Fig. 1.

Fig. 2.

Fig. 3.
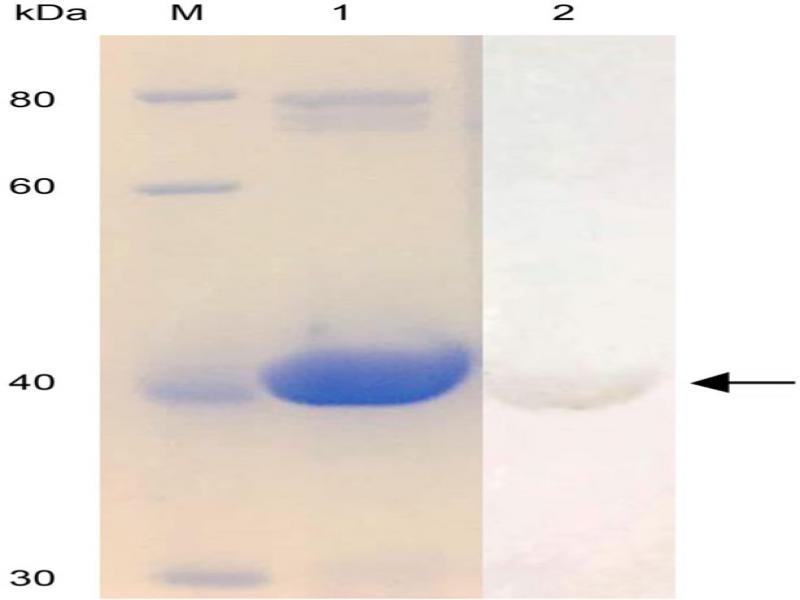
Fig. 4.

Fig. 5.

Fig. 6.

Fig. 7.

The results of antimicrobial activity of homogentisic acid and hydroquinone inhibitors against numerous isolates of Gram-positive and Gram-negative bacteria_
| Organism | aMIC (μg/ml) | |
|---|---|---|
| Hydro-quinone | Homogentisic Acid | |
| Streptococcus iniae HNM-1 | 25 (2.3) | 200 (5.6) |
| Escherichia coli DH5α | 130 (7.9) | 210 (8.4) |
| Salmonella typhimurium | 150 (8.7) | 180 (11.4) |
| Staphylococcus aureus | 210 (13.7) | 250 (14.1) |
| Acinetobacter baumannii | 180 (11.5) | 210 (12.3) |
| Pseudomonas aeruginosa | 0 | 0 |
Effect of Hydroxylamine, DTT and PLP on SiAlr Activity_
| Chemical | Concentration (mM) | Relative activity (%) |
|---|---|---|
| None | 100 (0.7) | |
| Hydroxylamine | 0.1 | 21 (1.2) |
| 1 | 11 (0.8) | |
| 10 | 9 (1.4) | |
| DTT | 1 | 27 (3.1) |
| 3 | 2 (0.8) | |
| PLP | 0.01 | 56 (2.4) |
| 0.04 | 83 (1.5) | |
| 0.06 | 96 (2.7) |
Strains and plasmids used in this study_
| Strains/Plasmids | Description | Source |
|---|---|---|
| Strains | ||
| Streptococcus iniae HNM-1 | Isolated from infected A. sinensis | This study |
| Escherichia coli DH5α | Used for cloning and propagation of plasmids | Novagen |
| Escherichia coli BL21(DE3) | Used for protein expression | Invitrogen |
| Salmonella typhimurium | This study | |
| Staphylococcus aureus | This study | |
| Acinetobacter baumannii | This study | |
| Pseudomonas aeruginosa | This study | |
| Plasmids | ||
| pMD19-T | Carries ampR gene; used for cloning PCR product with A at 3’ ends | Takara |
| pET 22b (+) | Carries ampR gene; used for expressing S. iniae Alanine racemase | Novagen |