Today, many countries are implementing the single-step model for their national genetic evaluations of dairy cattle. The main difference between the currently used multi-step and single-step models is the incorporation of genomic information available for calculating additive genetic covariances between individuals using phenotype, genotype, and pedigree information simultaneously. The single-step model incorporates the identical-by-descent or at least identical-by-state similarity between the genotyped fraction of the evaluated individuals, expressed by the similarity of their genotypes. Genotypic information is typically expressed by single nucleotide polymorphisms (SNPs). Two equivalent statistical formulations of such a single-step model comprise the genomic best linear unbiased prediction (ssG-BLUP) (Aguilar et al., 2010; Christensen and Lund, 2010; Fragomeni et al., 2015) and ssSNP-BLUP (Liu et al., 2016; Strandén and Garrick, 2009). The only difference between these models lies in the formulation of the additive genetic covariance between individuals, in particular, between individuals with genotypes. In ssG-BLUP, the covariance matrix of breeding values of all evaluated individuals comprises the components related to non-genotyped individuals constructed using the pedigree information, and to genotyped animals constructed as a weighted sum of pedigree and SNP genotype information. In the ssSNP-BLUP, the genomic relationship between the additive genetic effects of SNPs is incorporated as a separate component in addition to a pedigree-based relationship. However, despite different parameterizations both models are equivalent mathematically (Gao et al., 2018; Koivula et al., 2012).
However, practical applications for the evaluation of large national populations of dairy cattle are computationally very challenging. Three solvers have been developed and are used for national genetic evaluations. The ssG-BLUP model can be implemented either by the APY solver (Legarra et al., 2014) or the GT solver (Legarra et al., 2014). The APY (algorithm for proven and young) procedure does not use the entire genomic relationship matrix but only a part of it based on a predefined set of genotyped individuals (so-called core individuals) while the remainder, i.e. non-core genotyped individuals is fitted as genomically uncorrelated using a diagonal genomic relationship matrix. Therefore, the system for solving model equations only requires a computationally intensive inverse of the dense genomic submatrix for the core individuals and the submatrices corresponding to the covariance between core and noncore individuals. The GT procedure implements an inverse of the genomic relationship matrix as a function of the inverse of the pedigree-based numerator relationship matrix corresponding to genotyped individuals and the SNP genotype design matrix, calculated based on the Woodbury formula. In the ssSNP-BLUP (Gao et al., 2018), due to an alternative formulation of the covariance structure, solving the system of equations does not require inverting of the partial (APY) or full (GT) relationship matrices but only a computationally simple inverse of the SNP covariance matrix.
The purpose of this study was to compare G-BLUP, GT, and SNP-BLUP solvers in terms of model validation performance to predict breeding values, differences in predicted breeding values between solvers, as well as their computational efficiency and computational resources requirements, in the context of routine genetic evaluation.
The analyzed set (Table 1) represented data from the Polish national genetic evaluation of Holstein cattle from December 2021 for stature, which represents a highly heritable trait (h2=0.54), and foot angle, representing a low heritability trait (h2=0.09). It comprised 1,098,611 cows with stature phenotypes and 1,098,766 cows with foot angle phenotypes born between 1992 and 2019 as well as 141,397 bulls (stature) and 117,482 bulls (foot angle) born between 1986 and 2017 with pseudo-phenotypes expressed by their deregressed proofs (DRP) from the multiple across country evaluation (MACE) carried out by Interbull (interbull.org). Genomic data in the form of 46,118 SNP markers were available for 134,960 individuals, including 70,134 cows with phenotypes born between 2009 and 2021, as well as 64,826 bulls born between 1985 and 2021, of which 26,471 represented young individuals without pseudo-phenotypes and 38,355 were bulls with MACE-DRP. In our study, the pedigree of animals with phenotype data was truncated after the fifth generation, resulting in 1,555,995 individuals and 33 genetic groups.
Number of animals in the analyzed data sets
| Category | Sex | Number of animals |
|---|---|---|
| Phenotype data (Stature) | Females with phenotypes | 1,098,611 |
| Males with MACE DRP | 141,397 | |
| Phenotype data (Foot angle) | Females with phenotypes | 1,098,766 |
| Males with MACE DRP | 117,482 | |
| Genotype data | Females | 70,134 |
| Males (bulls and candidates) | 64,826 | |
| Pedigree data | Females | 1,368,487 |
| Males | 187,508 |
The following single-step single-trait models were considered for the prediction of GEBV.
The ssG-BLUP (Christensen and Lund, 2010):
The ssSNP-BLUP (Gao et al., 2018):
Variance components underlying the analyzed phenotypes
| Trait | σa2 | σe2 | σu2 |
|---|---|---|---|
| Stature | 5.50 | 4.63 | 20% |
| Foot angle | 0.11 | 1.06 | 20% |
Solutions for the effects fitted in the above models were obtained using the MiXBLUP 3.0 software (Liu et al., 2014; Vandenplass, 2022) that solves the following equation: D−1 M−1 Cx = D−1 M−1 p where C represents the coefficient matrix corresponding to the Mixed Model Equations (MME) for solving (1) or (2), x is the vector of model parameters, and p is the RHS of MME, while M and D respectively represent the first-level and the second-level preconditioning matrices. The computations were performed on a dedicated computing server running the Linux Red Hat operating system with 260GB of RAM, 16 Intel Xeon CPUs with 2.20GHz, and 600GB of hard disk space. For a large national dairy population, such as the one used in our study, the numerically severe challenge is to obtain the inverse of the HG and HS matrix, in particular of its component related to genotyped individuals – the dense sub-matrix G. In our study, the following approaches were considered.
The GT approach (Lourenco et al., 2020; Mäntysaari et al., 2020), in which the G−1 matrix is represented by
The APY approach (Legarra et al., 2014) that divides the genotyped individuals into a core and non-core sub-groups for which the inverse is handled differently in such a way that the exact inverse is computed only for the core animal sub-group (Gc) and the covariance between core and non-core individuals, while the part of the matrix corresponding to the non-core genotyped animals (Gn) is handled as a diagonal matrix:
The ssSNP-BLUP approach (Gao et al., 2018) does not meet the numerical burden of the models based on G, since in terms of the modelling of genomic information only the inverse of the diagonal matrix B is required:
For the validation of GEBV prediction, bulls born after 2013 (7,296 bulls for stature; 6,200 bulls for foot angle) and cows born after 2016 (96,772 cows for stature; 96,771 cows for foot angle) were removed from the phenotype vector (y) of equations (1) and (2). Based on such truncated data sets, the GEBVs of bulls with EDC greater than 20 were predicted by models (1) and (2). Prediction accuracy was assessed by a weighted linear regression (Mäntysaari et al., 2010): GEBVf = b0 + b1GEBVt + e, with GEBVf representing the vector of GEBVs predicted based on the full data set with all available individuals while GEBVt contains the GEBVs predicted based on the truncated data set. For i-th bull, weights were defined as
The above linear regression equation was fitted using the lm function and Pearson correlation coefficients, used to assess differences between GEBVs across scenarios were calculated using R software (www.rstudio.com).
The validation data set for stature and foot angle comprised 1,727 and 1,725 bulls with EDC > 20, respectively. For stature, generally, no marked differences in the estimated slope of the linear regression were observed between the models, varying between 0.94 (APY3000top) and 1.02 (APY10000random). Furthermore, except for APY3000, the differences between R2 corresponding to each model were small. For stature, the GT approach achieved the highest R2 of 0.83, while the lowest R2 corresponded to 3,000 core animal scenarios with 0.57 for APY3000top and 0.60 for APY3000random. Similar results were observed for foot angle, albeit with overall lower R2. GT and SNP-BLUP achieved the highest value of 0.76, while the lowest of 0.60 was attributed to APY3000random and 0.62 for APY3000top. Details of the validation results are summarized in Table 3. Pearson correlations (Figure 1) between GEBVs predicted for the validation bulls from the full and the truncated datasets were calculated for all pairs of models. For stature, they demonstrated a good agreement only between SNP-BLUP and GT, as expressed by correlations of 0.996. Interestingly, the lowest correlations of 0.810 were observed between both models implementing APY3000 (i.e. APY3000random and APY3000top), as well as between APY3000top and APY15000top. For foot angle correlations are generally higher, and the pattern is very similar, with the correlation between SNP-BLUP and GT reaching 0.999 and the lowest correlation of 0.870 observed between both APY3000 scenarios, between GT and APY3000top, as well as between GP and APY3000random. Notably, for both traits, the correlations within APY3000top and within APY3000random are markedly lower than for the other models
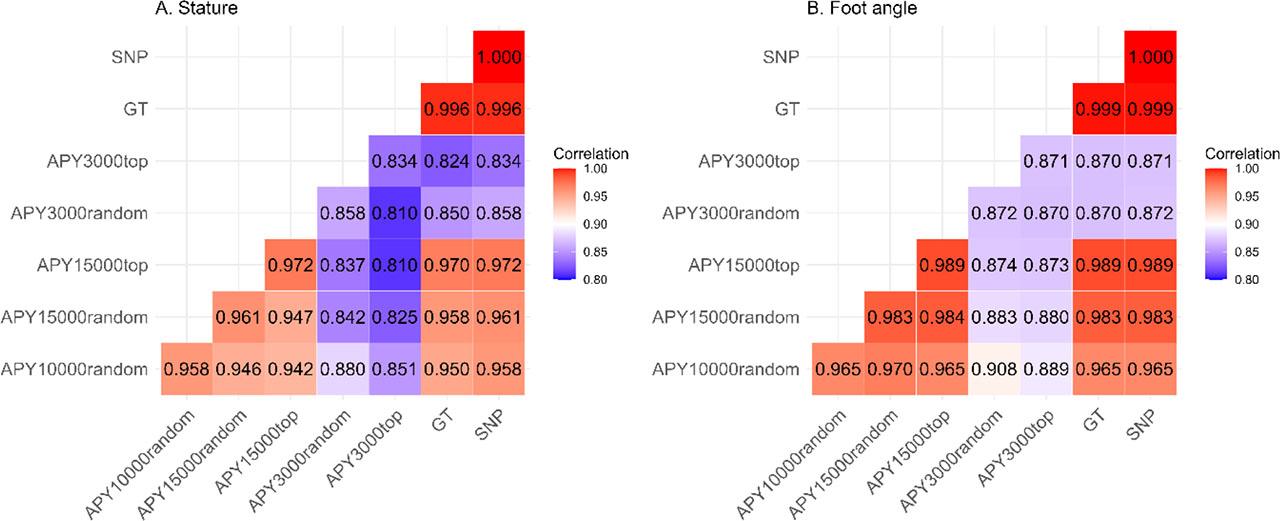
Pearson correlation coefficients of GEBVs predicted by different model variants
Validation of GEBV predictions
| Model variants |
|
|
|
| R2 |
|---|---|---|---|---|---|
| Stature, 1,727 validation bulls, h2 = 0.54 | |||||
| SNP-BLUP | −3.90 | 0.32 | 1.01 | 0.01 | 0.77 |
| GT | −4.40 | 0.27 | 1.01 | 0.01 | 0.83 |
| APY3000top | −1.81 | 0.44 | 0.94 | 0.02 | 0.57 |
| APY3000random | −2.20 | 0.44 | 0.99 | 0.02 | 0.60 |
| APY10000random | −3.74 | 0.36 | 1.02 | 0.02 | 0.72 |
| APY15000top | −2.59 | 0.32 | 0.96 | 0.01 | 0.75 |
| APY15000random | −2.32 | 0.33 | 0.96 | 0.01 | 0.73 |
| Foot angle, 1,725 validation bulls, h2 = 0.09 | |||||
| SNP-BLUP | −2.03 | 0.18 | 1.03 | 0.01 | 0.76 |
| GT | −1.96 | 0.18 | 1.04 | 0.01 | 0.76 |
| APY3000top | −0.68 | 0.21 | 0.97 | 0.02 | 0.62 |
| APY3000random | −0.52 | 0.26 | 1.03 | 0.02 | 0.60 |
| APY10000random | −2.04 | 0.21 | 1.02 | 0.02 | 0.69 |
| APY15000top | −2.15 | 0.18 | 1.02 | 0.01 | 0.75 |
| APY15000random | −2.10 | 0.19 | 1.03 | 0.02 | 0.73 |
Figure 2 shows the differences between the GEBVs predicted by ssSNP-BLUP compared to the GEBVs predicted by the six ssG-BLUP implementations. For the difference in GEBV between SNP-BLUP and GT, all genotyped animals were considered, while for all comparisons involving APY, only the core individuals were used. The original solutions were rescaled for each model by subtracting the mean of a cow base population to provide GEBVs comparable across all models. For stature, the differences were generally small across bull birth years (2A), however, for foot angle a different pattern emerged (2B) as for all comparisons, except GT and the most informative APY15000top, the differences were unstable across bull birth years. Furthermore, we compared correlations of GEBVs predicted by the full and the truncated models for individuals representing base cows (i.e. cows with phenotype records) and bulls (i.e. bulls, which have a minimum of one daughter). The dashed line divides the plot into animals defined as old and young in the validation process. For stature, regardless of the model, correlations between full and truncated data sets, for old animals, were very close to unity. For young candidates, we observed a declining trend with the lowest correlations calculated for APY3000random (Figure 3 A). For foot angle, similar results were obtained, except for APY3000random for females, where we observed a decreasing correlation trend as early as from 2004. From 2008, correlations dropped below 0.9 (Figure 3 B). Finally, we compared the overlap of bulls with the top 50 GEBV predictions resulting from different models (Figure 4). The number of bulls common across all models for stature (Figure 4 A) was 39, while the highest number of exclusive bulls (3) was observed in the top 50 list resulting from the APY15000top. Regarding foot angle (Figure 4 B), the number of common bulls was even lower, being 31, while 8 were exclusive for the APY3000random.
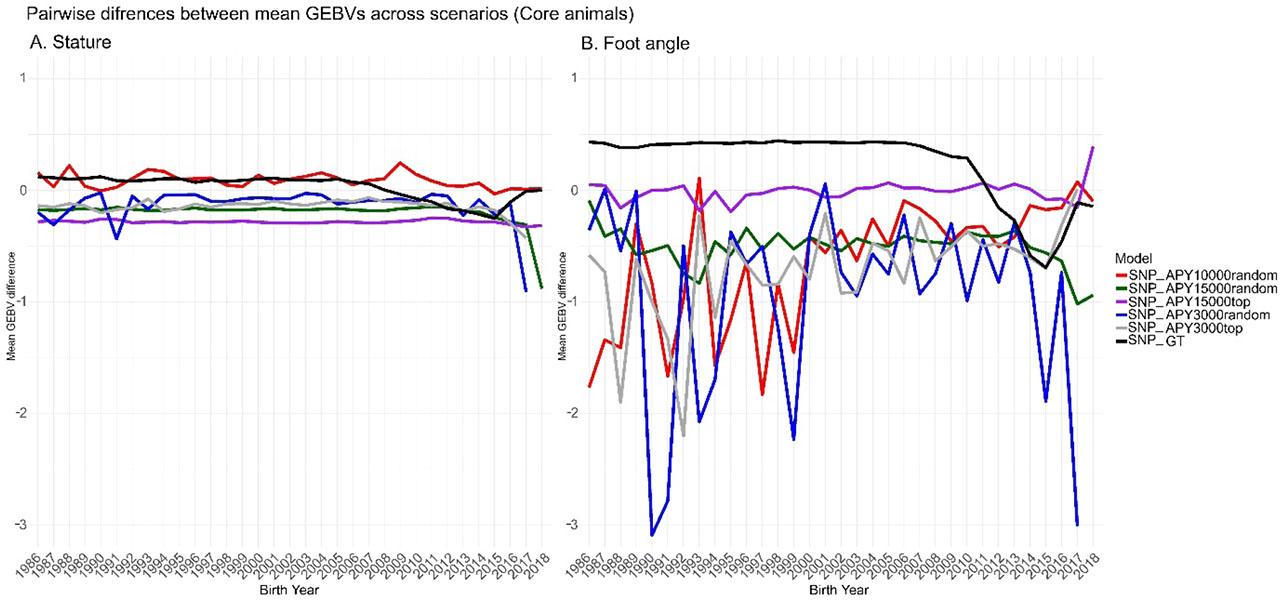
Mean GEBV difference divided by the genetic standard deviation between SNP-BLUP and GT, as well as SNP-BLUP and APY models implementing different scenarios of core individual selection. (A) stature, (B) foot angle
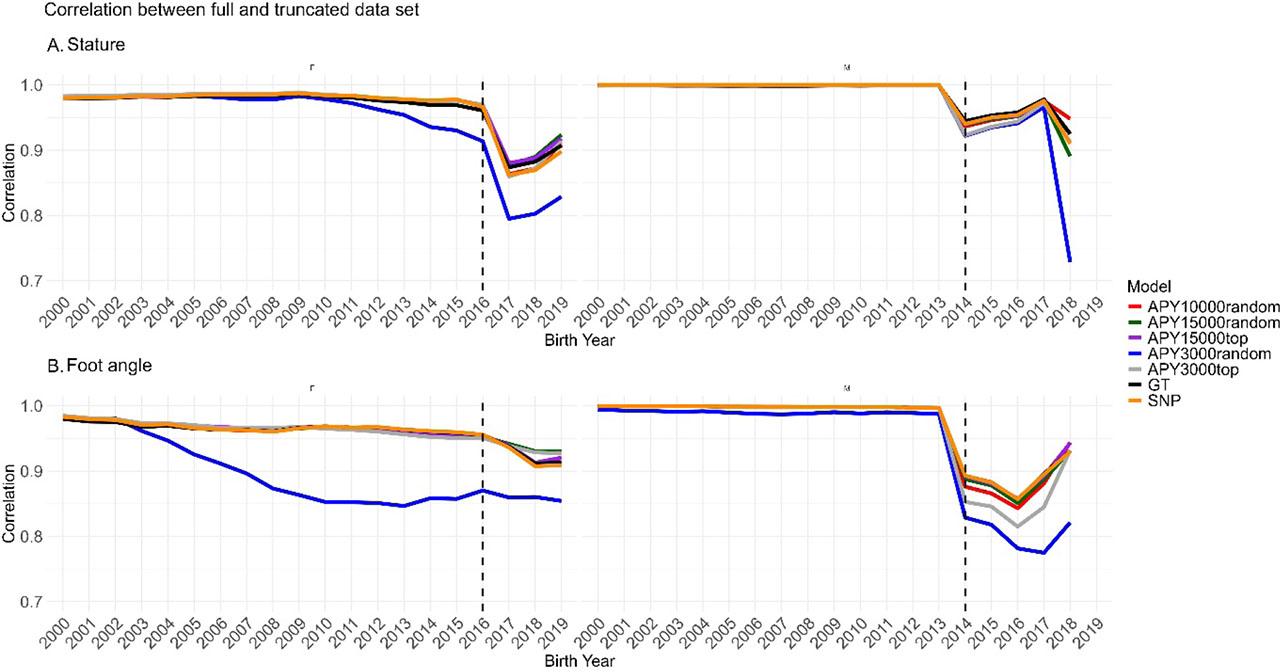
Correlation between full and truncated data set for cows with phenotype records and bulls with a minimum of one daughter. (A) stature, (B) foot angle
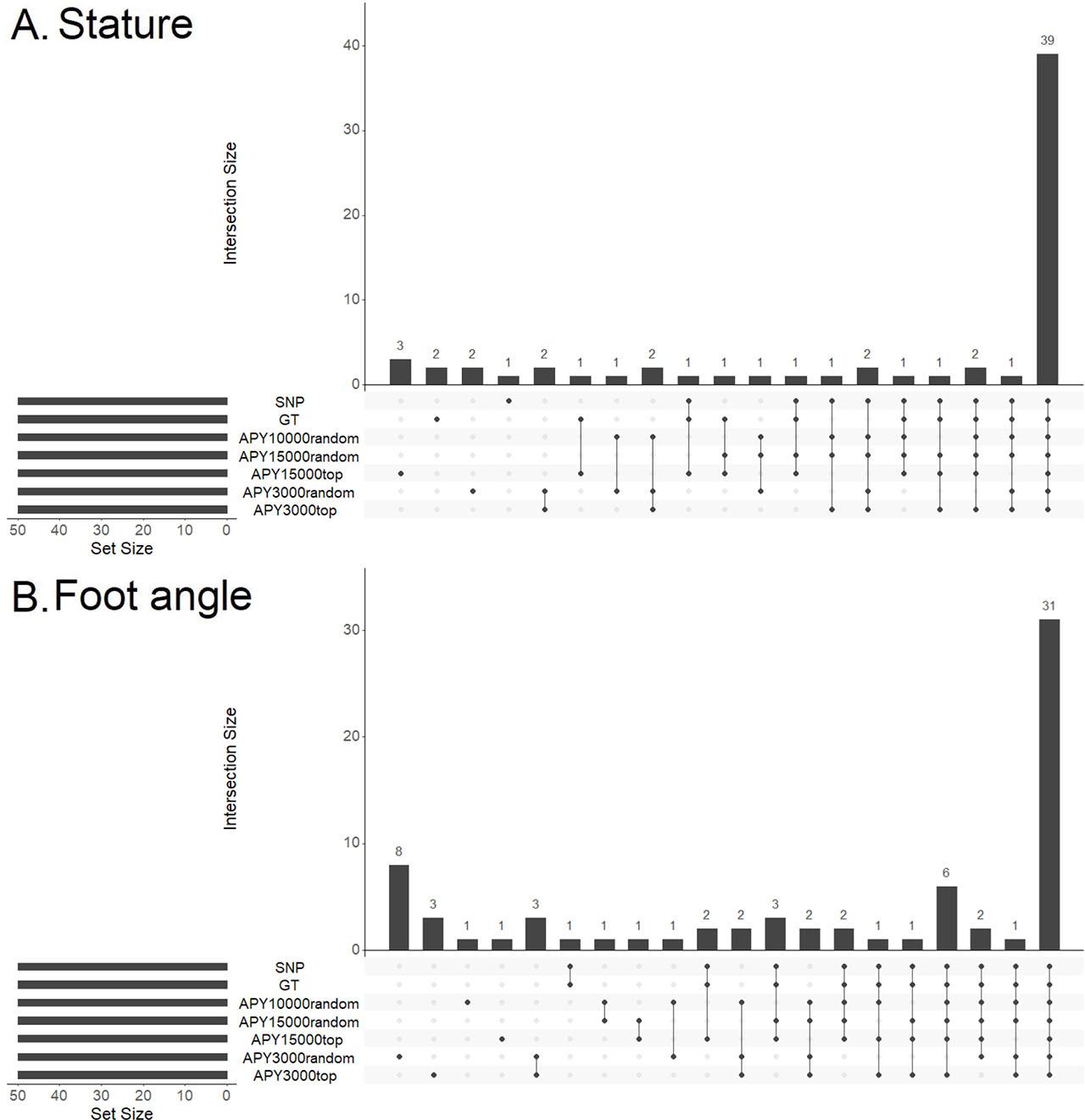
An upset plot of bulls with the top 50 GEBVs. (A) stature, (B) foot angle
The wall clock time corresponding to setting up and solving models (1) or (2) using the MiXBLUP software and the peak memory consumption varied considerably between solvers. The exact values are specified in Table 4, but generalizing for both traits, SNP-BLUP and APY3000 were the fastest, closely followed by APY10000 random. The wall clock time of GT was the longest, twice the time of APY15000top. The peak memory consumption by SNP-BLUP was on the order of ten times lower than for the remaining solvers. In the case of iterations, SNP-BLUP required the most iterations (673 for stature and 1,027 for foot angle) and an average of 2.3 seconds per iteration. The lowest number of 335 iterations and an average of 0.18 seconds per iteration were used for stature by APY3000top and 499 and 0.18 seconds for foot angle by APY3000random.
Computational resources utilized by the solvers
| Model variants | Wall clock time (min) | Peak RAM consumption (GB) | Number of iterations | |||
|---|---|---|---|---|---|---|
| stature | foot angle | stature | foot angle | stature | foot angle | |
| SNP-BLUP | 23 | 32 | 5.81 | 5.81 | 673 | 1027 |
| GT | 138 | 143 | 63.89 | 63.88 | 477 | 629 |
| APY3000top | 23 | 29 | 49.48 | 49.47 | 335 | 811 |
| APY3000random | 23 | 32 | 49.48 | 49.47 | 390 | 499 |
| APY10000random | 32 | 34 | 56.53 | 56.53 | 469 | 652 |
| APY15000top | 68 | 70 | 61.56 | 61.56 | 425 | 625 |
| APY15000random | 54 | 57 | 61.56 | 61.56 | 477 | 551 |
Many studies have been conducted related to comparing single-step genomic prediction models with con-ventional two-step approaches. However, the literature on comparisons within the single-step modelling frame is scarce. Koivula et al. (2012) considered genomic prediction models similar to (1) and (2). However, they addressed only the genotyped part of the population. Similarly to the results of our study, these authors observed very high correlations between models for predicted bull GEBVs or DGVs (direct genetic values) and similar validation results. Gao et al. (2018) also considered the validation performance of single-step models (Masuda et al., 2016). Although no marked differences were observed, the authors indicated that for APY-based solvers, not only the number but also the selection of core individuals was a crucial step influencing the prediction accuracy (Masuda et al., 2016; Misztal et al., 2016, 2014; Strandén and Garrick, 2009). Except worse prediction performance of APY3000random for cows, no marked differences due to the composition of the core animal set were observed in our study. Still, Macedo et al. (2022) demonstrated low robustness in predictions with ssG-BLUP model towards differential handling of missing parent information. Moreover, our results demonstrated that a large number of core individuals is recommended, provided computational resources, especially RAM, are available, to obtain stable prediction with ssG-BLUP. Similarly to the results of Misztal et al. (2014), we also did not observe a marked gain in the prediction quality of GEBV when using more than 10,000 core individuals.
Regarding the computational aspects, we observed very large differences in RAM consumption between SNP-BLUP and other solvers. GT needed over 10 times more RAM and fewer iterations than SNP-BLUP, as was also demonstrated by Vandenplas et al. (2023). Two possible ways to circumvent the problem of the optimal choice of core individuals are either the use of GT or SNP-BLUP solver-based prediction. The application of GT comes with the price of high memory requirements and long computing times. SNP-BLUP does not consume much memory and is computationally more efficient. Still the approaches did not yield identical rankings of top 50 bulls. Although 62% (foot angle) and 75% (stature) bulls were common in rankings across scenarios, there was still a considerable number of individuals that were ranked scenarios specific.
Regarding the prediction of GEBV on the active population scope, no marked differences between solvers (except the APY with only 3,000 core individuals) were observed, still GT and SNP-BLUP solvers resulted in the highest prediction accuracy, while the effectiveness of the APY model depends on the size of the set of core animals: small cores lead to a noticeable decrease in accuracy, while cores with large numbers minimize these differences. Another factor influencing the choice of the solver is its computational efficiency expressed by the computation time and memory resources required, which was also indicated by Koivula et al. (2012). However, it should be kept in mind that the ranking of the top bulls is not identical between models, which has implications for the breeding industry in terms of semen pricing and selection since, typically, the top-ranking bulls are the most widely used.
Considering the abovementioned aspects, the results of our study indicate that GT and SNP-BLUP solvers offer more modelling and computational advantages.