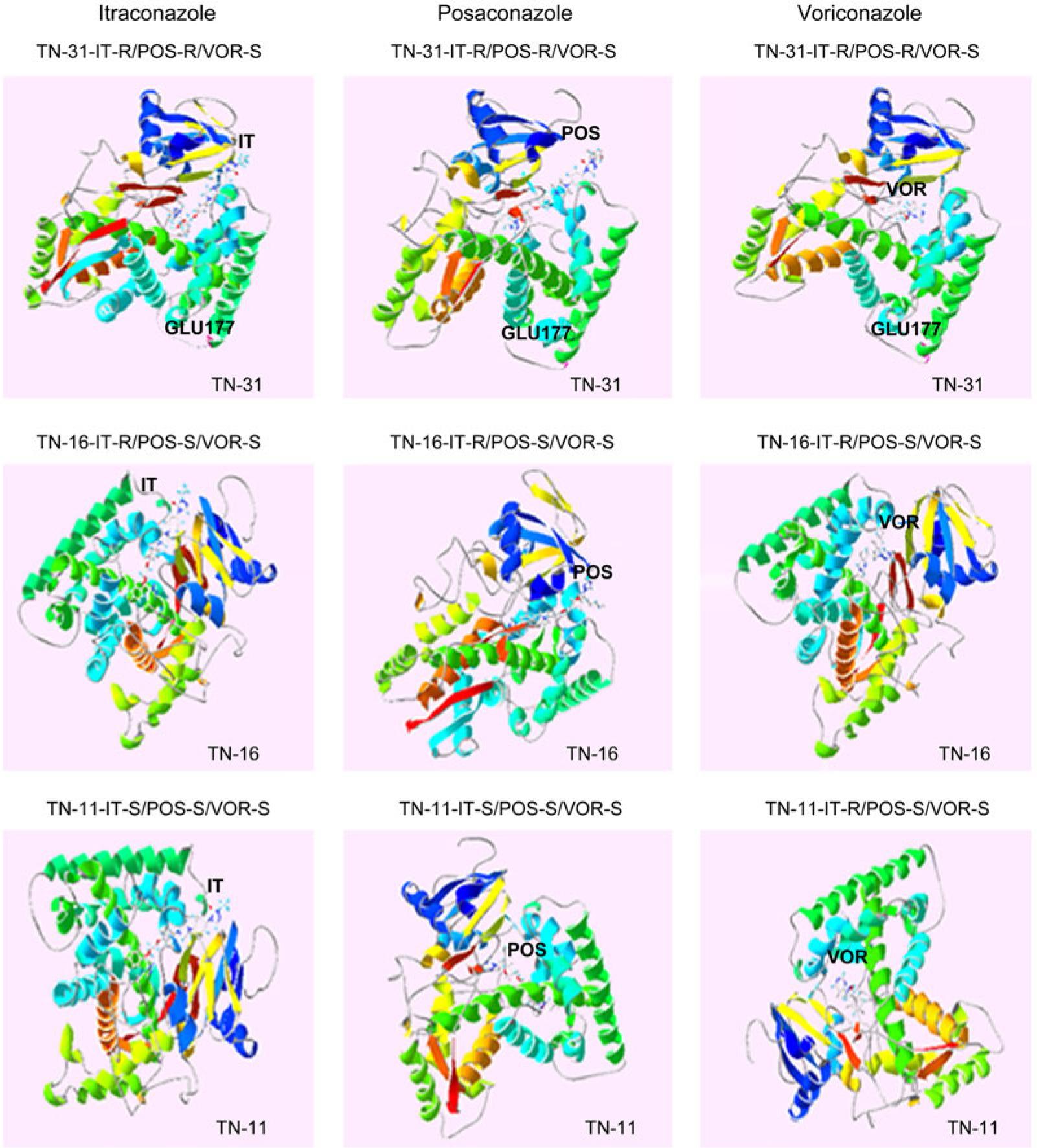
The sequences of primers and probes used in RT-qPCR_
| Gene | Primers and probes | |
|---|---|---|
| cyp51A | F | 5’-GCGCGCATGAGGGAGAT-3’ |
| R | 5’-CAATGCATGAGGTTCCAGATCA-3’ | |
| Probe | HEX-TCATTAACGAGCGCCGCAAGAACC-MGB | |
| cyp51B | F | 5’-ATTCGACTCGACATTTGCTGAA-3’ |
| R | 5’-GCATCACGCTTGCGGTTAT-3’ | |
| Probe | FAM-CATGATCTCGACATGGGTTTTGCCC-MGB | |
| ANXC4 | F | 5’-CCAACCCATAAACGCTCTGT-3’ |
| R | 5’-TGGTGGGAATCTTGGAGAAC-3’ | |
| Probe | CY5-ATCGAAGCAGCCTGTCTCAT-MGB | |
The sequences of primers used in PCR sequencing_
| Gene Primers | ||
|---|---|---|
| cyp51A | AP1F | 5’-ATGGCATCCTTCACTCTCGT-3’ |
| AP1R | 5’-CG ATCAACTTCATGCTTCCG-3’ | |
| AP2F | 5’-TCTGGAACCTCATGCATTGT-3’ | |
| AP2R | 5’-TCCCTCGAAACCAGCAATTA-3’ | |
| AP3F | 5’-GGCAGGGTCGAAATCACGGA-3’ | |
| AP3R | 5’-GCCCGGATGGAAGAACCCTT-3’ | |
| cyp51B | BP1F | 5’-CTTTTATCGGAAGTACCATC-3’ |
| BP1R | 5’-CTTTATCAGGAACAACTTCG-3’ | |
| BP2F | 5’-ACTCTTCGCATACATGCACC-3’ | |
| BP2R | 5’-CCAACGTGCATGATATTGCC-3’ | |
| BP3F | 5’-CGACTCGACATTTGCTGAAC-3’ | |
| BP3R | 5’-CTCTTCGCATACATGCACCA-3’ | |
Antifungal susceptibility to the azoles, relative quantification of gene expression, gene copy number of cyp51A and cyp51B genes, and mutation in Aspergillus flavus_
| Strain | GenBank accession | Isolation site | Isolation date | Itraconazole | Posaconazole | RNA relative quantification | DNA relative quantification | Cyp51A | Cyp51B | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| MIC (μg/ml) | R/S | MIC (μg/ml) | R/S | cyp51A | ±SD | cyp51B | ± SD | cyp51A | ±SD | cyp51B | ± SD | GenBank accession | Punctual mutation | GenBank accession | Punctual mutation | ||||||
| Patient 1 | Invasive aspergillosis | TN-1 | MN964040 | nasal | 15/10/2018 | 0.125 | S | 0.125 | S | 0.90 | ±0,02 | 1.16 | ±0,04 | 0.69 | ±0,05 | 0.57 | ±0,02 | ND | ND | ND | ND |
| TN-2 | MN964041 | sputum | 15/10/2018 | 0.125 | S | 0.125 | S | 2.15 | ±0,03 | 0.73 | ±0,02 | 0.89 | ±0,02 | 1.98 | ±0,04 | ND | ND | ND | ND | ||
| TN-3 | MN964042 | sputum | 22/10/2018 | 0.125 | S | 0.125 | S | 0.58 | ±0,01 | 0.90 | ±0,02 | 1.37 | ±0,02 | 0.97 | ±0,02 | ND | ND | ND | ND | ||
| Patient 2 | Invasive aspergillosis | TN-4 | MN964043 | nasal | 15/10/2018 | 0.125 | S | 0.19 | S | 1.53 | ±0,02 | 1.05 | ±0,04 | 1.27 | ±0,03 | 1.93 | ±0,02 | ND | ND | ND | ND |
| TN-5 | MN964044 | sputum | 15/10/2018 | 0.125 | S | 0.064 | S | 0.51 | ±0,04 | 0.05 | ±0,02 | 1.05 | ±0,02 | 1.92 | ±0,06 | ND | ND | ND | ND | ||
| TN-6 | MN964045 | nasal | 23/10/2018 | 0.032 | S | 0.094 | S | 0.95 | ±0,01 | 0.89 | ±0,02 | 1.69 | ±0,03 | 0.93 | ±0,02 | ND | ND | ND | ND | ||
| Patient 3 | Invasive aspergillosis | TN-7 | MN964046 | nasal | 30/10/2018 | 1.5 | R | 0.125 | S | 1.65 | ±0,02 | 0.73 | ±0,03 | 2.51 | ±0,02 | 1.98 | ±0,02 | MN964023 | C-T*(183) | MN964034 | No mutation |
| TN-8 | MN964047 | sputum | 30/10/2018 | 0.5 | S | 0.094 | S | 1.12 | ±0,02 | 0.73 | ±0,02 | 1.13 | ±0,07 | 5.97 | ±0,04 | ND | ND | ND | ND | ||
| Patient 4 | Invasive aspergillosis | TN-9 | MN964048 | sputum | 30/10/2018 | 0.75 | S | 0.125 | S | 4.48 | ±0,02 | 3.89 | ±0,02 | 2.71 | ±0,02 | 2.67 | ±0,02 | ND | ND | ND | ND |
| TN-10 | MN964049 | nasal | 30/10/2018 | 0.38 | S | 0.064 | S | 1.42 | ±0,02 | 1.97 | ±0,04 | 2.49 | ±0,03 | 2.01 | ±0,03 | ND | ND | ND | ND | ||
| Patient 5 | Invasive aspergillosis | TN-11 | MN964050 | sputum | 10/11/2018 | 0.5 | S | 0.125 | S | 5.65 | ±0,01 | 2.31 | ±0,02 | 1.87 | ±0,02 | 1.02 | ±0,02 | MN964018 | No mutation | MN964029 | No mutation |
| TN-12 | MN964051 | BAL | 17/11/2018 | 0.125 | S | 0.125 | S | 2.35 | ±0,02 | 2.31 | ±0,02 | 1.87 | ±0,02 | 1.02 | ±0,02 | MN964019 | C-T*(183) | MN964030 | No mutation | ||
| Patient 6 | Invasive aspergillosis | TN-13 | MN964052 | BAL | 03/12/2018 | 0.75 | S | 0.19 | S | 5.68 | ±0,02 | 2.35 | ±0,05 | 2.83 | ±0,05 | 3.95 | ±0,02 | ND | ND | ND | ND |
| TN-14 | MN964053 | sputum | 26/11/2018 | 0.5 | S | 0.125 | S | 0.63 | ±0,03 | 0.88 | ±0,06 | 6.95 | ±0,07 | 1.06 | ±0,02 | ND | ND | ND | ND | ||
| TN-15 | MN964054 | nasal | 26/11/2018 | 1 | R | 0.19 | S | 0.90 | ±0,02 | 0.76 | ±0,02 | 0.53 | ±0,02 | 1.65 | ±0,05 | MN964024 | C-T*(183) | MN964035 | No mutation | ||
| TN-16 | MN964055 | sputum | 10/11/2018 | 1 | R | 0.19 | S | 1.02 | ±0,02 | 1.22 | ±0,05 | 2.34 | ±0,04 | 2.84 | ±0,02 | MN964025 | C-T*(183) | MN964036 | No mutation | ||
| Patient 7 | Invasive aspergillosis | TN-17 | MN964056 | sputum | 01/03/2017 | 0.75 | S | 0.125 | S | 0.48 | ±0,02 | 1.10 | ±0,02 | 0.77 | ±0,02 | 2.07 | ±0,06 | ND | ND | ND | ND |
| TN-18 | MN964057 | BAL | 15/03/2017 | 0.5 | S | 0.125 | S | 2.18 | ±0,03 | 1.81 | ±0,02 | 2.11 | ±0,02 | 2.17 | ±0,02 | ND | ND | ND | ND | ||
| TN-19 | MN964058 | sputum | 08/03/2017 | 0.75 | S | 0.125 | S | 0.92 | ±0,02 | 0.76 | ±0,02 | 0.57 | ±0,02 | 1.11 | ±0,02 | ND | ND | ND | ND | ||
| TN-20 | MN964059 | sputum | 15/03/2017 | 0.38 | S | 0.064 | S | 2.27 | ±0,02 | 2.88 | ±0,05 | 0.64 | ±0,04 | 1.09 | ±0,02 | ND | ND | ND | ND | ||
| Patient 8 | Invasive aspergillosis | TN-21 | MN964060 | sputum | 08/03/2017 | 0.38 | S | 0.125 | S | 0.45 | ±0,02 | 0.88 | ±0,02 | 6.87 | ±0,02 | 1.82 | ±0,02 | MN964020 | C-T*(183) | MN964031 | No mutation |
| Patient 9 | Invasive aspergillosis | TN-22 | MN964061 | sputum | 01/03/2017 | 0.25 | S | 0.125 | S | 0.49 | ±0,02 | 0.55 | ±0,02 | 1.84 | ±0,02 | 1.68 | ±0,04 | ND | ND | ND | ND |
| Patient 10 | Invasive aspergillosis | TN-23 | MN964062 | sputum | 15/03/2017 | 0.38 | S | 0.125 | S | 0.69 | ±0,02 | 0.95 | ±0,03 | 1.73 | ±0,03 | 1.59 | ±0,02 | MN964021 | C-T*(183) | MN964032 | No mutation |
| Patient 11 | Invasive aspergillosis | TN-24 | MN964063 | sputum | 01/03/2017 | 0.38 | S | 0.125 | S | 0.51 | ±0,02 | 0.87 | ±0,02 | 1.62 | ±0,02 | 1.66 | ±0,09 | ND | ND | ND | ND |
| TN-25 | MN964064 | sputum | 08/03/2017 | 0.75 | S | 0.125 | S | 0.81 | ±0,02 | 0.75 | ±0,01 | 1.62 | ±0,01 | 1.48 | ±0,02 | ND | ND | ND | ND | ||
| TN-26 | MN964065 | BAL | 15/03/2017 | 0.75 | S | 0.125 | S | 2.21 | ±0,02 | 2.51 | ±0,02 | 0.67 | ±0,02 | 1.42 | ±0,02 | ND | ND | ND | ND | ||
| Patient 12 | Invasive aspergillosis | TN-27 | MN964066 | BAL | 03/12/2018 | 0.25 | S | 0.125 | S | 0.65 | ±0,02 | 0.78 | ±0,04 | 5.95 | ±0,03 | 1.03 | ±0,08 | ND | ND | ND | ND |
| TN-28 | MN964067 | nasal | 26/11/2018 | 0.25 | S | 0.125 | S | 1.73 | ±0,02 | 0.88 | ±0,02 | 1.95 | ±0,02 | 1.06 | ±0,02 | ND | ND | ND | ND | ||
| TN-29 | MN964068 | sputum | 26/11/2018 | 0.5 | S | 0.094 | S | 2.11 | ±0,02 | 0.48 | ±0,02 | 4.39 | ±0,02 | 1.95 | ±0,02 | MN964022 | No mutation | MN964033 | No mutation | ||
| Patient 13 | Invasive aspergillosis | TN-30 | MN964069 | sputum | 01/03/2017 | 0.5 | S | 0.094 | S | 2.37 | ±0,02 | 2.27 | ±0,08 | 0.68 | ±0,07 | 1.44 | ±0,03 | ND | ND | ND | ND |
| TN-31 | MN964070 | lung biopsy | 17/04/2017 | 1.5 | R | 0.75 | R | 12.99 | ±0,02 | 11.32 | ±0,02 | 3.07 | ±0,04 | 2.71 | ±0,02 | MN964026 | C-T*(183) | MN964037 | A-G*(529) | ||
| TN-32 | MN964071 | nasal | 01/03/2017 | 0.5 | S | 1 | R | 7.49 | ±0,02 | 3.51 | ±0,08 | 1.97 | ±0,02 | 0.77 | ±0,05 | MN964027 | C-T*(183) | MN964038 | No mutation | ||
| Patient 14 | Invasive aspergillosis | TN-33 | MN964072 | nasal | 21/03/2018 | 1 | R | 0.75 | R | 16.89 | ±0,02 | 2.48 | ±0,02 | 2.57 | ±0,06 | 1.71 | ±0,02 | MN964028 | G-A*(6I6) | MN964039 | No mutation |
| TN-34 | MN964073 | nasal | 28/03/2018 | 0.38 | S | 0.064 | S | 6.49 | ±0,02 | 1.52 | ±0,08 | 0.97 | ±0,02 | 0.77 | ±0,06 | ND | ND | ND | ND | ||