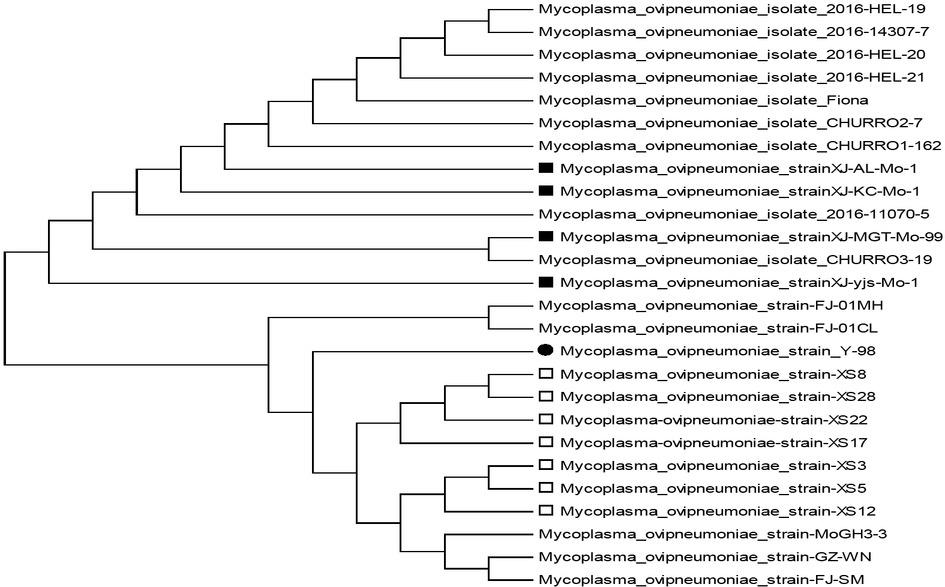
Fig. 1
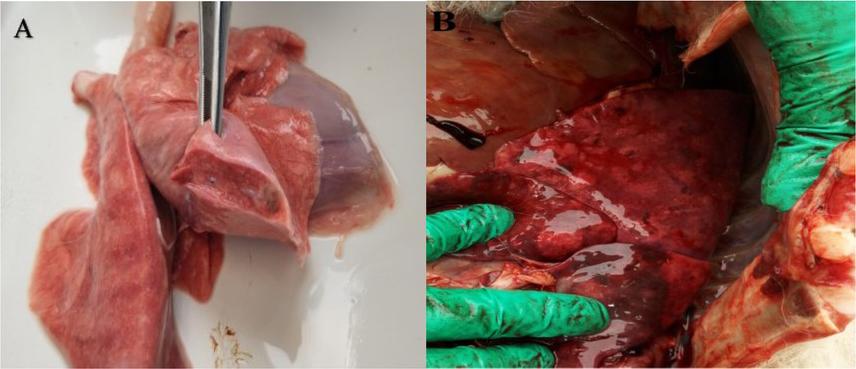
Fig. 2
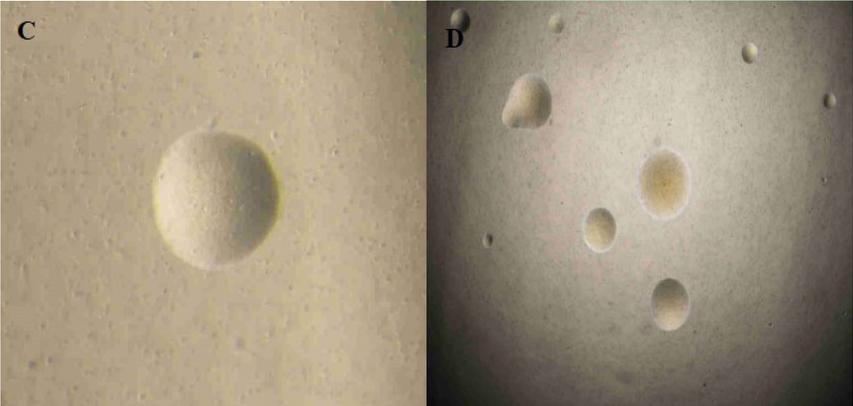
Fig. 3
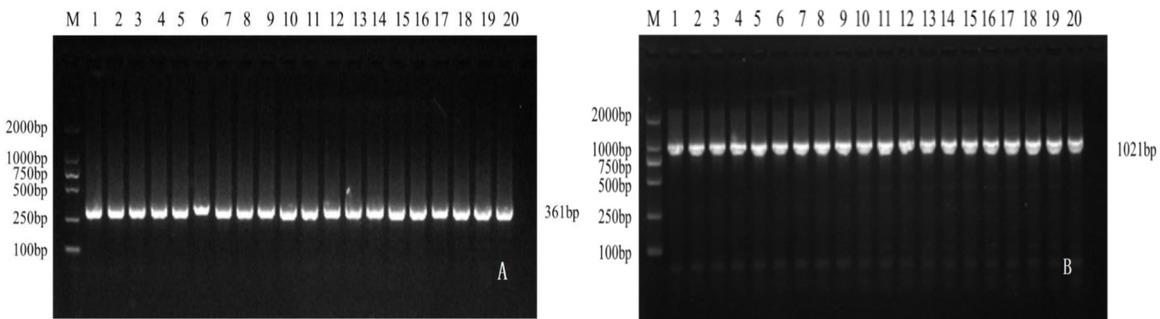
Fig. 4

Primers used in this study
| Species | Primer name | Primer sequence | Annealing temperature | Amplicon size (bp) | Reference |
|---|---|---|---|---|---|
| Mycoplasma genus | MGSO | TGCACCATCTGTCACTCTGTTAACCTC | 58°C | 1021 | (21) |
| M. ovipneumoniae | PP1 | GACTTCATCCTGCACTCTGT | 55°C | 361 | (17) |
M_ ovipneumoniae PCR detection results in nasal swabs
| Region | Examined | Positive | Prevalence (%) |
|---|---|---|---|
| Hotan | 200 | 76 | 38.00 |
| Kashgar | 242 | 128 | 52.89 |
| Aksu | 232 | 73 | 31.47 |
| Bazhou | 150 | 59 | 39.33 |
| Total | 824 | 336 | 40.78 |
M_ ovipneumoniae positive rates in nasal swabs by animal age
| Age (months) | Examined | Positive | Prevalence (%) | ||||||
|---|---|---|---|---|---|---|---|---|---|
| Kazak | Duolang | Total | Kazak | Duolang | Total | Kazak | Duolang | Total | |
| < 3 | 106 | 130 | 236 | 59 | 67 | 126 | 55.67 | 51.54 | 53.39 |
| 3-12 | 78 | 85 | 163 | 38 | 37 | 75 | 50.67 | 49.33 | 46.01 |
| >12 | 201 | 224 | 425 | 60 | 75 | 135 | 29.85 | 33.48 | 31.76 |
| Total | 385 | 439 | 824 | 147 | 189 | 336 | 38.10 | 43.05 | 40.78 |