Figure 1
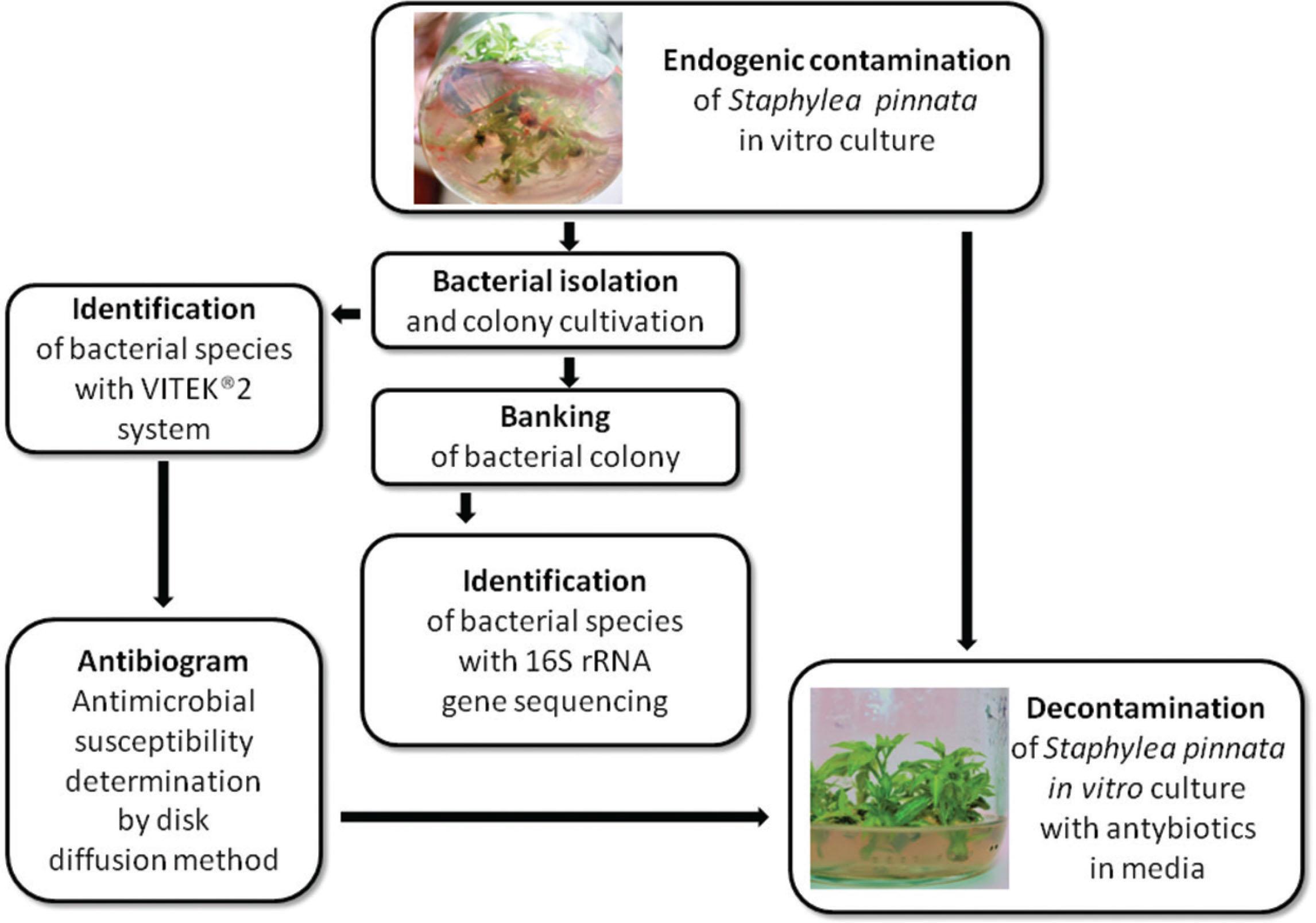
Figure 2
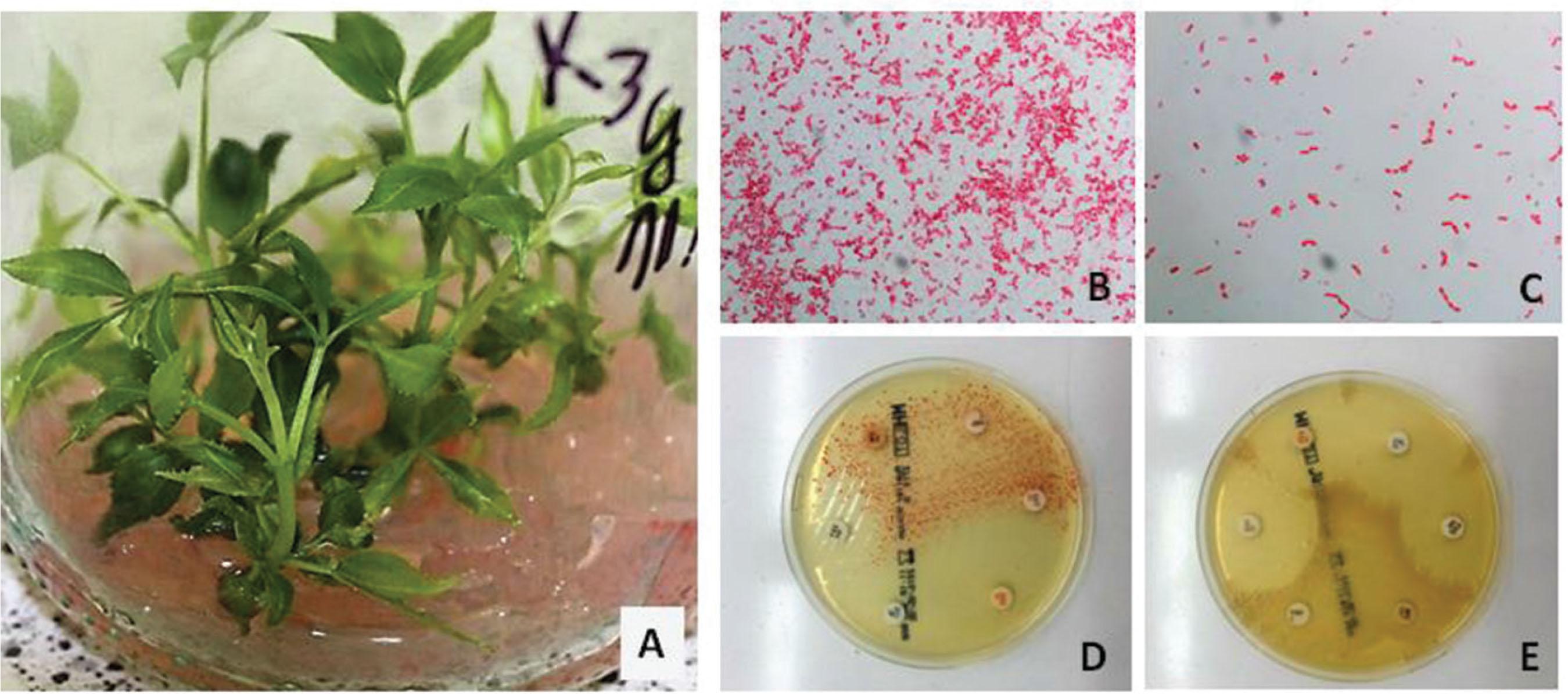
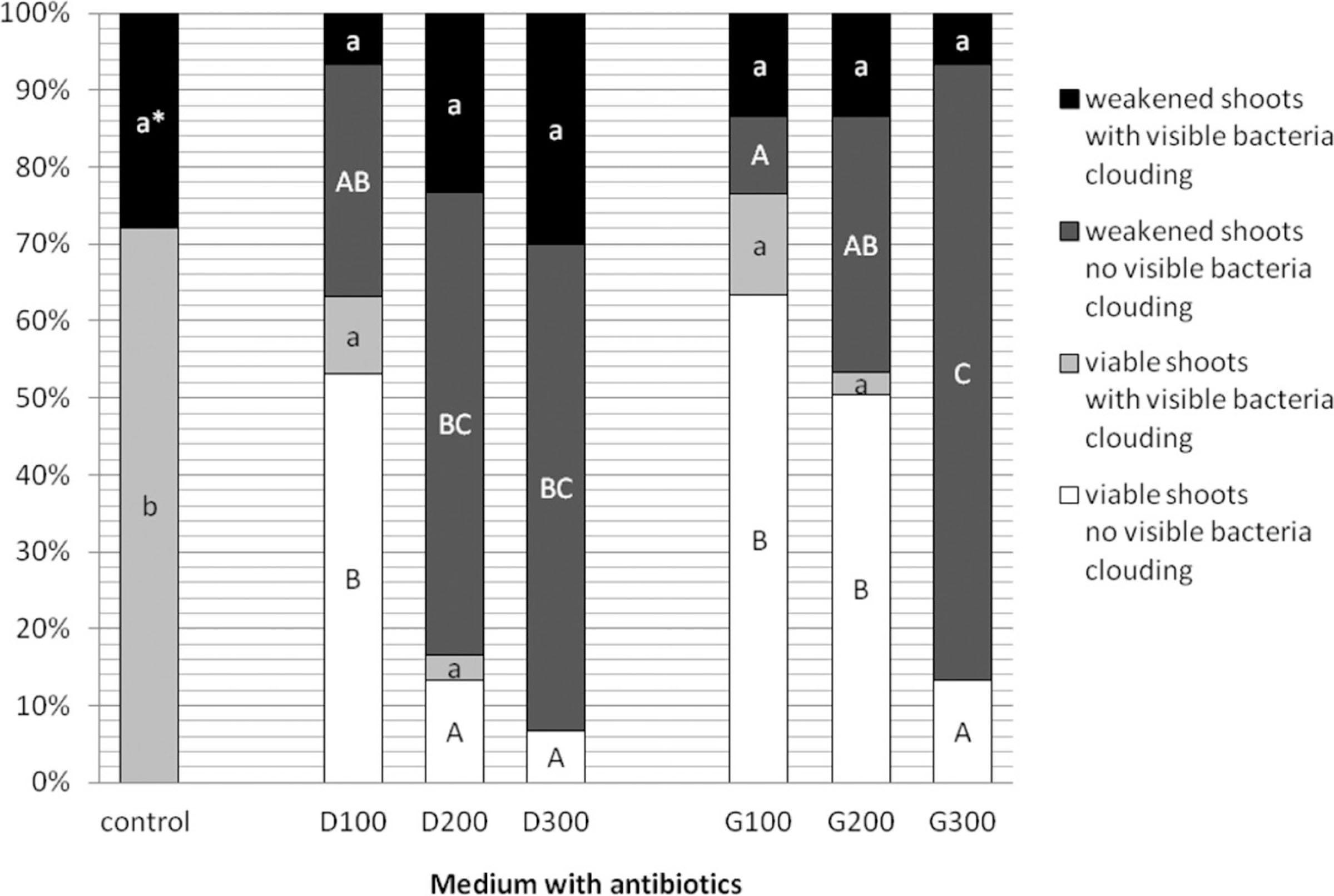
Figure 3
Basic bioinformatics parameters characterised by 16S rRNA sequencing analysis of the endogenous bacteria isolated from the Staphylea pinnata tissue culture
| Sample code | Primer | The strain with the highest compatibility | Compatibility (%) | NCBI access number |
|---|---|---|---|---|
| 16–31 | 63F | Acinetobacter johnsonii | 100 | NR_117624.1 |
| 1387R | Strain ATCC 17909 | 99 | ||
| 16–35 | 63F | Methylobacterium rhodesianum | 100 | NR_041028.1 |
| 1387R | Strain DSM 5687 | 100 |
PCR reaction mixture for amplification of gene coding 16S
| Volume (μl) | Final concentration | |
|---|---|---|
| GoTaq Green Master mix 2x | 25 | 1× |
| Upstream primer 63F | 5 | 1 μM |
| Downstream primer 1387R | 5 | 1 μM |
| Genomic DNA | 5 | < 250 ng |
| Nuclease-free water | 10 | – |
| Total | 50 |
Antibiotics susceptibility test with agar disc-diffusion method performed on bacterial strains isolated from in vitro cultures of Staphylea pinnata
| Antibiotic class | Antibiotic | AJ* | MR |
|---|---|---|---|
| β-lactams | Penicillin | R** | S |
| Amoxicillin | R | S | |
| Ampicillin+sulbactam | R | S | |
| Ampicillin | R | S | |
| Piperacillin | S | S | |
| Tikarcillin | S | S | |
| Cefuroxime | R | S | |
| Tetracyclines | Doksycycline | S | S |
| Tetracycline | S | S | |
| Fluoroquinolones | Ciprofloxacin | S | S |
| Levofloxacin | S | S | |
| Norfloxacin | S | S | |
| Ofloxacin | S | S | |
| Aminoglycosides | Amikacina | S | S |
| Gentamicin | S | S | |
| Netilmicin | S | S | |
| Tobramycin | S | S | |
| Macrolides, Lincosamides, Streptogramins | Azithromycin | R | S |
| Clarithromycin | R | S | |
| Erythromycin | R | S | |
| Clindamycin | R | R | |
| Others | Chloramphenicol | R | S |
| Nitrofurantoin | R | R | |
| Trimethoprim | R | R | |
| Co-trimoxazole | S | R | |