Fig. 1.

Fig. 2.

Fig. 3.
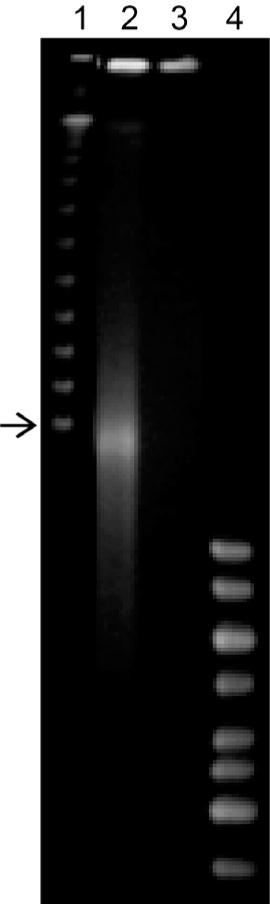

Fig. 4.
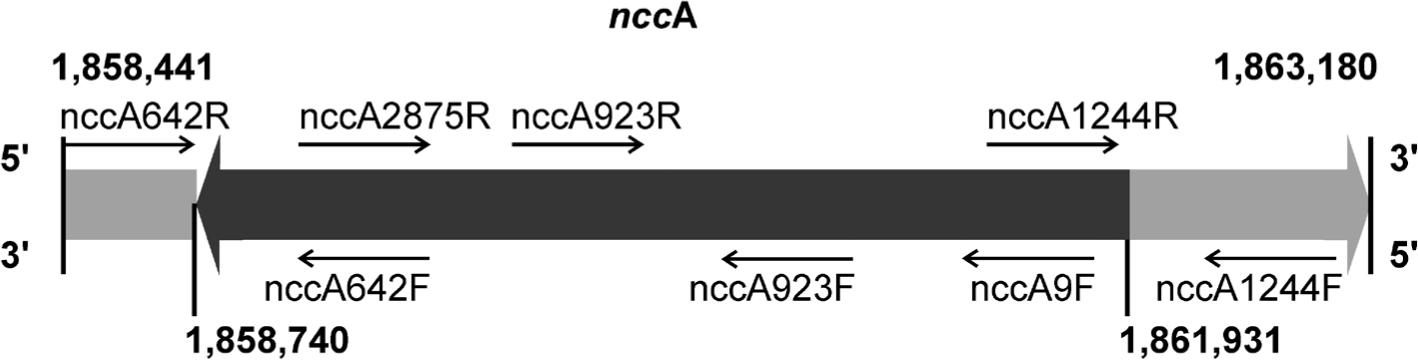
Fig. 5.

Primer sets used in this study_
| Probes | Sequence | Descriptiona |
|---|---|---|
| 27F | 5’ AGAGTTTGATCCTGGCTCAG 3’ | 16S rDNA universal primers, positions 8–27 and 704–685 and 1512–1492, resp. in the E. coli K12 [NC_000913] numbering system; (Lane, 1991) |
| 1492R | 5’ ACGGCTACCTTGTTACGACTT 3’ | |
| 2555nccAF | 5’ AGCCG (C,G) GA (C,G) AACGG CAAGCG 3’ | 2536–2555 and 3136–3117 degenerative nccA primers, positions on plazmid p9 in the Achromobacter xylosoxidans 31A [L31363] numbering system; (Karelová et al., 2011) |
| 3117nccAR | 5’ CCGATCACCACCGT (T,C) GC CAG 3’ | |
| nccA9F | 5’ ACGTATCATTAGTTTCGCCA 3’ | 1861923–1861904 and 1859057–1859076 nccA primers, positions on chromosome in the Ralstonia picketii 12J [CP001068] numbering system; This work |
| nccA2875R | 5’ ATCGGATAAACGACAGCATC 3’ | |
| nccA1244F | 5’ GCTCTCGAAAGAGGAAGGCA 3’ | 1862989–1862970 and 1861746–1861765 nccA primers, positions on chromosome in the Ralstonia picketii 12J [CP001068] numbering system; This work |
| nccA1244R | 5’ TTCGGTTTCGAGCGGTGAAT 3’ | |
| nccA642F | 5’ GCTAGTCTTCACGGGCATT 3’ | 1859211–1859193 and 1858570–1858589 nccA primers, positions on chromosome in the Ralstonia picketii 12J [CP001068] numbering system; This work |
| nccA642R | 5’ GCTCTTCGTCATGACACCAC 3’ | |
| nccA923F | 5’ GGTCGCTTCCATTAACCG 3’ | 1860996–1860979 and 1860074–1860091 nccA primers, positions on chromosome in the Ralstonia picketii 12J [CP001068] numbering system; This work |
| nccA923R | 5’ GATCGGATGCAATCTCCG 3’ | |
| nccA-F | 5’ GTCGCCTTGTTCATCGG 3’ | 1860425–1860409 and 1860301–1860319 nccA primers, positions on chromosome in the Ralstonia picketii 12J [CP001068] numbering system; This work |
| nccA-R | 5 GCAAACGTCAATACAACGG 3’ | |
| gdhA-F | 5’ CGTACTCAATGAACGAAGGC 3’ | 388722–388741 and 388866–388850 gdhA primers, positions on chromosome in the Ralstonia picketii 12D [CP001644] numbering system; This work |
| gdhA-R | 5’ TCGATGCCGAGATTGCG 3’ | |
| UP1 | 5’ GAAGTCATCATGACCGTTCTG CA(TC)GC(TCAG)GG(TCAG)GG (TCAG)AA(AG)TT(TC)GA 3’ | gyrB gene primers, positions 91–104 and 495–509 amino acid residues (the numbering corresponds to that of the E. coli K12 protein [(GYRB_ECOLI in the SWISS-PROT database)]) (Yamamoto and Harayama, 1995) |
| UP2r | 5’ AGCAGGGTACGGATGTGCGAG CC(AG)TC(TCAG)AC(AG)TC(TC AG)GC(AG)TC(TCAG)GTCAT 3’ | |
| UP-1S | 5’ GAAGTCATCATGACCGTTCT GCA 3’ | |
| UP-2Sr | 5’ AGCAGGGTACGGATGTGCG AGCC 3’ |
Expression of MR-CH-I2-nccA [KR476581] gene after heavy metal additions to the medium_
| Time (h) | Nickela | Cadmiuma | Cobalta | Coppera | Zinca | |||||
|---|---|---|---|---|---|---|---|---|---|---|
| ΔΔCt | RQ | ΔΔCt | RQ | ΔΔCt | RQ | ΔΔCt | RQ | ΔΔCt | RQ | |
| 0 | 0.00 | 1.00 | 0.00 | 1.00 | 0.00 | 1.00 | 0.00 | 1.00 | 0.00 | 1.00 |
| 2 | –4.08 | 16.91 | 0.53 | 0.69 | 3.23 | 0.11 | 5.50 | 0.02 | 1.81 | 0.29 |
| 4 | –0.17 | 1.13 | 6.32 | 0.01 | 5.18 | 0.03 | 6.61 | 0.01 | 3.17 | 0.11 |
| 6 | –0.51 | 1.42 | 5.82 | 0.02 | 5.22 | 0.03 | 9.38 | 0.002 | 3.65 | 0.08 |
| 8 | 0.67 | 0.63 | 6.13 | 0.01 | 7.61 | 0.005 | 7.52 | 0.005 | 2.81 | 0.14 |